Fungal Validation & Identification Pipeline
Project description
FunVIP 
FunVIP is now published please cite:
Seo CW, Yoo S, Cho Y, Kim JS, Steinegger M, Lim YW. FunVIP: Fungal Validation and Identification Pipeline based on phylogenetic analysis. J. Microbiol. 2025;63(4):e2411017.
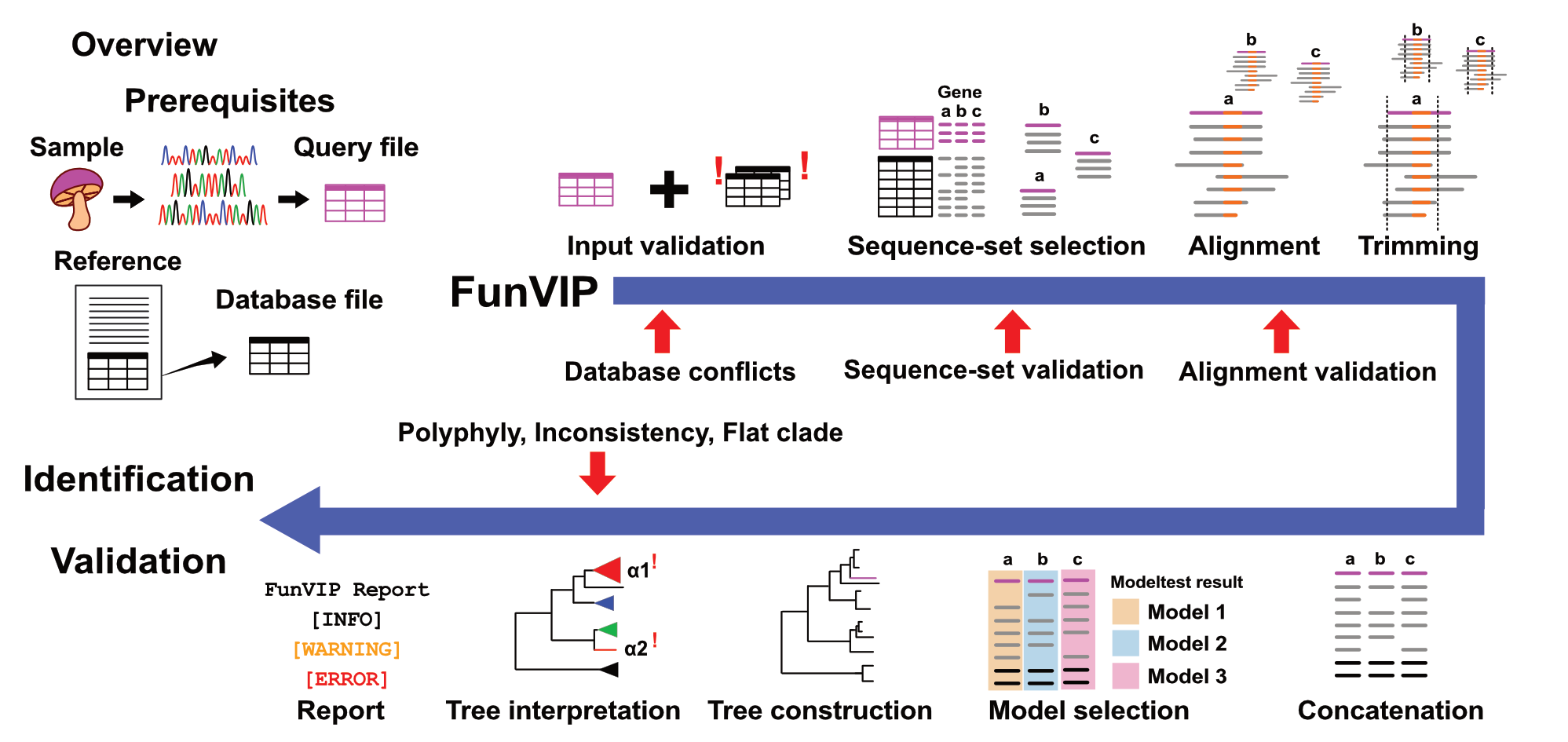
Fungal Validation & Identification Pipeline
An automatic tree-based sequence identification and validation pipeline for fungal (or maybe other) species
- Automatic tree-based identification
- Works with multiple genetic marker
- Database sequence validation algorithm implemented
Bug reports are always welcomed
IMPORTANT NOTICE: The python dependency for Linux platform has changed from 3.12 to 3.11 for TCS inclusion. Please remake conda environment for FunVIP 0.3.25 update
Tutorial
Documentation
- See Documentation for advanced usage !
Requirements
- Conda or Mamba environment
* See here for how to install conda environment
* Recently, Mamba is a lot faster than conda. See here to how to install mamba environment
Installation
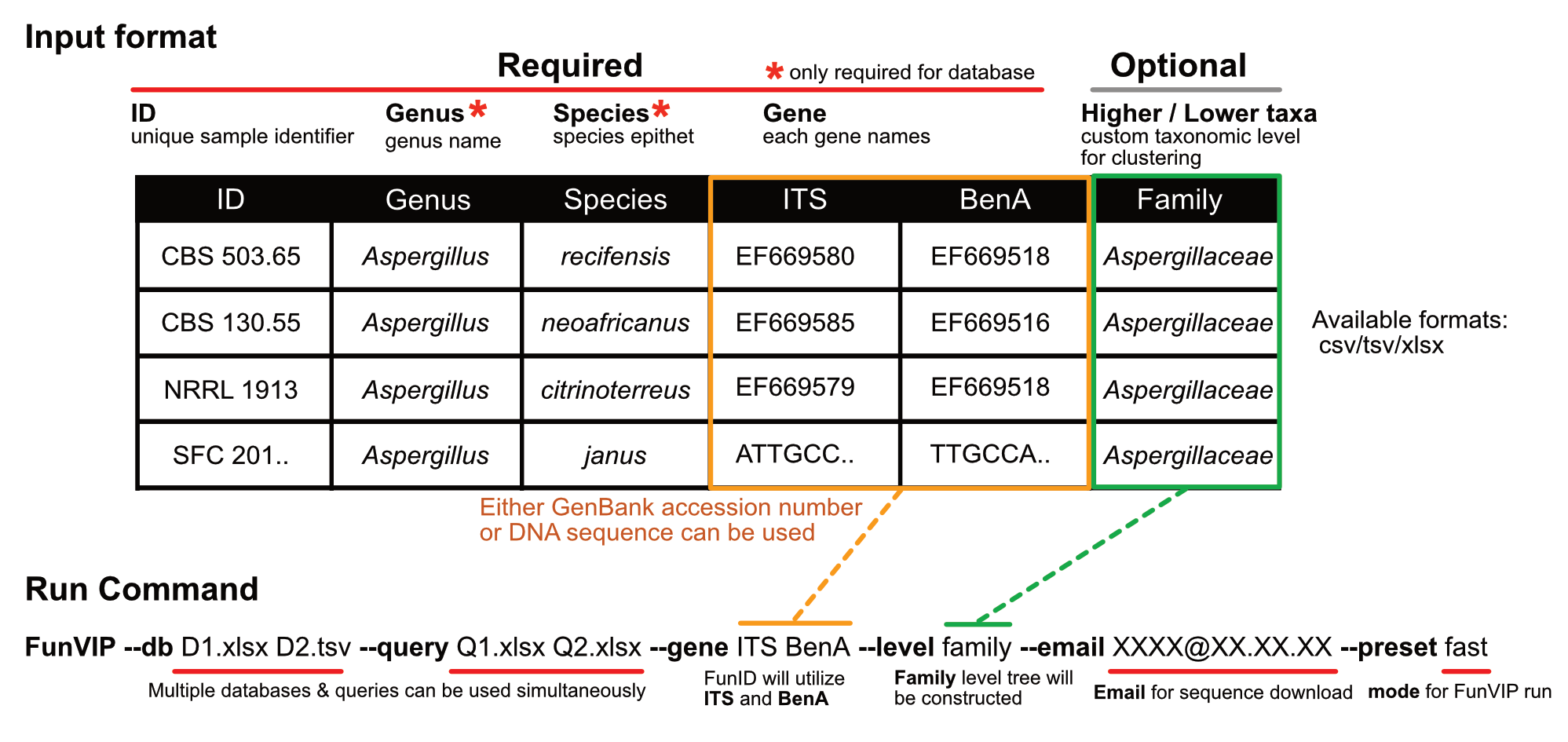
Usage
FunVIP --db {Your database file} --query {Your query file} --email {Your email} --gene {Your genes} --preset {fast or accurate}
Example
FunVIP --db Penicillium.xlsx --query Query.xlsx --email {Your email} --thread 8 --gene ITS BenA RPB2 CaM --preset fast
* See documentation for detailed usage
How to make database?
Results
-
Section Assignment.xlsx: Your clustering result is here. You can find which of your sequences are clustered to which section -
Identification_result.xlsx: Your final identification result. Shows how your sequences were assigned to species level through tree-based identification -
report.xlsx: overall statistics about the tree. If your find taxon ends with numbers, these taxon are found to be paraphyletic, so should be checked -
/Tree/{section}_{gene}.svg: Final collapsed tree in svg format. Can be edited in vector graphics programs, or in powerpoint (by ungroup) -
/Tree/{section}_{gene}_original.svg: Uncollapsed tree for inspection -
Example output tree of FunVIP
Scheduling
Beta release part 1 (2023 Feburary ~ As paper published, ver 0.3) Will be tested by our lab memebers to fix bugs and advance features- Beta release part 2 (As paper published ~ When pipeline gets stabled, ver 0.4) Will be tested by peer taxonomists
- Stable release (ver 1.0)
License
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file funvip-0.5.1.tar.gz.
File metadata
- Download URL: funvip-0.5.1.tar.gz
- Upload date:
- Size: 64.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d1c4d49ac8e57f3dadd580414e30c03ee0322adf2ccbd9617158c6b8f7bbb7a2
|
|
| MD5 |
90c9b9b46d4cfe0f4f23896c992dfbc4
|
|
| BLAKE2b-256 |
208553774bff266c5f3c5c13542ab19be6fa5a9d67fd09d148eb2617221539bc
|
File details
Details for the file funvip-0.5.1-py3-none-any.whl.
File metadata
- Download URL: funvip-0.5.1-py3-none-any.whl
- Upload date:
- Size: 64.5 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cb5689ac66ea58428b21f1da04a156a33b6aff63f5b5a7dec0ebe2fcd4cd68f7
|
|
| MD5 |
beb35fe81e803c2d8277c7168ce576dd
|
|
| BLAKE2b-256 |
d08a49f46a156d835f62c7b0116b966a3551c052f70f41cd15040b4f38913b0a
|