a flexible n-ary clustering package for all applications.
Project description
MDANCE: Molecular Dynamics Analysis with N-ary Clustering Ensembles
A transformative framework for analyzing molecular dynamics simulations through advanced clustering algorithms
Installation • The Problem • Our Solution • Key Features • Quick Start • Publications • Contributing
The Problem
Molecular Dynamics (MD) simulations generate terabytes of conformational data, but extracting meaningful biological insights remains challenging. Traditional clustering methods struggle with:
- Exponential complexity - MD datasets are massive.
- Poor initialization - leading to suboptimal clustering.
- Pathway ambiguity - difficulty identifying dominant biological pathways.
- Native structure prediction - accurately identifying biologically relevant states.
- Pairwise similarity limitations - traditional methods only compare pairs of objects, causing performance bottlenecks.
- Stochastic variability - lack of reproducibility across clustering runs.
Our Solution
MDANCE introduces a novel n-ary similarity framework that transforms how we analyze MD trajectories. Our algorithms provide:
- Linear scaling - from O(N²) to O(N) complexity.
- Deterministic results - reproducible science.
- Biological relevance - algorithms designed for structural biology.
- Unprecedented accuracy - validated against experimental structures.
- Extended similarity techniques - swift identification of high and low-density regions in linear time.
Key Features
🪄 NANI - Smart k-means Initialization
Breakthrough: Deterministic centroid initialization using n-ary comparisons to identify high-density regions and select diverse initial conformations.
Key Advantages:
- Solves the seed selection challenge in k-means clustering.
- Creates compact, well-separated clusters that accurately find metastable states.
- Provides consistent cluster populations across replicates.
- Dramatically reduces runtime: clusters 1.5 million HP35 frames in ~40 minutes.
🧩 HELM - Scalable Hierarchical Clustering
Breakthrough: Combines k-means efficiency with hierarchical flexibility using n-ary difference functions.
Performance:
- Retains k-means computational efficiency while enabling arbitrary partitions.
- Successfully analyzes simulations with over 1.5 million frames.
- Achieves in ~34 minutes what traditional HAC requires 29 hours for 1.5 million frames.
- Builds hierarchy without expensive pairwise distance matrices.
🌳 DIVINE - Deterministic Divisive Clustering
Breakthrough: Top-down hierarchical clustering framework that recursively splits clusters based on n-ary similarity principles.
Key Features:
- Completely avoids O(N²) pairwise distance matrices.
- Deterministic anchor initialization with NANI.
- Multiple cluster selection criteria including weighted variance metric.
- Single-pass design enables efficient resolution exploration.
- Matches or exceeds bisecting k-means quality with reduced runtime.
🌿 mdBIRCH - Online Clustering for MD Data
Innovation: Adapts BIRCH CF-tree to molecular dynamics data with RMSD-calibrated merge tests.
Key Capabilities:
- Online clustering that processes frames as they arrive.
- Merge test calibrated directly to RMSD for physical interpretability.
- Completely avoids pairwise distance matrices.
- Scales near-linearly with number of frames.
- Two practical protocols: RMSD-anchored runs and blind sweep analysis.
- Processes hundreds of thousands of frames on a single CPU core in seconds.
🔍 SHINE - Pathway Analysis
Transformative: Hierarchical clustering that identifies dominant biological pathways from enhanced sampling data.
Key Advantages:
- Streamlines analysis of pathway ensembles from multiple MD simulations.
- Integrates n-ary similarity with cheminformatics-inspired tools.
- Identifies most representative pathway within each pathway class.
- Provides insight into dominant biomolecular transformation mechanisms.
- Lower computational cost than Fréchet distance approaches.
- Successfully applied to alanine dipeptide and adenylate kinase systems.
🎯 eQual - O(N) Clustering
Innovation: Transforms O(N²) Radial Threshold Clustering into O(N) algorithm with novel seed selection and tie-breaking.
Key Features:
- Uses k-means++ for efficient seed selection.
- Employs extended similarity indices for deterministic results.
- Eliminates memory-intensive pairwise RMSD matrices.
- Produces compact and well-separated clusters matching RTC quality.
📊 CADENCE - Density-Based Clustering
Novelty: Bridges the gap between efficient k-means and robust density-based clustering using n-ary similarity framework.
Key Advantages:
- Swiftly pinpoints high and low-density regions in linear O(N) time.
- Enables focused exploration of rare events.
- Identifies most representative conformations efficiently.
- Overcomes limitations of pairwise similarity operations.
🏆 PRIME - Native Structure Prediction
|
Game Changer: Predicts native protein structures from simulation data with unprecedented accuracy. Scientific Validation: PRIME (Protein Retrieval via Integrative Molecular Ensembles) perfectly mapped all structural motifs in benchmark studies and consistently identified native structures within 2Å RMSD of experimental data. |
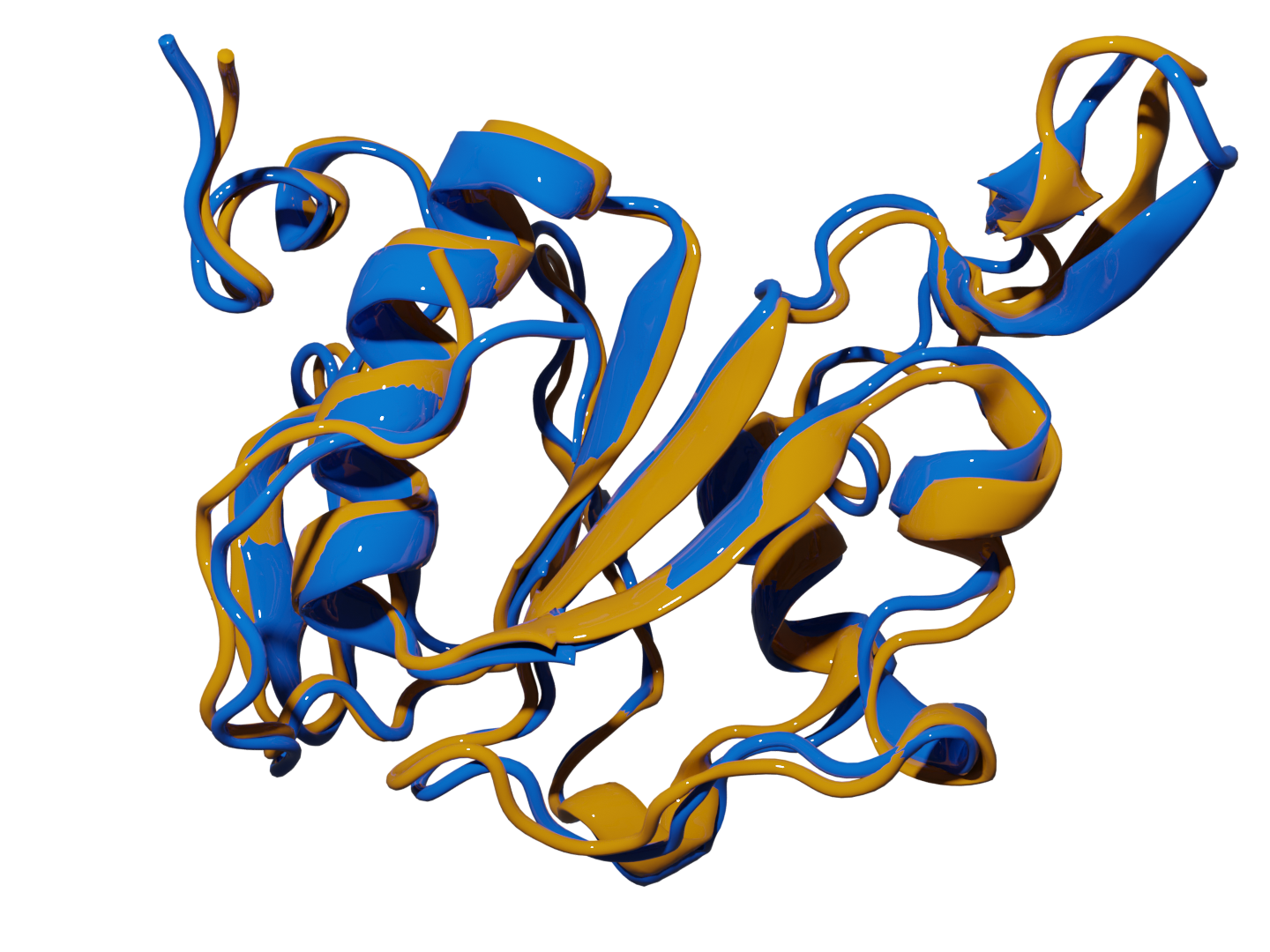
|
Algorithm Comparison
| Algorithm | Complexity | Type | Key Feature | Best Use Case |
|---|---|---|---|---|
| NANI | O(N) | Initialization | Deterministic centroids | k-means improvement |
| HELM | O(N) | Hybrid hierarchical | k-means + hierarchical fusion | Large-scale analysis |
| DIVINE | O(N) | Divisive hierarchical | Top-down splitting | Multi-resolution analysis |
| mdBIRCH | O(N) | Online clustering | Streaming data processing | Large-scale trajectories |
| SHINE | O(N) | Hierarchical | Pathway analysis | Enhanced sampling |
| eQual | O(N) | Flat clustering | Linear RTC replacement | General purpose |
| CADENCE | O(N) | Density-based | n-ary density estimation | Rare event detection |
| PRIME | O(N) | Post-processing | Native structure prediction | Structure validation |
Quick Start
Installation
pip install mdance
Basic Usage
import mdance
import numpy as np
# Load your MD trajectory data
data = np.load('trajectory.npy')
# Use NANI for optimal clustering initialization
from mdance.cluster.nani import KmeansNANI
nani = KmeansNANI(data, n_clusters=5, metric='MSD')
optimal_centroids = nani.initiate_kmeans()
# Cluster with standard *k*-means
from sklearn.cluster import KMeans
kmeans = KMeans(5, init=optimal_centroids[:5], n_init=1)
labels = kmeans.fit_predict(data)
Tutorials
- NANI Tutorial - Smart k-means initialization.
- HELM Tutorial - Scalable hierarchical clustering.
- DIVINE Scripts- Deterministic divisive clustering. 1-
run_divine.py, 2-analysis_db.ipynb, 3-assign_labels.py. - mdBIRCH Script - Online clustering for streaming MD data.
- SHINE Script - Pathway analysis.
- eQual Tutorial - Linear-time clustering.
- CADENCE Tutorial - Density-based clustering (to be added).
- PRIME Tutorial - Native structure retrieval.
Publications
Our methods are backed by peer-reviewed research:
- NANI: J. Chem. Theory Comput. 2024
- HELM: J. Chem. Inf. Model. 2025
- DIVINE: BioRxiv 2025
- mdBIRCH: BioRxiv 2025
- SHINE: J. Chem. Inf. Model. 2025
- eQual: J. Chem. Inf. Model. 2025
- CADENCE: J. Chem. Inf. Model. 2025
- PRIME: J. Chem. Theory Comput. 2024
Impact
MDANCE is enabling researchers to:
- Accelerate drug discovery by rapidly identifying biologically relevant conformations.
- Understand disease mechanisms through precise pathway analysis.
- Validate computational models against experimental structures.
- Scale analyses to massive simulation datasets.
Contributing
We welcome collaborations and contributions! Whether you're a:
- Computational biologist with novel analysis needs.
- Method developer interested in extending our framework.
- Structural biologist with challenging datasets.
Get involved:
- Open an issue for bug reports or feature requests.
- Submit a pull request for improvements.
- Reach out to discuss research collaborations.
Funding
This research was supported by the National Institute of General Medical Sciences of the National Institutes of Health under award number R35GM150620.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mdance-0.3.9.tar.gz.
File metadata
- Download URL: mdance-0.3.9.tar.gz
- Upload date:
- Size: 31.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6da04eb6af8167006973627f3491a011b83542d4469df5de334132ecb1fd198c
|
|
| MD5 |
ee4f76f674c9d08b77cbd2dd583675d3
|
|
| BLAKE2b-256 |
efb67667c66916a7d4347d943fa32928e2c22dde947ae0ef8f70db7f2b082707
|
File details
Details for the file mdance-0.3.9-py3-none-any.whl.
File metadata
- Download URL: mdance-0.3.9-py3-none-any.whl
- Upload date:
- Size: 31.1 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6861d970bed294e4bad2aa709648cab22b70fb23fc11ff8058c67c2813fe5006
|
|
| MD5 |
54b33e6dcb5470e7cf84fea150881bff
|
|
| BLAKE2b-256 |
099559d59b514d70df20d1f9e6d392bbd09e60f0ec0ef3987f900f793b9ae59c
|











