Transform materials research data into FAIR-compliant RDF Data. Align your datasets with MDS-Onto and convert them into Linked Data, enhancing interoperability and reusability for seamless data integration. See the README or vignette for more information. This tool is used by the SDLE Research Center at Case Western Reserve University.
Project description
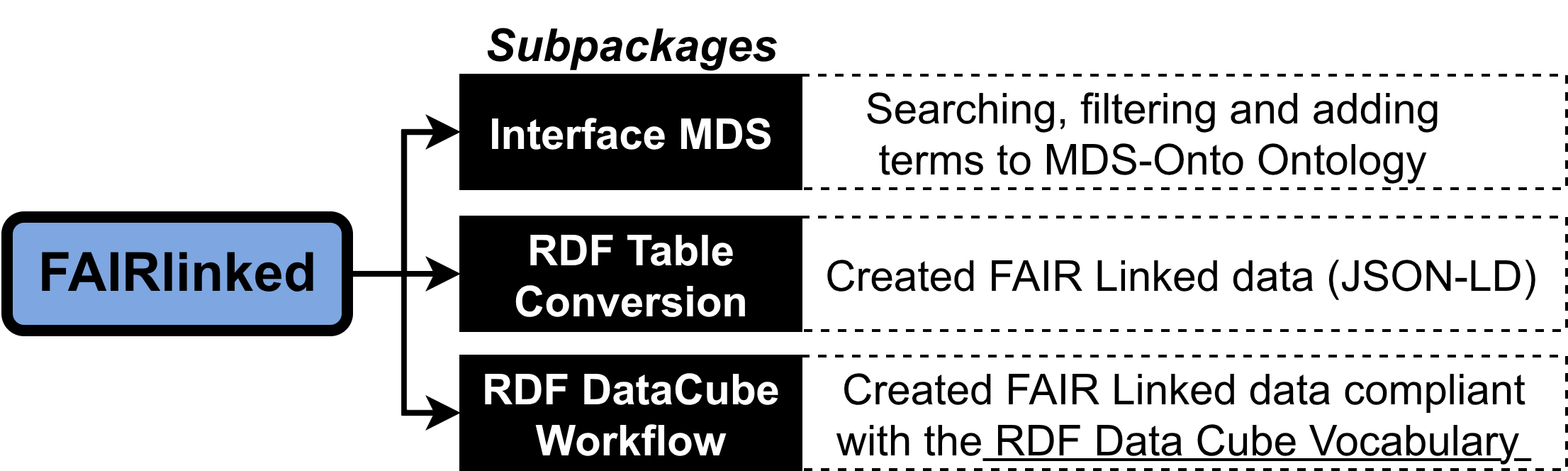
FAIRLinked
FAIRLinked is a powerful tool for transforming research data into FAIR-compliant RDF. It helps you align tabular or semi-structured datasets with the MDS-Onto ontology and convert them into Linked Data formats, enhancing interoperability, discoverability, and reuse.
With FAIRLinked, you can:
- Convert CSV/Excel/JSON into RDF, JSON-LD, or OWL
- Automatically download and track the latest MDS-Onto ontology files
- Add or search terms in your ontology files with ease
- Generate metadata summaries and RDF templates
- Prepare datasets for FAIR repository submission
This tool is actively developed and maintained by the SDLE Research Center at Case Western Reserve University and is used in multiple federally funded projects.
Documentations of how to use functions in FAIRLinked can be found here
✍️ Authors
- Van D. Tran
- Brandon Lee
- Ritika Lamba
- Henry Dirks
- Balashanmuga Priyan Rajamohan
- Gabriel Ponon
- Quynh D. Tran
- Ozan Dernek
- Yinghui Wu
- Erika I. Barcelos
- Roger H. French
- Laura S. Bruckman
🏢 Affiliation
Materials Data Science for Stockpile Stewardship Center of Excellence, Cleveland, OH 44106, USA
🐍 Python Installation
You can install FAIRLinked using pip:
pip install FAIRLinked
or directly from the FAIRLinked GitHub repository
git clone https://github.com/cwru-sdle/FAIRLinked.git
cd FAIRLinked
pip install .
👀 Public Datasets
PMMA (poly(methyl methacrylate)) dataset generated using FAIRLinked.
⏰ Quick Start
This section provides example runs of the serialization and deserialization processes. All example files can be found in the GitHub repository of FAIRLinked under resources or can be directly accessed here. Command-line version of the functions below can be found here and in updates.
Metadata management and analysis provenance with RDFTableConversion.MDS_DF
Serializing and deserializing with RDFTableConversion.MDS_DF
The main.py module contains the MatDatSciDf class, which serves as an object that contains both the data and metadata associated with that data. It bridges tabular Pandas DataFrames and Linked Data (RDF) by maintaining synchronized metadata templates and ontological mappings.
A semantic wrapper for Pandas DataFrames that enforces FAIR principles by validating researcher identity (ORCID) and ensuring ontological consistency before serialization.
Initialize the semantic wrapper. Setting metadata_rows=True instructs the class to ignore the first three rows. The method template_generator can be used to parse the metadata.
import pandas as pd
from FAIRLinked import MatDatSciDf
# Load the microindentation data
raw_df = pd.read_csv("resources/worked-example-RDFTableConversion/microindentation/sa17455_00.csv")
mds_df = MatDatSciDf(
df=raw_df,
metadata_template={},
orcid="0000-0001-2345-6789",
df_name="PMMA_Hardness_Test",
metadata_rows=True
)
template, matched, unmatched = mds_df.template_generator(skip_prompts=True)
mds_df.metadata_template = template
To serialize the data, run serialize_row or serialize_bulk method:
row_graphs = mds_df.serialize_row(
output_folder="resources/worked-example-RDFTableConversion.MDS_DF/individual_pmma_records",
format='json-ld',
row_key_cols=["Measurement", "Sample"], # Used to name files and generate IDs
license="CC0-1.0" # Defaults to public domain
)
master_graph = mds_df.serialize_bulk(
output_path="resources/worked-example-RDFTableConversion.MDS_DF/master_pmma_record.jsonld",
format='json-ld',
row_key_cols=["Measurement", "Sample"],
license="MIT"
)
To turn serialized JSON-LDs back into an MDS_DF, run from_rdf_dir:
import os
from FAIRLinked import MatDatSciDf
from FAIRLinked.InterfaceMDS.load_mds_ontology import load_mds_ontology_graph
# 1. Setup environment and load the reference ontology
mds_graph = load_mds_ontology_graph()
input_directory = "resources/worked-example-RDFTableConversion.MDS_DF/individual_pmma_records"
# 3. Use the method to reconstruct.
reconstructed_mds_df = MatDatSciDf.from_rdf_dir(
input_dir=input_directory,
orcid="0000-0001-2345-6789",
df_name="Restored_PMMA_Study",
ontology_graph=mds_graph
)
| Method | Description | Key Arguments |
|---|---|---|
template_generator |
Parses the isolated first 3 rows of the CSV to map Types, Units, and Stages to the JSON-LD template. | skip_prompts |
serialize_row |
Transforms each individual row of the DataFrame into its own RDF graph/file (e.g., .jsonld). |
output_folder, row_key_cols, license |
serialize_bulk |
Aggregates all row-level data into a single master graph file while preserving prefix context. | output_path, row_key_cols, license |
from_rdf_dir |
A factory method that builds a new MatDatSciDf object from a directory of RDF files. |
input_dir, orcid, ontology_graph |
save_mds_df |
Exports the data to CSV (with semantic headers), Parquet, or Arrow. | output_dir, metadata_in_output_df |
Metadata management
Users can update their metadata template using update_metadata (which handles an already existing entry), add_column_metadata (which adds a new entry to the template), or delete_column_metadata (deletes an entry from the metadata template)
mds_df.update_metadata(
col_name="Hardness (GPa)",
field="definition",
value="Vickers hardness value measured on PMMA sample."
)
mds_df.add_column_metadata(
col_name="YoungModulus",
rdf_type="mds:YoungsModulus",
unit="GigaPA",
definition="Elastic modulus of the polymer.",
study_stage="Result"
)
mds_df.add_column_metadata(
col_name="YoungModulus"
)
| Method | Description | Key Arguments |
|---|---|---|
update_metadata |
Updates a specific semantic field (e.g., definition, unit, or type) for an existing column entry. | col_name, field, value |
add_column_metadata |
Manually defines semantic metadata for a new column or one found during an audit. | col_name, rdf_type, unit, definition |
delete_metadata |
Removes a column's entire semantic definition from the internal RDF graph and template. | col_name |
view_metadata |
Renders the current metadata template as a table or raw JSON-LD. | format ("table" or "json") |
validate_metadata |
Performs a two-way check to ensure DataFrame columns and metadata entries are aligned. | None |
Data relations Management
micro_relations = {
"is about": [
("Hardness (GPa)", "Sample"),
("YoungModulus", "Sample")
],
"mds:measuredBy": [
("Hardness (GPa)", "Indenter_ID"),
("Load (Newton)", "Indenter_ID")
]
}
mds_df.add_relations(micro_relations)
mds_df.delete_relation("mds:measuredBy", ("Hardness (GPa)", "Indenter_ID"))
mds_df.delete_relation("is about")
| Method | Description | Key Arguments |
|---|---|---|
add_relations |
Ingests a dictionary of links (e.g., Hardness is about Sample) and validates them against the ontology. |
data_relations (dict) |
delete_relation |
Removes a specific subject-object pair or an entire property group. | prop_key, pair (optional) |
view_data_relations |
Generates a visual validation report of all current semantic links. | None |
get_relations |
Scans the active ontology to find all available OWL Object and Datatype properties. | None |
validate_data_relations |
Verifies that predicates are ontologically valid and columns exist in the DataFrame. | None |
Analysis Provenance
Batch Analysis with AnalysisGroup
The AnalysisGroup class allows you to execute a function multiple times while capturing the provenance and data metadata for every iteration. It then aggregates these into a master report and a unified JSON-LD graph.
Example: Batch Lattice Parameter Trend Analysis In this scenario, we perform a linear regression on lattice parameter data across different temperature ranges and track each fitting step as a distinct event.
import numpy as np
from sklearn.linear_model import LinearRegression
from fairlinked import AnalysisGroup
# 1. Initialize the Group
# This will manage individual trackers and aggregate their results.
group = AnalysisGroup(
proj_name="Lattice_Trend_Study",
home_path="resources/worked-example-RDFTableConversion.MDS_DF/analysis/test_research_analysis_data",
orcid="0000-0001-2345-6789"
)
# 2. Define the Analysis Function
def perform_regression(X_data, y_data):
"""
Fits a linear model to lattice data and returns regression coefficients.
The tracker will automatically audit these inputs and the dictionary output.
"""
model = LinearRegression()
X_reshaped = np.array(X_data).reshape(-1, 1)
model.fit(X_reshaped, y_data)
return {
"slope": model.coef_[0],
"intercept": model.intercept_,
"r_squared": model.score(X_reshaped, y_data)
}
# 3. Run the Batch
# We simulate a rolling window of temperature data.
temperatures = [300, 400, 500, 600]
lattice_params = [3.98, 3.99, 4.01, 4.02]
for i in range(len(temperatures)):
sub_x = temperatures[:i+2]
sub_y = lattice_params[:i+2]
# Use run_and_track to create a new tracker for this specific iteration.
group.run_and_track(perform_regression, X_data=sub_x, y_data=sub_y)
# 4. Aggregate and Export Results
# Flattens all run variables into a single tabular DataFrame.
master_df = group.create_group_arg_df()
# Creates a semantic wrapper (MatDatSciDf) for the aggregated data.
mds_obj = group.create_MatDatSciDf()
group.save_report() # Consolidates individual Markdown reports into one.
group.save_jsonld() # Generates a master graph linking all runs.
Serializing and deserializing with RDFTableConversion directly from CSV
To start serializing with FAIRLinked, we first make a template using jsonld_template_generator from FAIRLinked.RDFTableConversion.csv_to_jsonld_mapper. In your CSV, make sure to have some (possibly empty or partially filled) rows reserved for metadata about your variable.
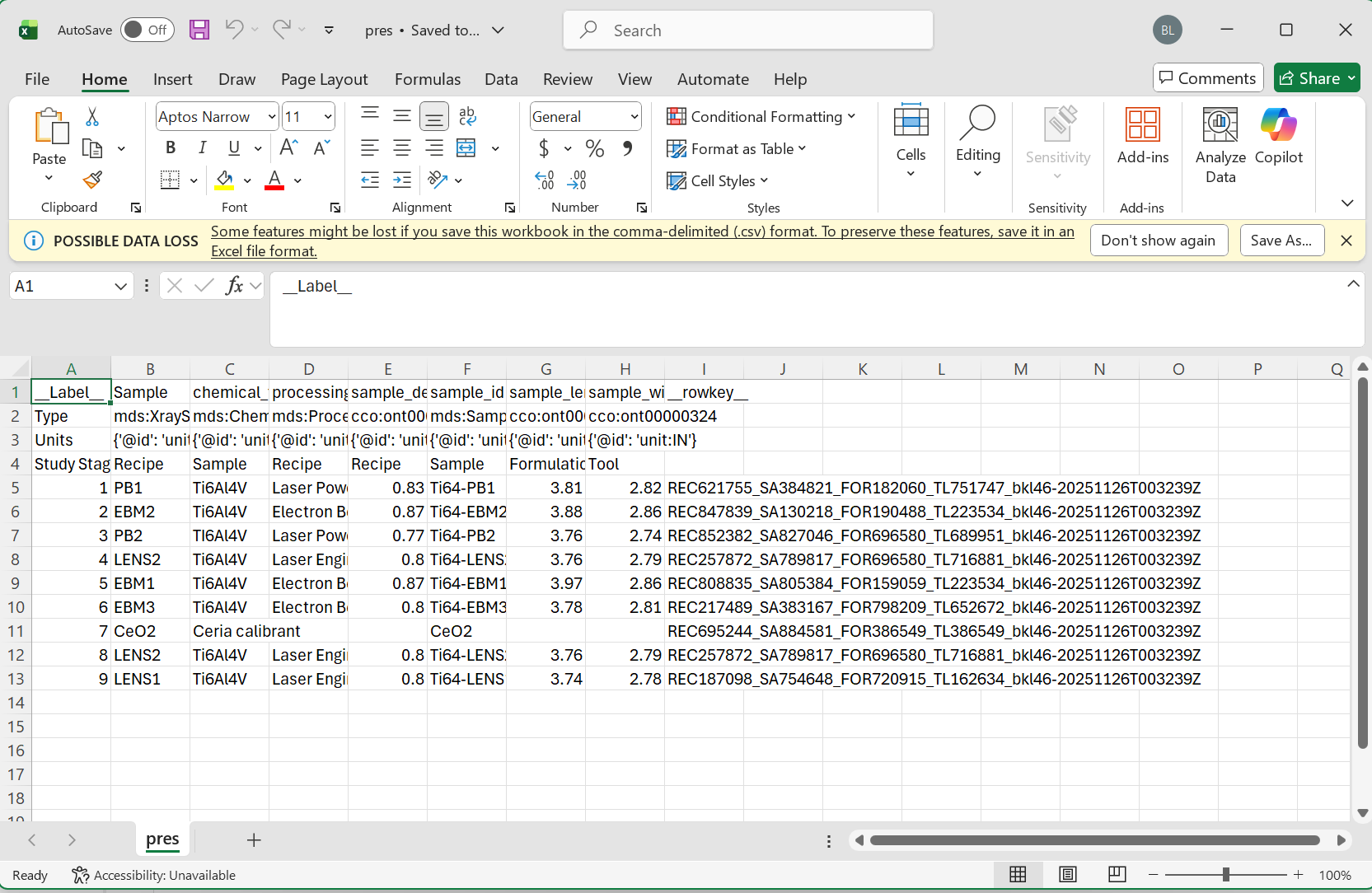
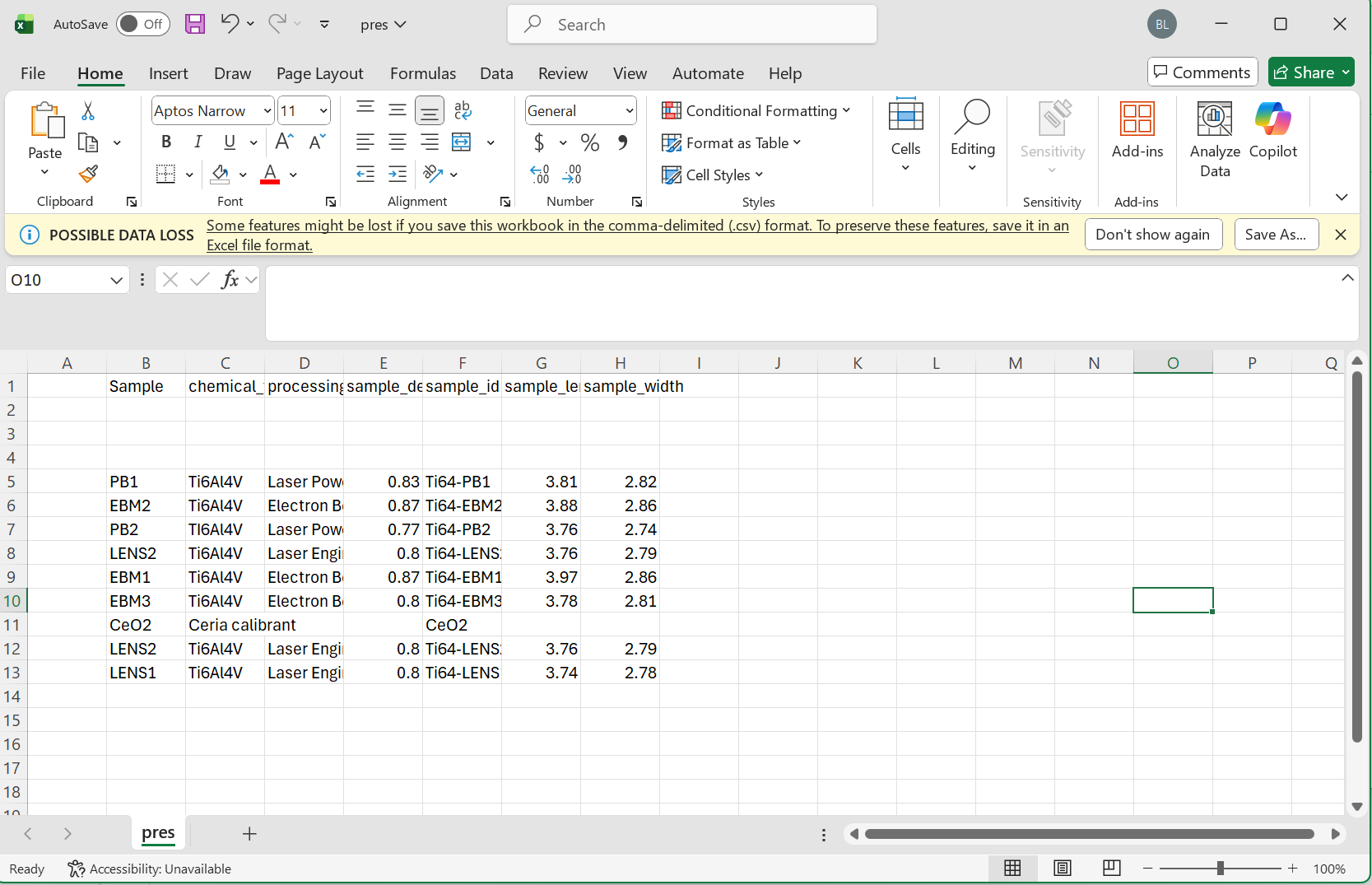
Note Please make sure to follow the proper formatting guidelines for input CSV file.
- Each column name should be the "common" or alternative name for this object
- The following three rows should be reserved for the type, units, and study stage in that order
- if values for these are not available, the space should be left blank
- data for each sample can then begin on the 5th row
Please see the following images for reference
Minimum Viable Data
During the template generating process, the user may be prompted for data for different columns. When no units are detected, the user will be prompted for the type of unit, and then given a list of valid units to choose from.
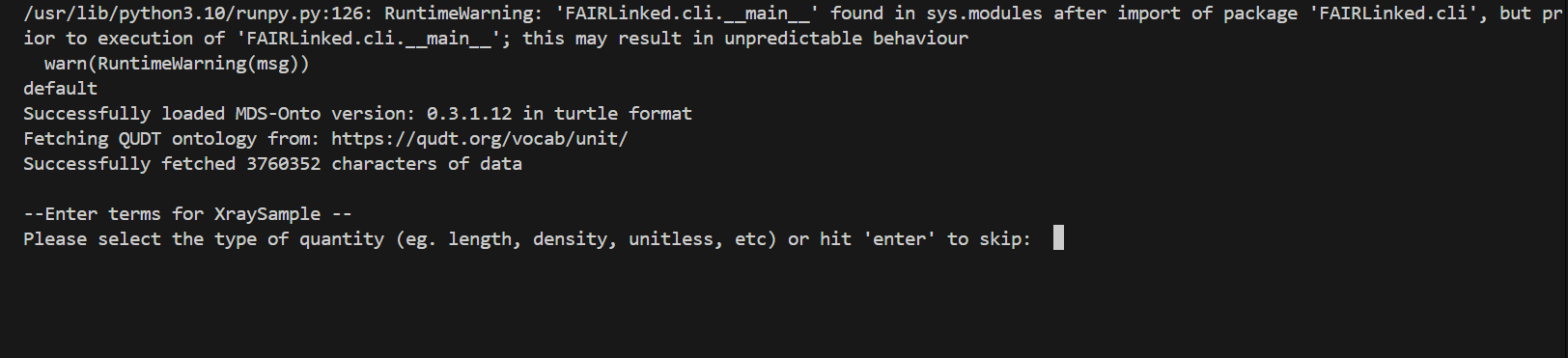
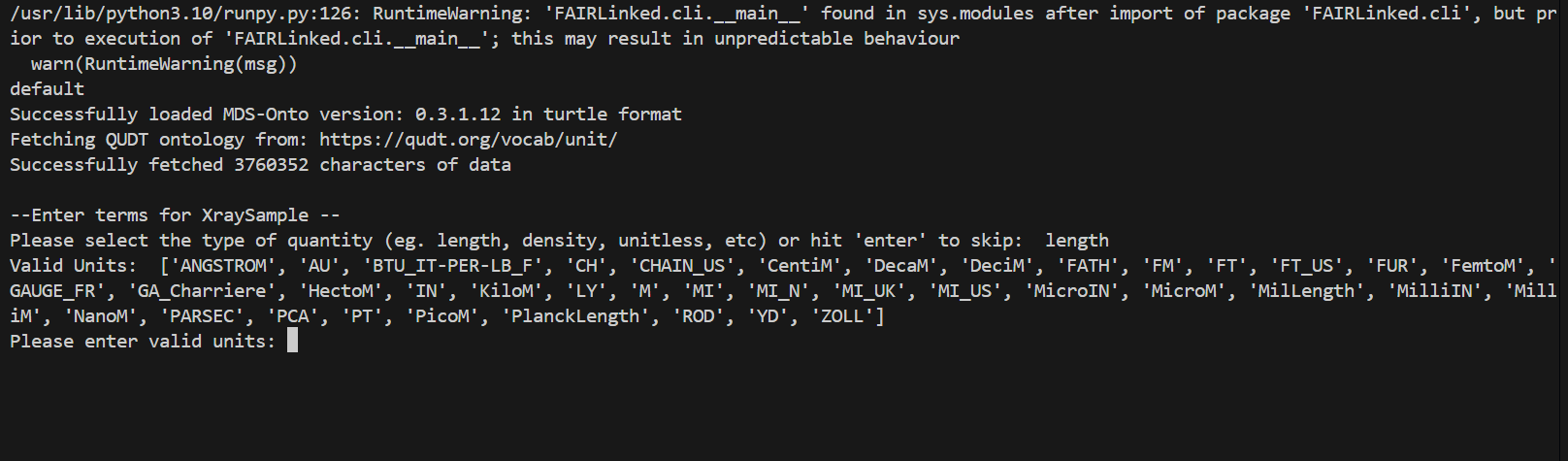

IN THIS FIRST EXAMPLE, we will use the microindentation data of a PMMA, or Poly(methyl methacrylate), sample.
from FAIRLinked.InterfaceMDS.load_mds_ontology import load_mds_ontology_graph
from FAIRLinked.RDFTableConversion.csv_to_jsonld_mapper import jsonld_template_generator
mds_graph = load_mds_ontology_graph()
jsonld_template_generator(csv_path="resources/worked-example-RDFTableConversion/microindentation/sa17455_00.csv",
ontology_graph=mds_graph,
output_path="resources/worked-example-RDFTableConversion/microindentation/output_template.json",
matched_log_path="resources/worked-example-RDFTableConversion/microindentation/microindentation_matched.txt",
unmatched_log_path="resources/worked-example-RDFTableConversion/microindentation/microindentation_unmatched.txt",
skip_prompts=False)
The template is designed to caputre the metadata associated with a variable, including units, study stage, row key, and variable definition. If the user do not wish to go through the prompts to put in the metadata, set skip_prompts to True.
After creating the template, run extract_data_from_csv using the template and CSV input to create JSON-LDs filled with data instances.
from FAIRLinked.RDFTableConversion.csv_to_jsonld_template_filler import extract_data_from_csv
import json
from FAIRLinked.InterfaceMDS.load_mds_ontology import load_mds_ontology_graph
mds_graph = load_mds_ontology_graph()
with open("resources/worked-example-RDFTableConversion/microindentation/output_template.json", "r") as f:
metadata_template = json.load(f)
prop_col_pair_dict = {"is about": [("PolymerGrade", "Sample"),
("Hardness (GPa)", "Sample"),
("VickersHardness", "YoungModulus"),
("Load (Newton)", "Measurement"),
("ExposureStep","Measurement"),
("ExposureType","Measurement"),
("MeasurementNumber","Measurement")]}
extract_data_from_csv(metadata_template=metadata_template,
csv_file="resources/worked-example-RDFTableConversion/microindentation/sa17455_00.csv",
orcid="0000-0001-2345-6789",
output_folder="resources/worked-example-RDFTableConversion/microindentation/test_data_microindentation/output_microindentation",
row_key_cols=["Measurement", "Sample"],
id_cols=["Measurement", "Sample"],
prop_column_pair_dict=prop_col_pair_dict,
ontology_graph=mds_graph,
license="CC0-1.0")
The arguments row_key_cols, id_cols, prop_column_pair_dict, and ontology_graph are all optional arguments. row_key_cols identify columns in which concatenation of values create row keys which are used to identify the columns, while id_cols are columns whose value specify identifiers of unique entities which must be kept track across multiple rows. prop_column_pair_dict is a Python dictionary specifying the object properties or data properties (specified by the value of rdfs:label of that property) which will be used in the resulting RDF graph and the instances connected by those properties. Finally, ontology_graph is a required argument if prop_column_pair_dict is provided, and this is the source of the properties available to the user. The license argument is an optional argument. If not provided, the function defaults to the permissive public domain license (CC0-1.0). For a list of available licenses see https://spdx.org/licenses/.
To view the list of properties in MDS-Onto, run the following script:
from FAIRLinked.InterfaceMDS.load_mds_ontology import load_mds_ontology_graph
from FAIRLinked.RDFTableConversion.csv_to_jsonld_template_filler import generate_prop_metadata_dict
mds_graph = load_mds_ontology_graph()
view_all_props = generate_prop_metadata_dict(mds_graph)
for key, value in view_all_props.items():
print(f"{key}: {value}")
To deserialize your data, use jsonld_directory_to_csv, which will turn a folder of JSON-LDs (with the same data schema) back into a CSV with metadata right below the column headers.
import rdflib
from rdflib import Graph
import FAIRLinked.RDFTableConversion.jsonld_batch_converter
from FAIRLinked.RDFTableConversion.jsonld_batch_converter import jsonld_directory_to_csv
jsonld_directory_to_csv(input_dir="resources/worked-example-RDFTableConversion/microindentation/test_data_microindentation/output_microindentation",
output_basename="sa17455_00_microindentation",
output_dir="resources/worked-example-RDFTableConversion/microindentation/test_data_microindentation/output_deserialize_microindentation")
Serializing and deserializing using RDF Data Cube with QBWorkflow
The RDF Data Cube Workflow is better run in bash.
$ FAIRLinked data-cube-run
This will start a series of prompts for users to serialize their data using RDF Data Cube vocabulary.
Welcome to FAIRLinked RDF Data Cube 🚀
Do you have an existing RDF data cube dataset? (yes/no): no
Answer 'yes' to deserialize your data from linked data back to tabular format. If you do not wish to deserialize, answer 'no'. After answering 'no', you will be asked whether you are currently running an experiment. To generate a data template, answer 'yes'. Otherwise, answer 'no'.
Are you running an experiment now? (yes/no): yes
Once you've answered 'yes', you will be prompted to provide two ontology files.
Do you have these ontology files (lowest-level, MDS combined)? (yes/no): yes
This question is asking if you have the following two turtle files: 'lowest-level' and 'MDS combined'. A 'lowest-level' ontology file is a turtle file that contains all the terms you want to use in your dataset, while 'MDS combined' is the general MDS-Onto which can be downloaded from our website https://cwrusdle.bitbucket.io/. If you answer 'yes', you will be prompted to provide file paths to these files. If you answer 'no', then a generic template will be created with unspecified variable name.
Enter the path to the Lowest-level MDS ontology file: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/Low-Level_Corrected.ttl
Enter the path to the Combined MDS ontology file: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/MDS_Onto_Corrected.ttl
This will generate a template in the current working directory. For the result of this run, see resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/data_template.xlsx.
To serialize your data, start the workflow again.
$ FAIRLinked data-cube-run
Answer 'no' to the first question.
Welcome to FAIRLinked RDF Data Cube 🚀
Do you have an existing RDF data cube dataset? (yes/no): no
When asked if you are running an experiment, answer 'no'.
Are you running an experiment now? (yes/no): no
For most users, the answer to the question below should be 'no'. However, if you are working with a distributed database where 'hotspotting' could be a potential problem (too many queries directed towards one node), then answering 'yes' will make sure serialized files are "salted". If you answer 'yes', each row in a single sheet will be serialized, and the file names will be "salted" with two random letters in front of the other row identifiers.
Do you have data for CRADLE ingestion? (yes/no): no
Next, you will be prompted for a namespace template (which contains a mapping of all the prefixes to the proper namespace in the serialized files), your filled-out data file, and the path to output the serialized files.
Enter ORCID iD: 0000-0001-2345-6789
Enter the path to the namespace Excel file: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/namespace_template.xlsx
Enter the path to the data Excel file: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/mock_xrd_data.xlsx
Enter the path to the output folder: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/output_serialize
The next question asks for the mode of conversion. If 'entire', then the full table will be serialized into one RDF graph. If 'row-by-row', the each row will be serialized into its own graph. In this example, we will choose row-by-row.
Do you want to convert the entire DataFrame as one dataset or row-by-row? (entire/row-by-row): row-by-row
You will then be prompted to select your row identifiers and the FAIRLinked will automatically exits.
The following columns appear to be identifiers (contain 'id' in their name):
Include column 'ExperimentId' in the row-based dataset naming? (yes/no): yes
Include column 'DetectorWidth' in the row-based dataset naming? (yes/no): no
Approved ID columns for naming: ['ExperimentId']
Conversion completed under mode='row-by-row'. Outputs in: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/output_serialize/Output_0000000123456789_20260216110630
FAIRLinked exiting
To deserialize your data, start the workflow and answer 'yes' to the first question:
Welcome to FAIRLinked RDF Data Cube 🚀
Do you have an existing RDF data cube dataset? (yes/no): yes
Enter the path to your RDF data cube file/folder (can be .ttl/.jsonld or a directory): resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/output_serialize/Output_0000000123456789_20260216110630/jsonld
Enter the path to the output folder: resources/worked-example-QBWorkflow/test_data/Final_Corrected_without_DetectorName/output_deserialize
💡 Acknowledgments
This work was supported by:
- U.S. Department of Energy’s Office of Energy Efficiency and Renewable Energy (EERE) under the Solar Energy Technologies Office (SETO) — Agreement Numbers DE-EE0009353 and DE-EE0009347
- Department of Energy (National Nuclear Security Administration) — Award Number DE-NA0004104 and Contract Number B647887
- U.S. National Science Foundation — Award Number 2133576
🤝 Contributing
We welcome new ideas and community contributions! If you use FAIRLinked in your research, please cite the project or reach out to the authors.
Let us know if you'd like to include:
- Badges (e.g., PyPI version, License, Docs)
- ORCID links or contact emails
- Example datasets or a GIF walkthrough
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file fairlinked-0.3.3.0.tar.gz.
File metadata
- Download URL: fairlinked-0.3.3.0.tar.gz
- Upload date:
- Size: 143.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
213fc973f23010f2c436b6deec36ead466e742972fc552184ff8183a680f6520
|
|
| MD5 |
ef502c4eb169f34f7feec201b6eb5937
|
|
| BLAKE2b-256 |
dbfb09ec37fdcf8b6625bea2942e4af1bcb07dd7a7d7c8aadf0f3ebe28d22ea9
|
File details
Details for the file fairlinked-0.3.3.0-py3-none-any.whl.
File metadata
- Download URL: fairlinked-0.3.3.0-py3-none-any.whl
- Upload date:
- Size: 154.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5c70e49e57b1d272bcde00e7e7590f93b141c2918bedaab771f65aa6c665678d
|
|
| MD5 |
c26dea6057a6d3842258bf62bcb3036a
|
|
| BLAKE2b-256 |
81daa12400f81374ef72d06f01bbb273a67787606e505729f00ce08545d8a807
|