An updated version of the SHAP-based Boruta feature selection algorithm with some new features.
Reason this release was yanked:
First Project. Minor erros in readme
Project description
Boruta-Shap-Plus
This is a fork of Boruta-Shap by Eoghan Keany. The original library was not maintained anymore, and this fork includes the following updates:
- Updated to use the latest versions of numpy and scipy. The original library had some depreciated imports that threw errors when used with newer versions of those libraries.
- Added the ability to do a time-aware train-test split.
- Made test set size parameterizable.
Boruta-Shap
BorutaShap is a wrapper feature selection method which combines both the Boruta feature selection algorithm with shapley values. This combination has proven to out perform the original Permutation Importance method in both speed, and the quality of the feature subset produced. Not only does this algorithm provide a better subset of features, but it can also simultaneously provide the most accurate and consistent global feature rankings which can be used for model inference too. Unlike the orginal R package, which limits the user to a Random Forest model, BorutaShap allows the user to choose any Tree Based learner as the base model in the feature selection process.
Despite BorutaShap's runtime improvements the SHAP TreeExplainer scales linearly with the number of observations making it's use cumbersome for large datasets. To combat this, BorutaShap includes a sampling procedure which uses the smallest possible subsample of the data availble at each iteration of the algorithm. It finds this sample by comparing the distributions produced by an isolation forest of the sample and the data using ks-test. From experiments, this procedure can reduce the run time up to 80% while still creating a valid approximation of the entire data set. Even with these improvments the user still might want a faster solution so BorutaShap has included an option to use the mean decrease in gini impurity. This importance measure is independent of the size dataset as it uses the tree's structure to compute a global feature ranking making it much faster than SHAP at larger datasets. Although this metric returns somewhat comparable feature subsets, it is not a reliable measure of global feature importance in spite of it's wide spread use. Thus, I would recommend to using the SHAP metric whenever possible.
Algorithm
-
Start by creating new copies of all the features in the data set and name them shadow + feature_name, shuffle these newly added features to remove their correlations with the response variable.
-
Run a classifier on the extended data with the random shadow features included. Then rank the features using a feature importance metric the original algorithm used permutation importance as it's metric of choice.
-
Create a threshold using the maximum importance score from the shadow features. Then assign a hit to any feature that had exceeded this threshold.
-
For every unassigned feature preform a two sided T-test of equality.
-
Attributes which have an importance significantly lower than the threshold are deemed 'unimportant' and are removed them from process. Deem the attributes which have importance significantly higher than than the threshold as 'important'.
-
Remove all shadow attributes and repeat the procedure until an importance has been assigned for each feature, or the algorithm has reached the previously set limit of runs.
If the algorithm has reached its set limit of runs and an importance has not been assigned to each feature the user has two choices. Either increase the number of runs or use the tentative rough fix function which compares the median importance values between unassigned features and the maximum shadow feature to make the decision.
Installation
Use the package manager pip to install BorutaShap.
pip install BorutaShap
Usage
For more use cases such as alternative models, sampling or changing the importance metric please view the notebooks here.
Using Shap and Basic Random Forest
from BorutaShap import BorutaShap, load_data
X, y = load_data(data_type='regression')
X.head()
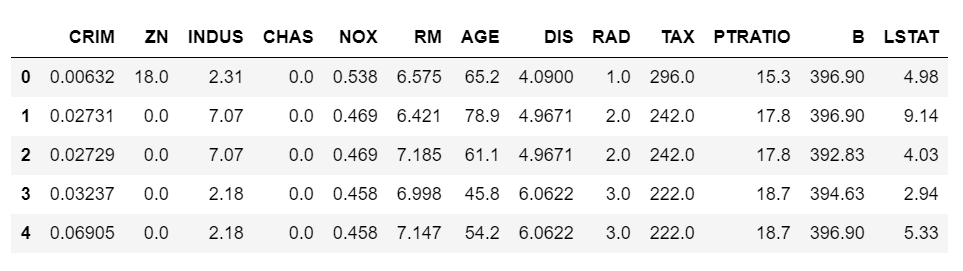
# no model selected default is Random Forest, if classification is True it is a Classification problem
Feature_Selector = BorutaShap(importance_measure='shap',
classification=False)
'''
Sample: Boolean
if true then a rowise sample of the data will be used to calculate the feature importance values
sample_fraction: float
The sample fraction of the original data used in calculating the feature importance values only
used if Sample==True.
train_or_test: string
Decides whether the feature improtance should be calculated on out of sample data see the dicussion here.
https://slds-lmu.github.io/iml_methods_limitations/pfi-data.html
normalize: boolean
if true the importance values will be normalized using the z-score formula
verbose: Boolean
a flag indicator to print out all the rejected or accepted features.
'''
Feature_Selector.fit(X=X, y=y, n_trials=100, sample=False,
train_or_test = 'test', normalize=True,
verbose=True)

# Returns Boxplot of features
Feature_Selector.plot(which_features='all')
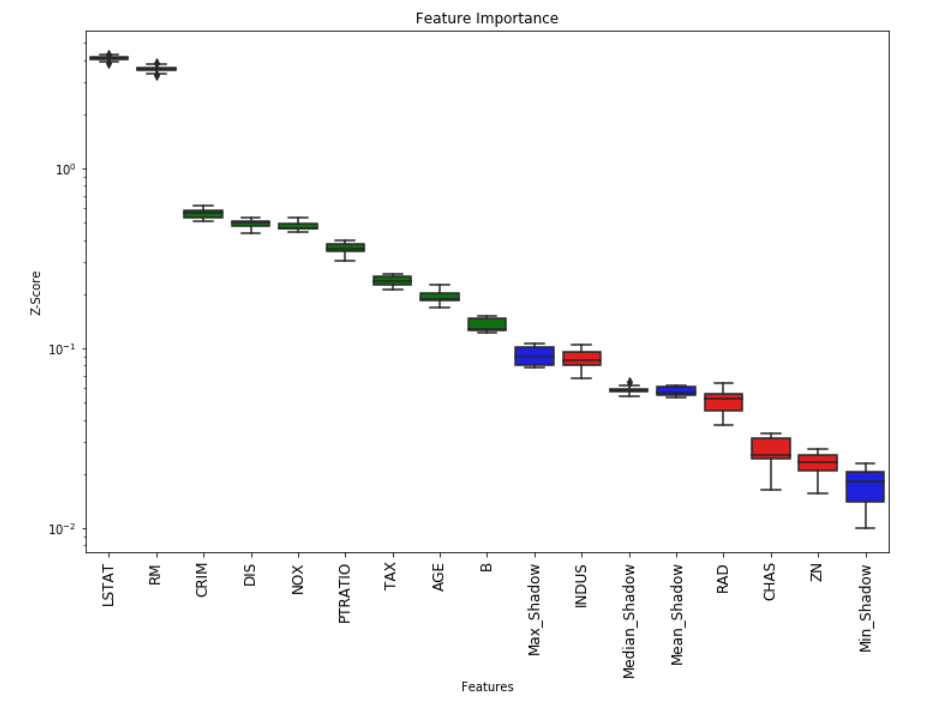
# Returns a subset of the original data with the selected features
subset = Feature_Selector.Subset()
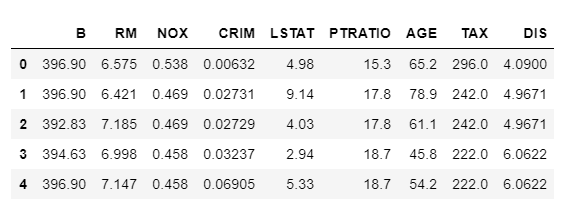
Using BorutaShap with another model XGBoost
from BorutaShap import BorutaShap, load_data
from xgboost import XGBClassifier
X, y = load_data(data_type='classification')
X.head()
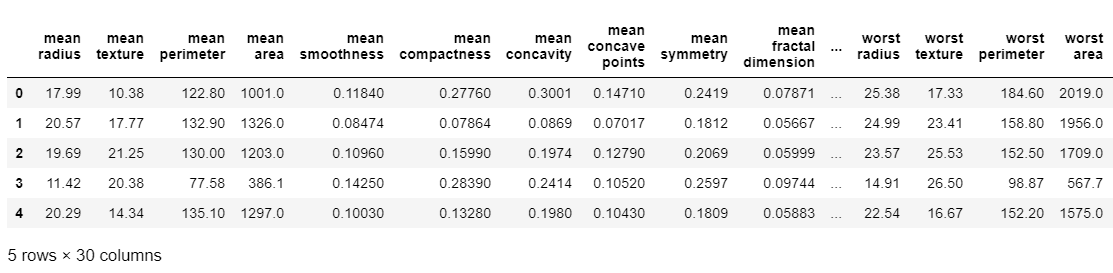
model = XGBClassifier()
# if classification is False it is a Regression problem
Feature_Selector = BorutaShap(model=model,
importance_measure='shap',
classification=True)
Feature_Selector.fit(X=X, y=y, n_trials=100, sample=False,
train_or_test = 'test', normalize=True,
verbose=True)
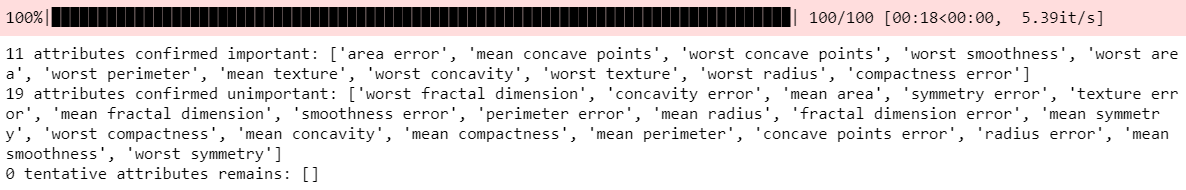
# Returns Boxplot of features
Feature_Selector.plot(which_features='all')
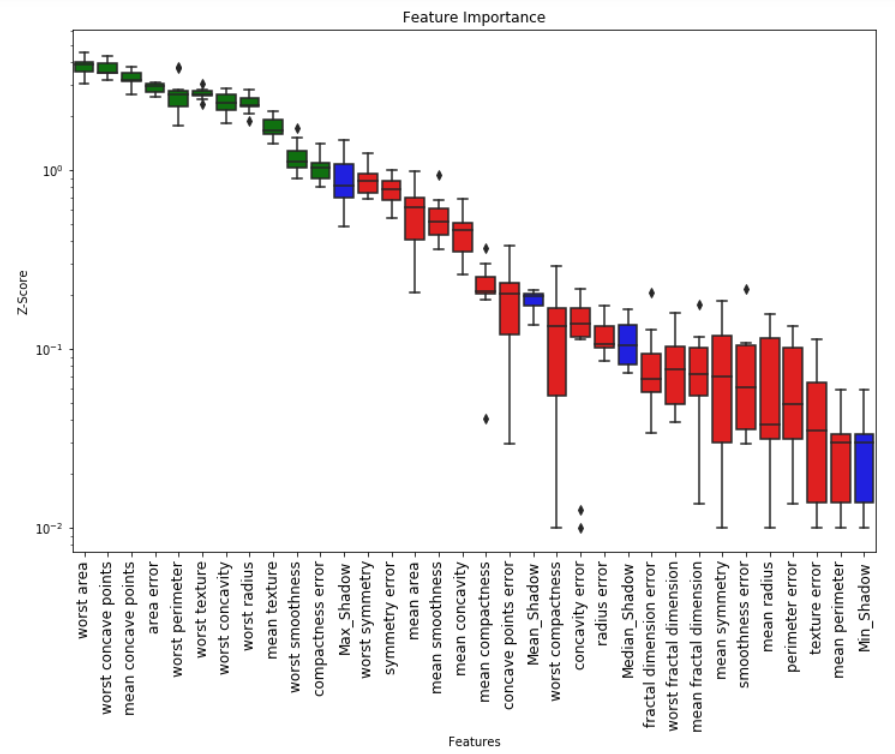
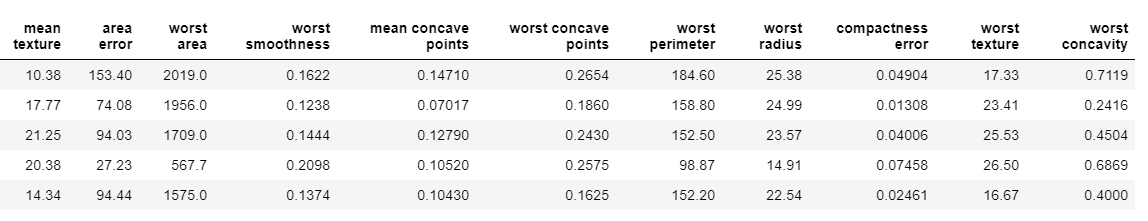
# Returns a subset of the original data with the selected features
subset = Feature_Selector.Subset()
Contributing
Pull requests are welcome. For major changes, please open an issue first to discuss what you would like to change.
License
If you wish to cite this work please click on the zenodo badge at the top of this READme file MIT
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file borutashapplus-0.1.2.tar.gz.
File metadata
- Download URL: borutashapplus-0.1.2.tar.gz
- Upload date:
- Size: 18.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.10.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a32d2709b76d1d95792025a817152bf3c2eb6adaf8c7b406f9b077e464039f38
|
|
| MD5 |
b8c91ca32cc3e3af3efa2ac733c04abf
|
|
| BLAKE2b-256 |
d49b245323c23cc5ebd119fb9ce79ce0e51d0aff787ec6b04ba22c71e96eed25
|
File details
Details for the file BorutaShapPlus-0.1.2-py3-none-any.whl.
File metadata
- Download URL: BorutaShapPlus-0.1.2-py3-none-any.whl
- Upload date:
- Size: 15.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.10.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ba29866a0e5b09a2f2773f479f2d87e3455f7409953f0a3ead5560bf1f2dd21c
|
|
| MD5 |
9c06ae953b2ecb30fecd09ec860f8410
|
|
| BLAKE2b-256 |
f256895f75524753eba45caab9afcbf33cf9423ca588a97c3ec420340daf024e
|












