"This software identifies anomalous H-Alpha observations."
Project description
HAlphaAnomalyzer
(Part of the MLEcoFi Project)
HAlphaAnomalyzer is a software for detecting anomalies in H-Alpha full-disk solar observations. Leveraging a grid-based approach and advanced statistical analysis, it computes optimal pixel average range values for each grid cell to identify anomalies effectively.
Try the demo online
Table of Contents
- Installation
- Importing Required Library
- Initializing Anomalyzer
- Computing Best Ranges
- Finding Corrupt Images
- Plotting Image Likelihoods with Corrupt Cells
- Example Outputs
User Manual
Installation
To install the required library, run:
pip install HAlphaAnomalyzer
Importing Required Class
Import the Anomalyzer class from the library's anomalyzer module into your Python script with:
from HAlphaAnomalyzer.anomalyzer import Anomalyzer
Initializing Anomalyzer Object
Create an instance of Anomalyzer with a specified grid size:
anomalyzer = Anomalyzer(grid_size=8)
Computing Best Ranges
Calculate optimal range values for each grid cell using your image paths.
anomalyzer.compute_best_ranges(
non_anomalous_paths=['path/to/non-anomalous/image1.png', 'path/to/non-anomalous/image2.png'],
anomalous_paths=['path/to/anomalous/image1.png', 'path/to/anomalous/image2.png']
)
Parameters:
non_anomalous_paths: List of paths to non-anomalous image files.anomalous_paths: List of paths to anomalous image files.lower_range_end: End of candidate lower ranges, default is 20.upper_range_start: Start of candidate upper ranges, default is 80.step_size: Step size for candidate ranges, default is 2.
Finding Corrupt Images
Detect corrupt images based on computed best ranges, with options to set likelihood thresholds and minimum corrupt cells:
corrupt_images_labels = anomalyzer.find_corrupt_images(
image_paths=['path/to/image1.png', 'path/to/image2.png'],
likelihood_threshold=0.6,
min_corrupt_cells=1,
verbose=True
)
Parameters:
image_paths: List of paths to image files.likelihood_threshold: Threshold for considering a cell as corrupt, default is 0.5.min_corrupt_cells: Minimum number of corrupt cells required to classify an image as corrupt, default is 0.verbose: If True, prints the number of detected corrupt images, default is False.
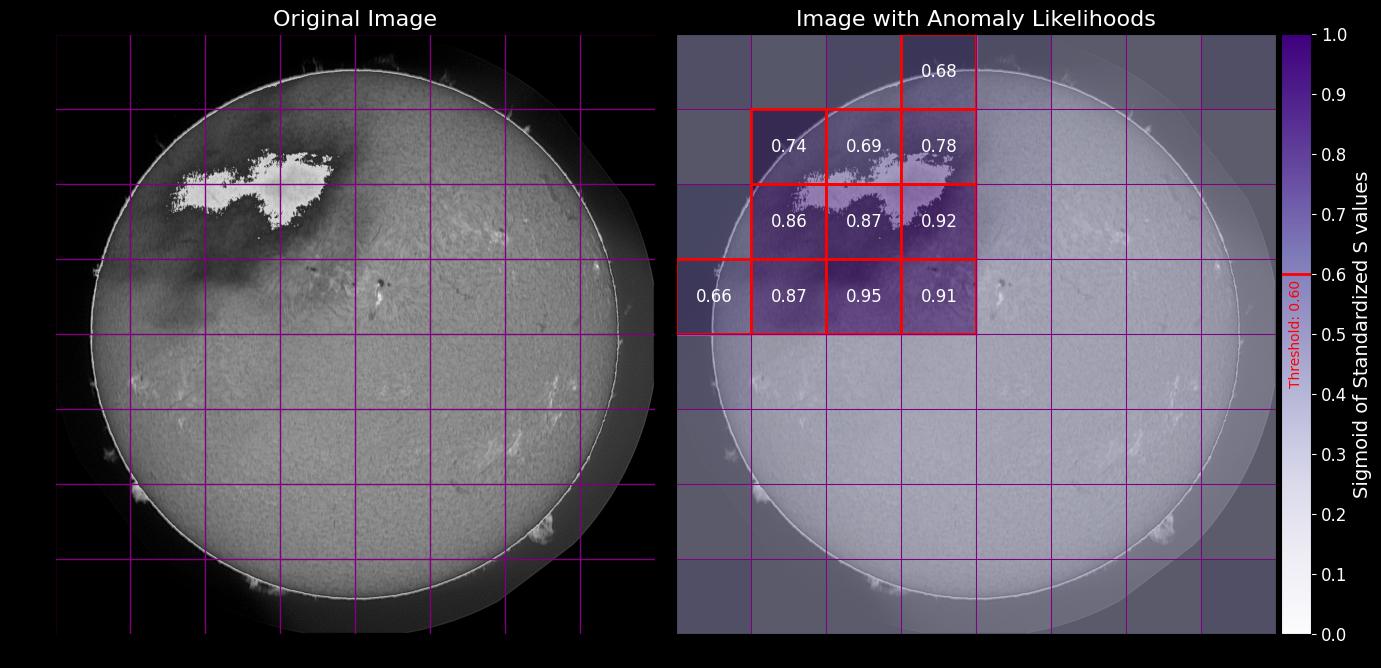
Plotting Image Likelihoods with Corrupt Cells
Visualize the anomaly likelihoods with an option to highlight corrupt cells:
anomalyzer.plot_image_likelihoods(
image_path='path/to/image.png',
likelihood_threshold=0.6
)
Parameters:
image_path: Path to the image file.likelihood_threshold: Optional threshold for identifying corrupt cells. If provided, corrupt cells will be outlined.
Example Outputs
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file HAlphaAnomalyzer-1.1.0.tar.gz.
File metadata
- Download URL: HAlphaAnomalyzer-1.1.0.tar.gz
- Upload date:
- Size: 11.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.12.1 pkginfo/1.6.0 requests/2.24.0 setuptools/50.3.2 requests-toolbelt/0.9.1 tqdm/4.50.2 CPython/3.7.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e23ec1c5c444ca1da94a2993886f4234b4c2c26015e19a90df2bbf44dc858cbf
|
|
| MD5 |
a5cd1858ece3f146f08d2eea3b1fb6b9
|
|
| BLAKE2b-256 |
f91b333a120a4aad73d01eae717ec862a67f7b5ff84f74f32fac4c9a145be1ae
|