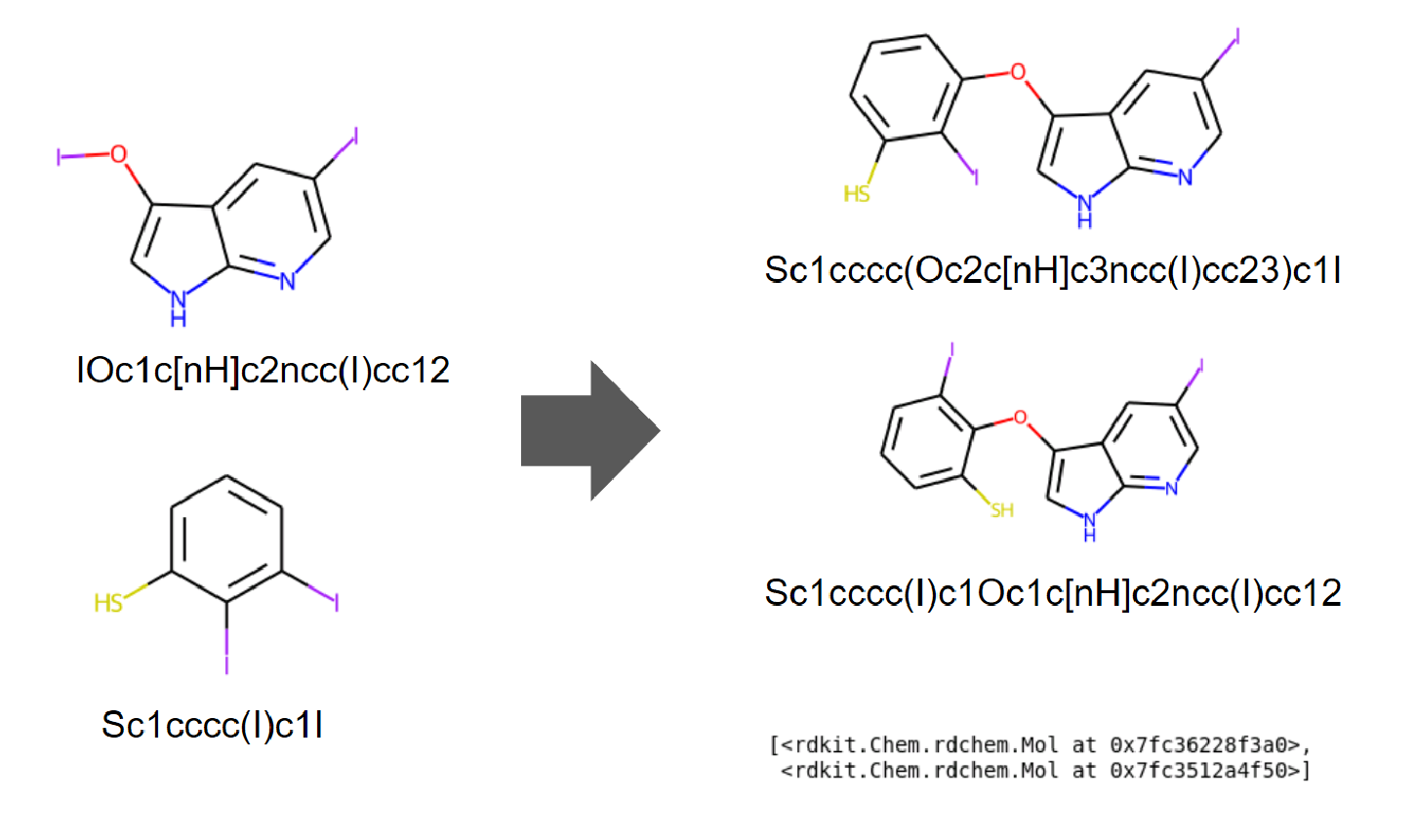
Easy to combine two molecules
Project description
Welcome to CombineMols
CombineMols is a python package for easily combining two molecules.
This intuitively aids in the combining of molecules using dummy atoms.
The easiest way to install the dependencies is to install the Anaconda 3 Python distribution and use conda to set up an environment with RDKit.
Dependencies
rdkit / mendeleev
Installation
Step 1 :
Make a new conda environment and install RDKit.
conda create -c rdkit -n my-rdkit-env rdkit
Then activate this new environment.
conda activate my-rdkit-env
Step 2 :
Inside this environment install CombineMols.
Using pip
pip install CombineMols
Examples
from rdkit import Chem
from CombineMols.CombineMols import CombineMols
Two molecule classes can be combined.
mol1 = Chem.MolFromSmiles('IOc1c[nH]c2ncc(I)cc12')
mol2 = Chem.MolFromSmiles('Sc1cccc(I)c1I')
CombineMols(mol1, mol2, "I")
Molecules can be entered directly in SMILES form.
mol1 = 'IOc1c[nH]c2ncc(I)cc12'
mol2 = 'Sc1cccc(I)c1I'
CombineMols(mol1, mol2, "I")
Dummy atom can be entered by atomic number.
mol1 = 'IOc1c[nH]c2ncc(I)cc12'
mol2 = 'Sc1cccc(I)c1I'
CombineMols(mol1, mol2, 53)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file CombineMols-1.0.1.0-py3-none-any.whl.
File metadata
- Download URL: CombineMols-1.0.1.0-py3-none-any.whl
- Upload date:
- Size: 4.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.0.1 pkginfo/1.6.1 requests/2.24.0 requests-toolbelt/0.9.1 tqdm/4.50.2 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cd7d988e82a7f7b40b19e4fb8d07a293e015c8ee91012e17da51ef70b1b64ecc
|
|
| MD5 |
22173cc2bfc1e7442632ffdc7f705619
|
|
| BLAKE2b-256 |
b56cc49bef56b81de91cfefee0495ea0c66647a4367a362eabbc191de4c117fa
|