Helper utilities for the Chemical Functional Use Taxonomy SQLite database.
Project description
Chemical Function Taxonomy (ChemFuncT)
The Analytical Methods and Open Spectra (AMOS) Database's Chemical Function Taxonomy (ChemFuncT) contains mappings between chemicals (via name and DTXSID) and their functional uses. This repository contains a snapshot of the SQLite database and a Python class to help query the database.
Sources
This dataset was compiled using information from:
- Wikipedia – General-purpose encyclopedia entries on chemical uses.
- ChemExpo – A web application that surfaces reported chemical function use from the EPA's CPDat database.
- DrugBank – Pharmaceutical chemical uses and mechanisms.
- APPRIL – The EPA's Active Pesticide Product Registration Informational Listing.
ChemFuncT Database
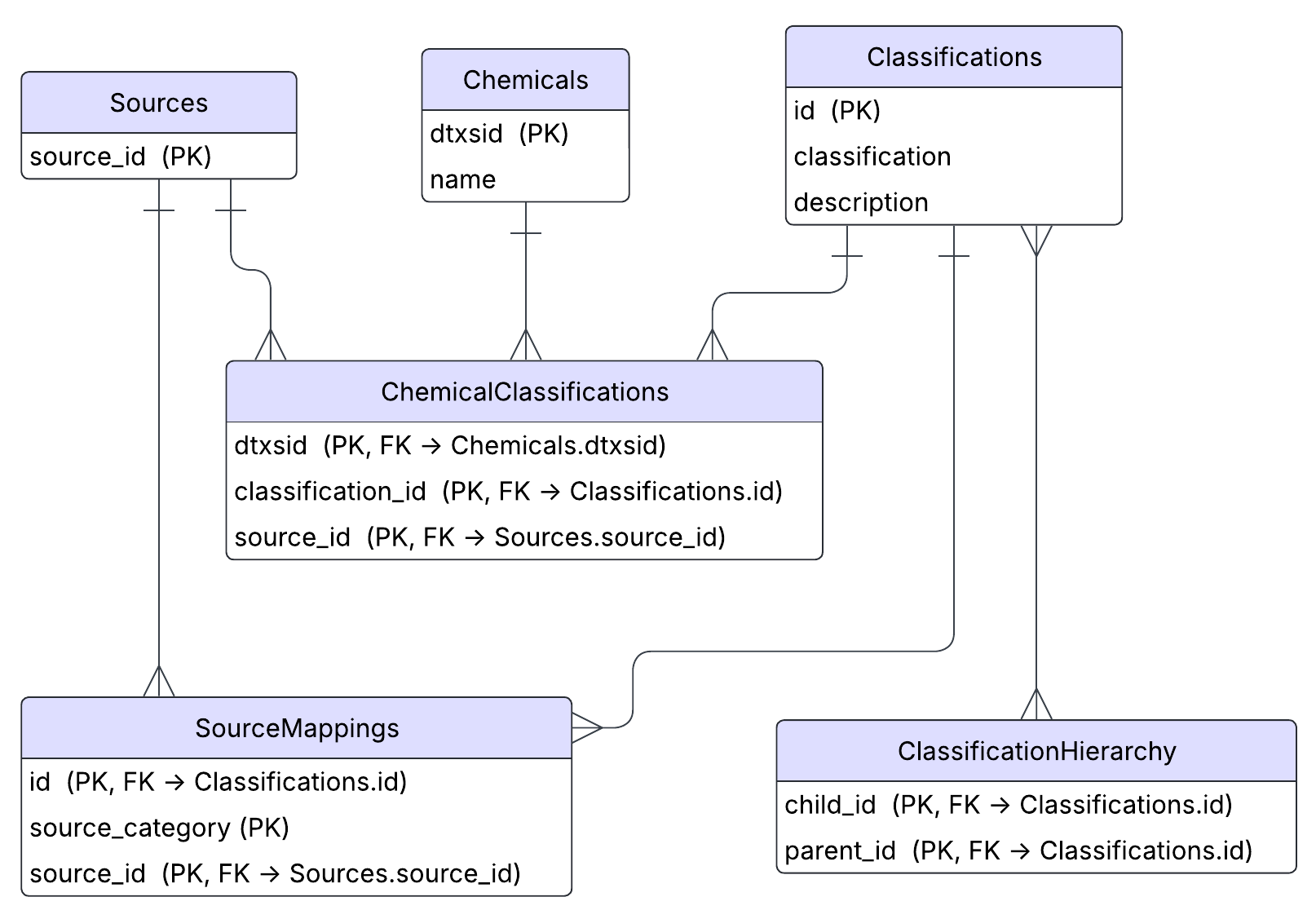
A snapshot of the data is contained in data/ChemFuncT.db. Here is an ER diagram representation of the database.
Descriptions of each table and their variables follow:
| Table | Description |
|---|---|
| Sources | Contains unique identifiers for each source that data was pulled from. |
| Chemicals | Maps the unique chemical identifier (DTXSID) to its chemical name. |
| Classifications | Maps a unique classification ID to a human readable class name (e.g., Pharmaceuticals) and a description of that class. |
| ChemicalClassifications | Maps chemicals (by DTXSID) to their classifications, and which source that classification came from. |
| SourceMappings | Contains the raw category pulled from a given source mapped to a harmonized ChemFuncT class. |
| ClassificationHierarchy | Contains parent/child mappings for each classification. |
Sources Table
| Variable | Role | Description |
|---|---|---|
| source_id | PK | A unique identifier for each source. E.g., "wikipedia", "drugbank". |
Chemicals Table
| Variable | Role | Description |
|---|---|---|
| dtxsid | PK | The unique DTXSID for each chemical. |
| name | The EPA's preferred name for each chemical. |
Classifications Table
| Variable | Role | Description |
|---|---|---|
| id | PK | A unique identifier for each classification of the form func_0001. |
| classification | A human readable name for each classification (e.g., Pharmaceuticals). | |
| description | A description for each classification. |
ChemicalClassifications Table
| Variable | Role | Description |
|---|---|---|
| dtxsid | CPK, FK $\rightarrow$ Chemicals.dtxsid |
The unique DTXSID chemical identifier. |
| classification_id | CPK, FK $\rightarrow$ Classifications.id |
A unique identifier for each classification. |
| source_id | CPK, FK $\rightarrow$ Sources.source_id |
The unique source_id for each source. |
SourceMappings Table
| Variable | Role | Description |
|---|---|---|
| id | CPK, FK $\rightarrow$ Classifications.id |
A unique identifier for each classification. |
| source_category | CPK | The raw category pulled from the source that was mapped to the given id. |
| source_id | CPK, FK $\rightarrow$ `Sources.source_id | The unique source_id for each source. |
ClassificationHierarchy
| Variable | Role | Description |
|---|---|---|
| child_id | CPK, FK $\rightarrow$ Classifications.id |
The unique classification identifier for the child node. |
| parent_id | CPK, FK $\rightarrow$ Classifications.id |
The unique classification identifier for the parent node. |
Usage
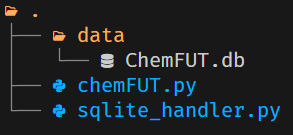
For the ChemFuncT.ChemFuncTHelper class to work by default, you must use the following directory structure:
For more information about any given method and its parameters, check the respective docstring in the source code.
Connecting to ChemFuncT.db
If using the recommended directory structure, the path to ChemFuncT.db is correctly set within the sqlite_handler.SqliteHandler.chem_func_uses_path attribute such that instantiating an instance of ChemFuncT.ChemFuncTHelper will automatically connect to the database and set the resulting sqlite3.Connection and sqlite3.Cursor objects to the ChemFuncT.ChemFuncTHelper.conn and ChemFuncT.ChemFuncTHelper.cursor attributes, respectively.
from chemFuncT import ChemFuncTHelper
FuncDB = ChemFuncTHelper()
If using a different directory structure, you can specify the path of your .db file when instantiating ChemFuncT.ChemFuncTHelper as a string or a pathlib.Path object.
FuncDB = ChemFuncTHelper("./path/to/ChemFuncT.db")
Print Database/Table Descriptions
ChemFuncTHelper.print_db_description()
This method will print a description of each table with its columns to the console.
FuncDB = ChemFuncTHelper()
FuncDB.print_db_description()
Example output:
Table: Chemicals
Column: dtxsid, Type: TEXT, Not Null: 0, Default: None, Primary Key: 1
Column: name, Type: TEXT, Not Null: 1, Default: None, Primary Key: 0
----------------------------------------
Table: Classifications
Column: id, Type: TEXT, Not Null: 0, Default: None, Primary Key: 1
Column: classification, Type: TEXT, Not Null: 1, Default: None, Primary Key: 0
Column: description, Type: TEXT, Not Null: 0, Default: None, Primary Key: 0
----------------------------------------
Table: ChemicalClassifications
Column: dtxsid, Type: TEXT, Not Null: 1, Default: None, Primary Key: 1
Column: classification_id, Type: TEXT, Not Null: 1, Default: None, Primary Key: 2
Column: source_id, Type: TEXT, Not Null: 1, Default: None, Primary Key: 3
----------------------------------------
Table: ClassificationHierarchy
Column: child_id, Type: TEXT, Not Null: 1, Default: None, Primary Key: 1
Column: parent_id, Type: TEXT, Not Null: 0, Default: None, Primary Key: 2
----------------------------------------
Table: SourceMappings
Column: id, Type: TEXT, Not Null: 1, Default: None, Primary Key: 1
Column: source_category, Type: TEXT, Not Null: 1, Default: None, Primary Key: 2
Column: source_id, Type: TEXT, Not Null: 1, Default: None, Primary Key: 3
----------------------------------------
Table: Sources
Column: source_id, Type: TEXT, Not Null: 1, Default: None, Primary Key: 1
----------------------------------------
ChemFuncTHelper.print_table()
This method will print the contents of a table from the database. The user specifies the number of rows they want printed. To print the first 10 rows of the SourceMappings table:
FuncDB = ChemFuncTHelper()
FuncDB.print_table(table_name="SourceMappings", limit=10)
To print every row of the Sources table:
FuncDB = ChemFuncTHelper()
FuncDB.print_table(table_name="Sources", limit=None)
Making Queries
ChemFuncTHelper.query_hierarchy_paths()
This method will return every possible hierarchical path starting from each root node. The returned data structure is by default a tuple of two lists of strings. The first list contains the paths with the classes encoded with their classification id (e.g., func_0004), the second list contains the paths with the class names. Each element represents one possible path (e.g., ['Drugs -> Pharmaceuticals -> Respiratory Drugs -> Anti-allergic Agents', 'Drugs -> Pharmaceuticals -> Respiratory Drugs -> Anti-allergic Agents -> Antihistamines']). Notice that the delimiter ' -> ' points from parent to child.
This method might be useful for further processing, things like determining the longest path from a root node to a leaf node.
FuncDB = ChemFuncTHelper()
id_paths, class_name_paths = FuncDB.query_hierarchy_paths()
ChemFuncTHelper.get_chem_name()
Takes a DTXSID as a required parameter and returns the chemical name.
FuncDB = ChemFuncTHelper()
dtxsid = "DTXSID9020112"
chem_name = FuncDB.get_chem_name(dtxsid)
print(chem_name)
Atrazine
ChemFuncTHelper.get_class_id_from_name()
Takes a class name and returns its class id.
FuncDB = ChemFuncTHelper()
class_name = "Pharmaceuticals"
class_id = FuncDB.get_class_id_from_name(class_name)
print(class_id)
func_0231
ChemFuncTHelper.get_class_name_from_id()
Takes a class id and returns its name.
FuncDB = ChemFuncTHelper()
class_id = "func_0231"
class_name = FuncDB.get_class_id_from_name(class_id)
print(class_name)
Pharmaceuticals
ChemFuncTHelper.get_chem_classes()
Returns the classes that a chemical (from DTXSID) is a member of. This method is fairly robust in that it allows many alterations of your output through the optional parameters.
It should be noted that the hierarchy of classes is not retained in the returned value - it is either a list of classes or a semicolon delimited string.
To return all classes that Atrazine is a member of, from all sources:
FuncDB = ChemFuncTHelper()
dtxsid = "DTXSID9020112"
atrazine_classes = FuncDB.get_chem_classes(dtxsid)
print(atrazine_classes)
Additives; Biocides; Biologicals; Fertilizers; Herbicides; Hormones; Industrial Chemicals; Pesticides; Soil Additives; Xenohormones
To only get the classes for Atrazine that came from APPRIL in a list:
FuncDB = ChemFuncTHelper()
dtxsid = "DTXSID9020112"
atrazine_classes = FuncDB.get_chem_classes(dtxsid, sources=["appril"], as_str=False)
print(atrazine_classes)
['Additives', 'Biocides', 'Fertilizers', 'Herbicides', 'Industrial Chemicals', 'Pesticides', 'Soil Additives']
To return the classification IDs instead of the class names:
FuncDB = ChemFuncTHelper()
dtxsid = "DTXSID9020112"
atrazine_classes = FuncDB.get_chem_classes(dtxsid, names=False, sources=["appril"], as_str=False)
print(atrazine_classes)
['func_0005', 'func_0087', 'func_0153', 'func_0181', 'func_0189', 'func_0227', 'func_0269']
ChemFuncTHelper.get_chem_classes_batch()
This function is a wrapper for ChemFunctHelper.get_chem_classes() that allows you to return the functional use classes for a list of DTXSIDs.
FuncDB = ChemFuncTHelper()
dtxsids = ["DTXSID9020112", "DTXSID2020006"]
batch_classes = FuncDB.get_chem_classes(dtxsids)
ChemFuncTHelper.get_class_parents()
Returns the direct parents of a given class (accepts either class name or class id).
FuncDB = ChemFuncTHelper()
class_name = "Antinematodal Agents"
parents = FuncDB.get_class_parents(class_name)
print(parents)
['Anthelmintics', 'Nematicides']
ChemFuncTHelper.get_class_children()
Returns the direct children of a given class (accepts either class name or class id).
FuncDB = ChemFuncTHelper()
class_name = "Biocides"
children = FuncDB.get_class_parents(class_name)
print(children)
['Acaricides', 'Algicides', 'Antifouling Agents', 'Antimicrobial Agents', 'Antimycotics', 'Antiparasitics', 'Avicides', 'Chemosterilants', 'Fumigants', 'Fungicides', 'Fungistats', 'Herbicides', 'Insecticides', 'Molluscicides', 'Nematicides', 'Spermicides', 'Sporicide', 'Sterilizing Agents']
ChemFuncTHelper.export_db_to_excel()
This method generates a data dump of ChemFuncT.db in the form of a .xlsx file. This requires the openpyxl library.
FuncDB = ChemFuncTHelper()
FuncDB.export_db_to_excel("./path/to/ChemFuncT_datadump.xlsx")
Installation
Option 1 - install from PyPI
pip install ChemFuncT
Option 2 - install from source
git clone https://github.com/carret1268/AMOS-ChemFuncT.git
cd AMOS-ChemFuncT
pip install -e
This uses your local source files directly, so any changes you make are reflected immediately without reinstalling.
License

Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file chemfunct-1.0.1.tar.gz.
File metadata
- Download URL: chemfunct-1.0.1.tar.gz
- Upload date:
- Size: 3.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2885f5a195277efdf0a45903787a9e2afeb3f5aecf409d07f802f4f52005b076
|
|
| MD5 |
6a5675d5dbc455ceb8b998a616db32e5
|
|
| BLAKE2b-256 |
4d743be746fc79e60672397a7787e9d3a7246ef4f35548bbd833bb6b7913c429
|
File details
Details for the file chemfunct-1.0.1-py3-none-any.whl.
File metadata
- Download URL: chemfunct-1.0.1-py3-none-any.whl
- Upload date:
- Size: 3.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ca5bceafd8bf85c00486ea6334dcd59f35eef8779f188abdeea5460b07bf078f
|
|
| MD5 |
7f29cfd8931423c0c3931351eb5ab70f
|
|
| BLAKE2b-256 |
d939de789174d6c586b2d00b294440450671dd0fa1c185b8d5a23cb12754fd23
|