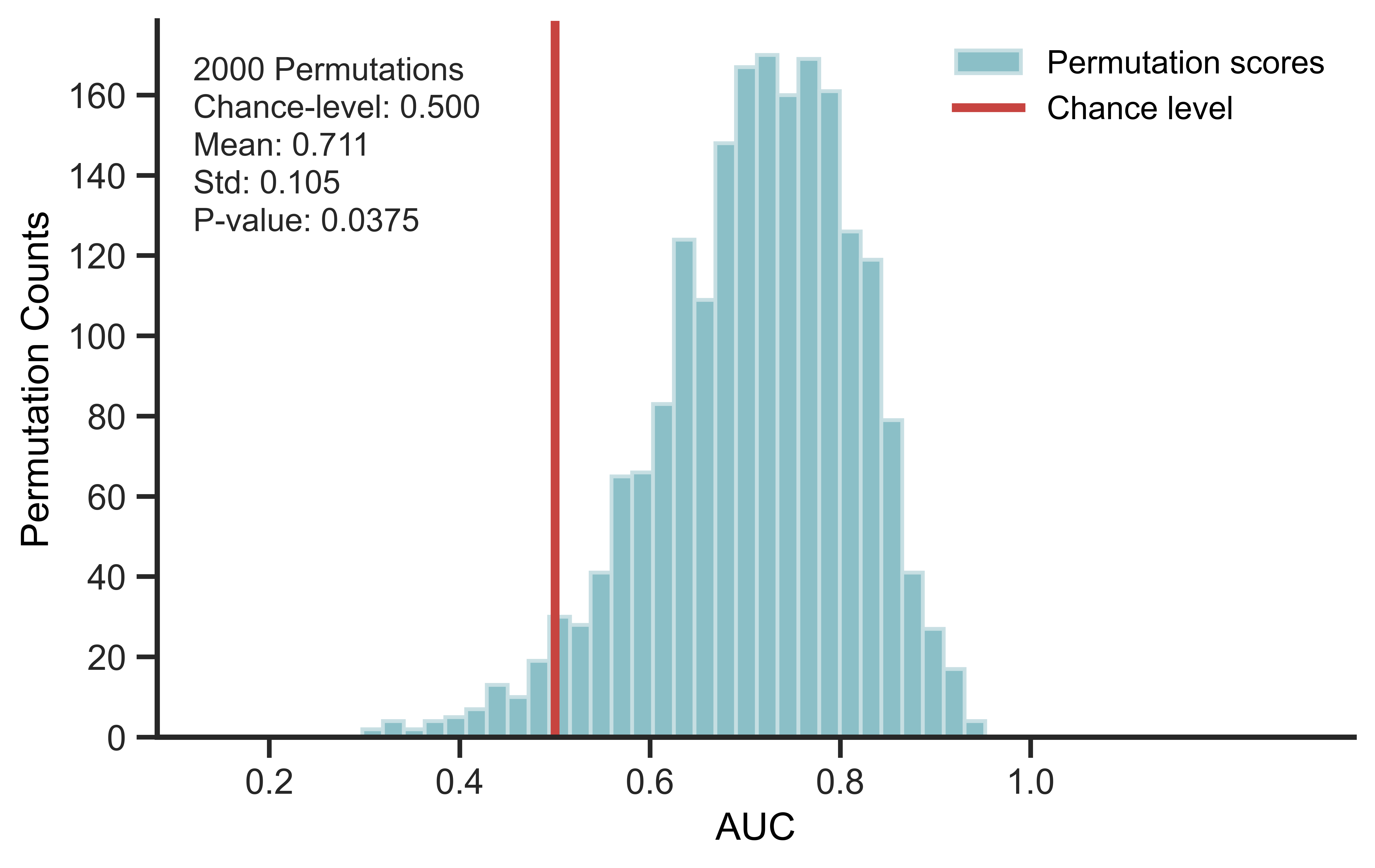
Leakage checks for machine-learning pipelines using permutation tests.
Project description

Leakly: Leakage checks for any machine-learning pipeline
Leakly uses label permutation to test whether a pipeline still performs above
chance when no true signal is present.
If it does, the pipeline may be leaking test-set information through preprocessing, feature selection, tuning, or another step.
Principle
- Permute labels to remove real signal.
- Run the full pipeline exactly as a user would run it.
- Compare the score distribution with chance level.
- Above-chance permuted performance suggests possible leakage.
Leakly includes example configurations for a leaky pipeline and a non-leaky pipeline so users can see the effect immediately.
Install
pip install Leakly
For notebook environments that need the optional notebook dependencies:
pip install "Leakly[notebook]"
For the current GitHub checkout:
git clone https://github.com/DeMONLab-BioFINDER/Leakly.git
cd Leakly
pip install -e .
Quick Start
Key Python snippet
from leakly import (
MLPipeline,
SummaryPlotter,
load_example_leakage_config,
permute_label,
)
scores = []
for seed in range(100):
permuted_y = permute_label(data.y, random_state=seed)
score = (
# user could replace with any pipeline
MLPipeline(
data.X,
permuted_y,
covariates=data.covariates,
config=load_example_leakage_config(),
).fit()).evaluate()
scores.append(score)
SummaryPlotter(scores, chance_level=0.5).plot("assets/AUC.png")
FAQ
Can Leakly check my own pipeline?
Yes. Leakly can evaluate any pipeline that takes X, y, optional covariates, and returns a test score. The key is to run the full pipeline exactly as in the real analysis, including preprocessing, feature selection, tuning, and evaluation.
Why can a leaky pipeline score well on permuted labels?
After label permutation, there should be no real biological, clinical, or statistical link between features and outcomes. A valid pipeline should therefore perform near chance.
A leaky pipeline may still score well if information from the full dataset enters the analysis before the train/test split or outside the cross-validation loop. Common sources include feature selection, scaling, imputation, covariate adjustment, dimensionality reduction, or hyperparameter tuning performed on all samples.
This is especially problematic in high-dimensional data, such as neuroimaging, omics, or biomarker studies, where random label-specific patterns can appear meaningful by chance. If the test set influences preprocessing or feature selection, the model may "remember" these random patterns and show inflated performance.
How many permutations should I run?
Use 100 for a quick check. Use 1,000 or more for publication-level evidence.
License
MIT. See LICENSE.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file leakly-0.1.0.tar.gz.
File metadata
- Download URL: leakly-0.1.0.tar.gz
- Upload date:
- Size: 255.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
847bc505ce93e997826c228c73b26cf83841e3a1ade512f90bbc50fb497c8a84
|
|
| MD5 |
b608858d56ea8e82784310e64b57ce89
|
|
| BLAKE2b-256 |
95ebf29c0d257bcc708d22e2a00c5024784462a31ecd9af2b9105f426f2cd93c
|
Provenance
The following attestation bundles were made for leakly-0.1.0.tar.gz:
Publisher:
publish.yml on DeMONLab-BioFINDER/Leakly
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
leakly-0.1.0.tar.gz -
Subject digest:
847bc505ce93e997826c228c73b26cf83841e3a1ade512f90bbc50fb497c8a84 - Sigstore transparency entry: 1526306111
- Sigstore integration time:
-
Permalink:
DeMONLab-BioFINDER/Leakly@6e224e0c2aa3c5826ca0a7979de7ca1167af3459 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/DeMONLab-BioFINDER
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@6e224e0c2aa3c5826ca0a7979de7ca1167af3459 -
Trigger Event:
release
-
Statement type:
File details
Details for the file leakly-0.1.0-py3-none-any.whl.
File metadata
- Download URL: leakly-0.1.0-py3-none-any.whl
- Upload date:
- Size: 29.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
53f3f0f88ca06de22865d94958723cd17c736ce19182ffbd2f50ce08872674a3
|
|
| MD5 |
a370c4ec74bd0e7505bd166ef93de7a2
|
|
| BLAKE2b-256 |
d976069c63ecfd0371bd9df46260c81066b7fc45b201e29f74fd803a68aeecd7
|
Provenance
The following attestation bundles were made for leakly-0.1.0-py3-none-any.whl:
Publisher:
publish.yml on DeMONLab-BioFINDER/Leakly
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
leakly-0.1.0-py3-none-any.whl -
Subject digest:
53f3f0f88ca06de22865d94958723cd17c736ce19182ffbd2f50ce08872674a3 - Sigstore transparency entry: 1526306217
- Sigstore integration time:
-
Permalink:
DeMONLab-BioFINDER/Leakly@6e224e0c2aa3c5826ca0a7979de7ca1167af3459 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/DeMONLab-BioFINDER
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@6e224e0c2aa3c5826ca0a7979de7ca1167af3459 -
Trigger Event:
release
-
Statement type: