MetaXTools is a novel tool for linking peptide sequences with taxonomic and functional information in Metaproteomics.
Project description
MetaX
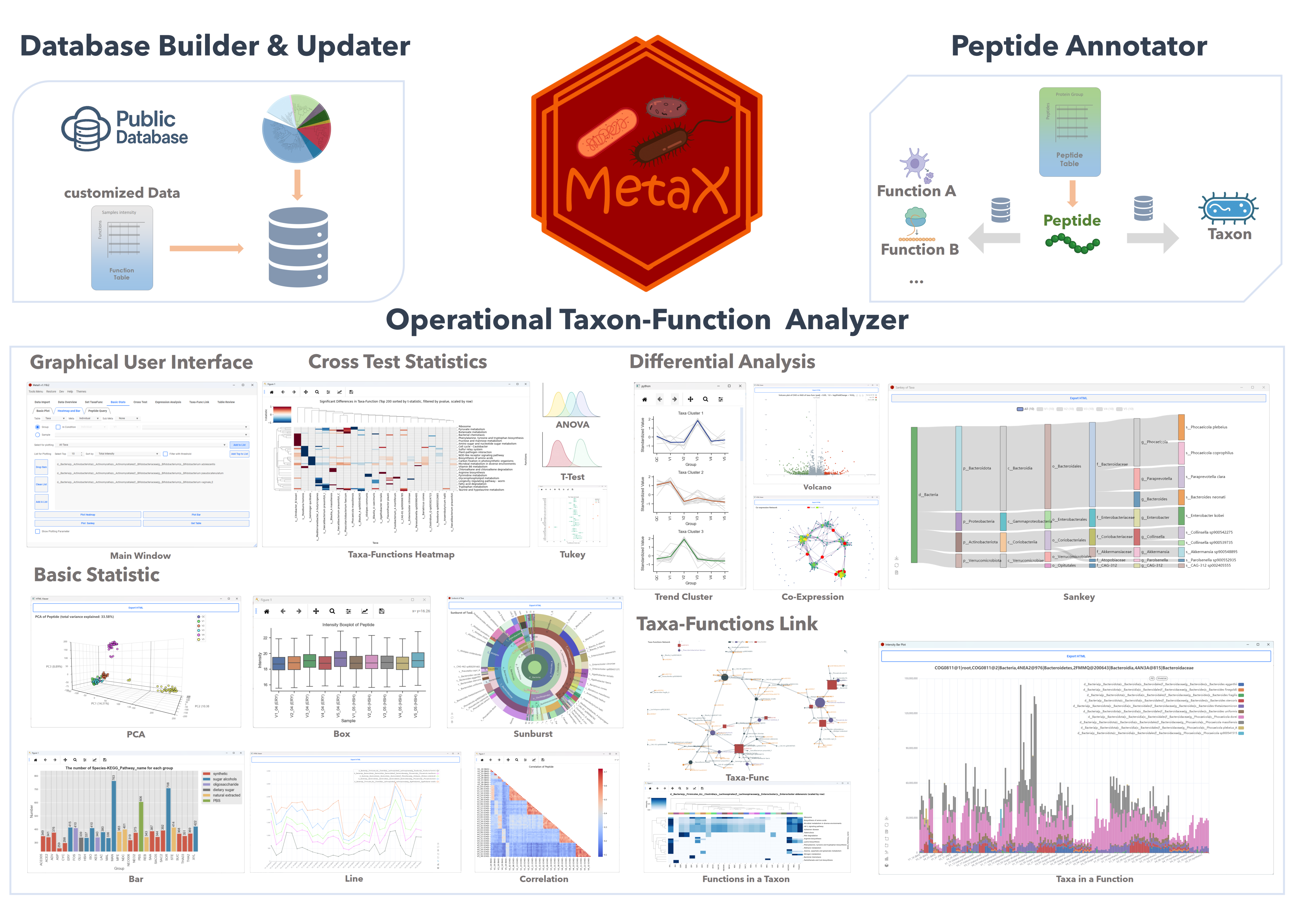
MetaX is a novel tool for linking peptide sequences with taxonomic and functional information in Metaproteomics. We introduce the Operational Taxon-Function (OTF) concept to explore microbial roles and interactions ("who is doing what and how") within ecosystems.
MetaX also features statistical modules and plotting tools for analyzing peptides, taxa, functions, proteins, and taxon-function contributions across groups.
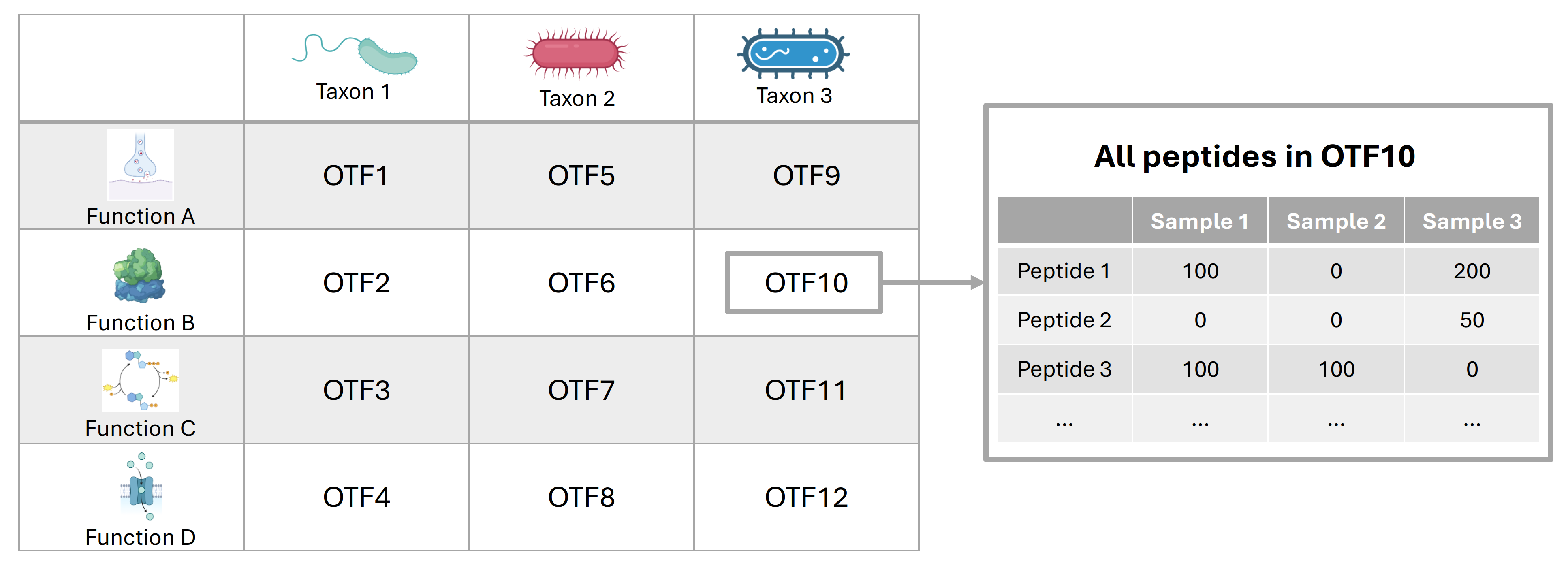
Operational Taxon-Function (OTF)
Operational Taxon-Function (OTF) Unit: An operational unit which represents the association between specific taxa and biological functions.
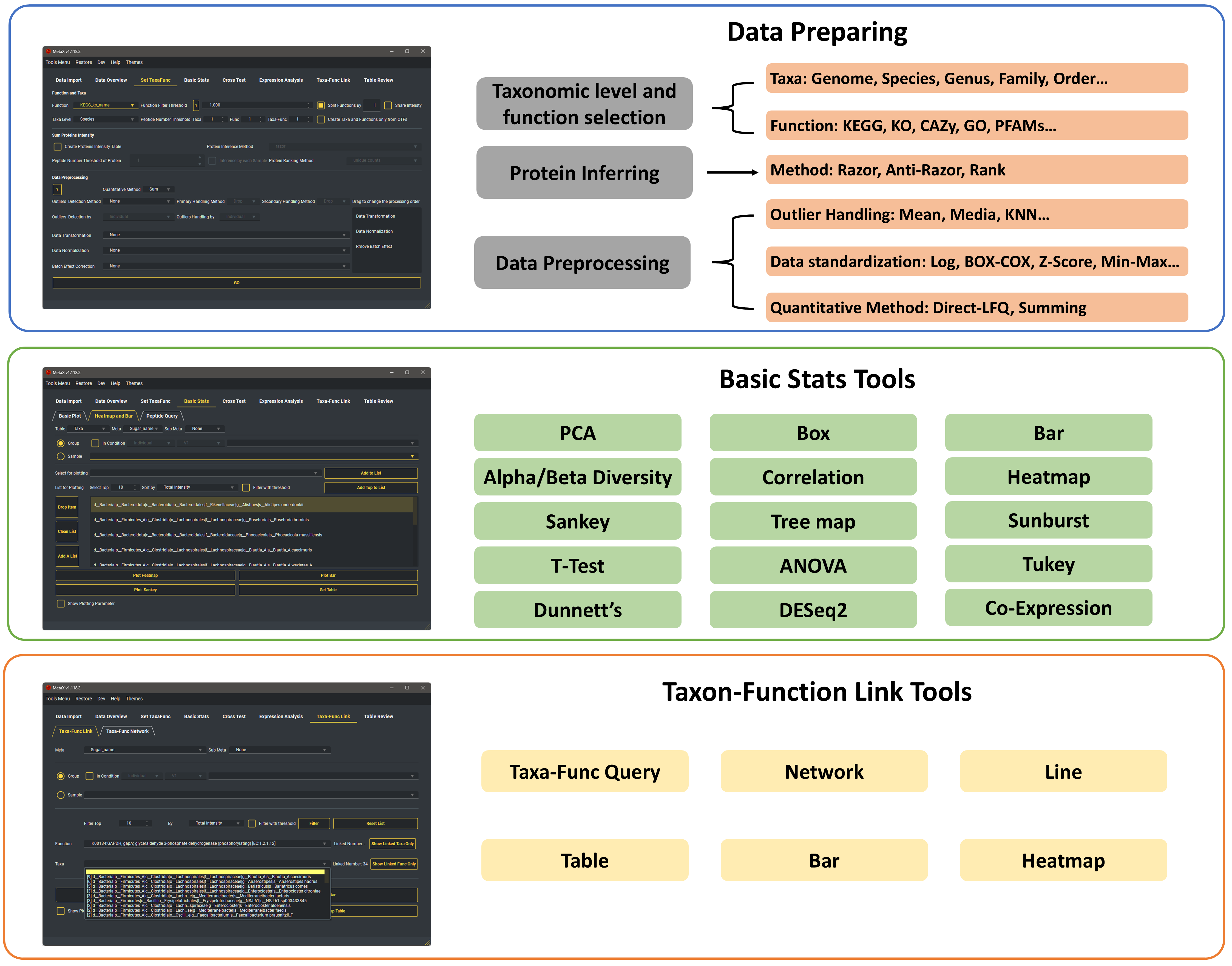
OTF Analyzer
The Tools in OTF Analyzer
OTF Examples
Linking Taxa and Functions in different levels of the hierarchy, and various functional categories. e.g., Species-KO, Genus-CAZy, Phylum-EC, etc. Lots of visualization tools can be selected.
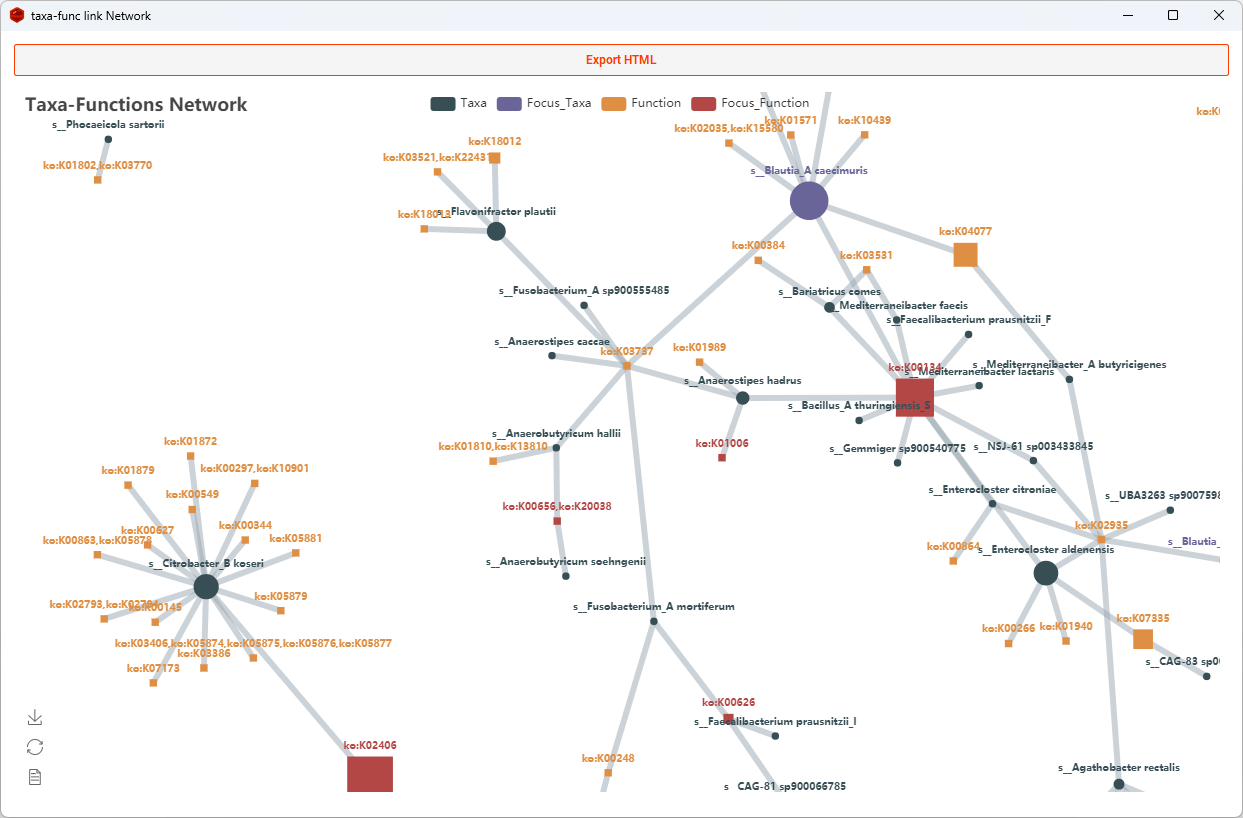
Network
Dots represent taxa and squares represent functions in the network.
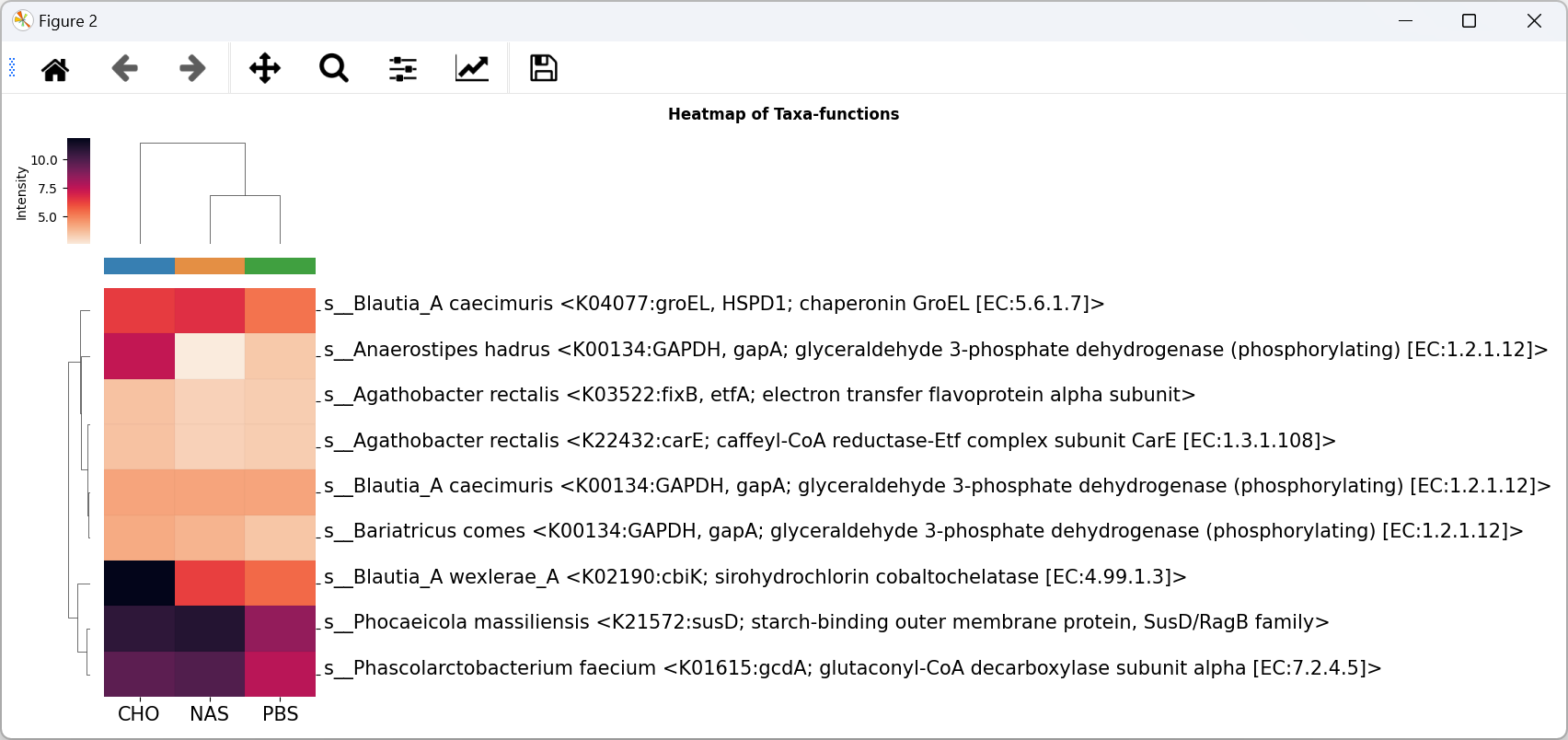
Heatmap
Show all Taxa of a function.
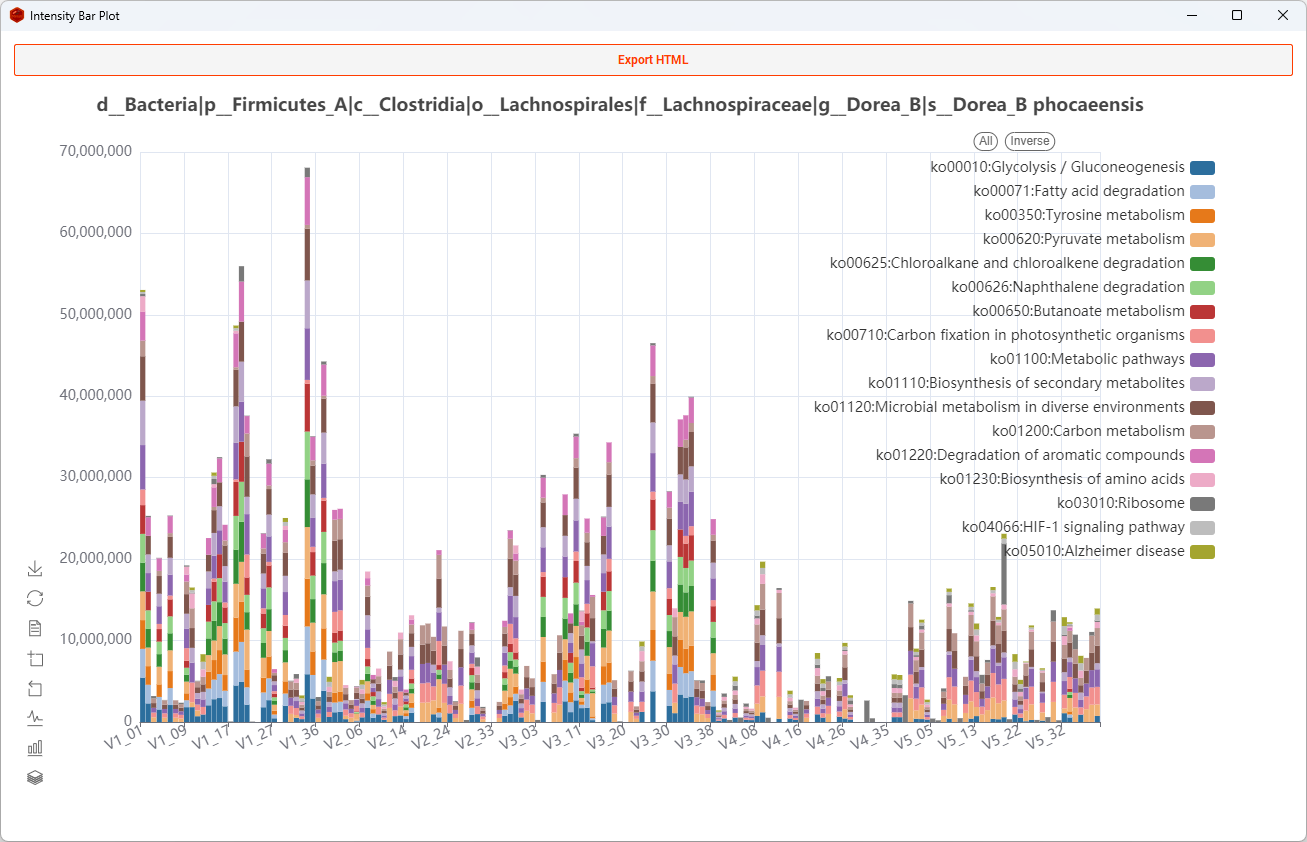
Bar
Show all functions of a taxon
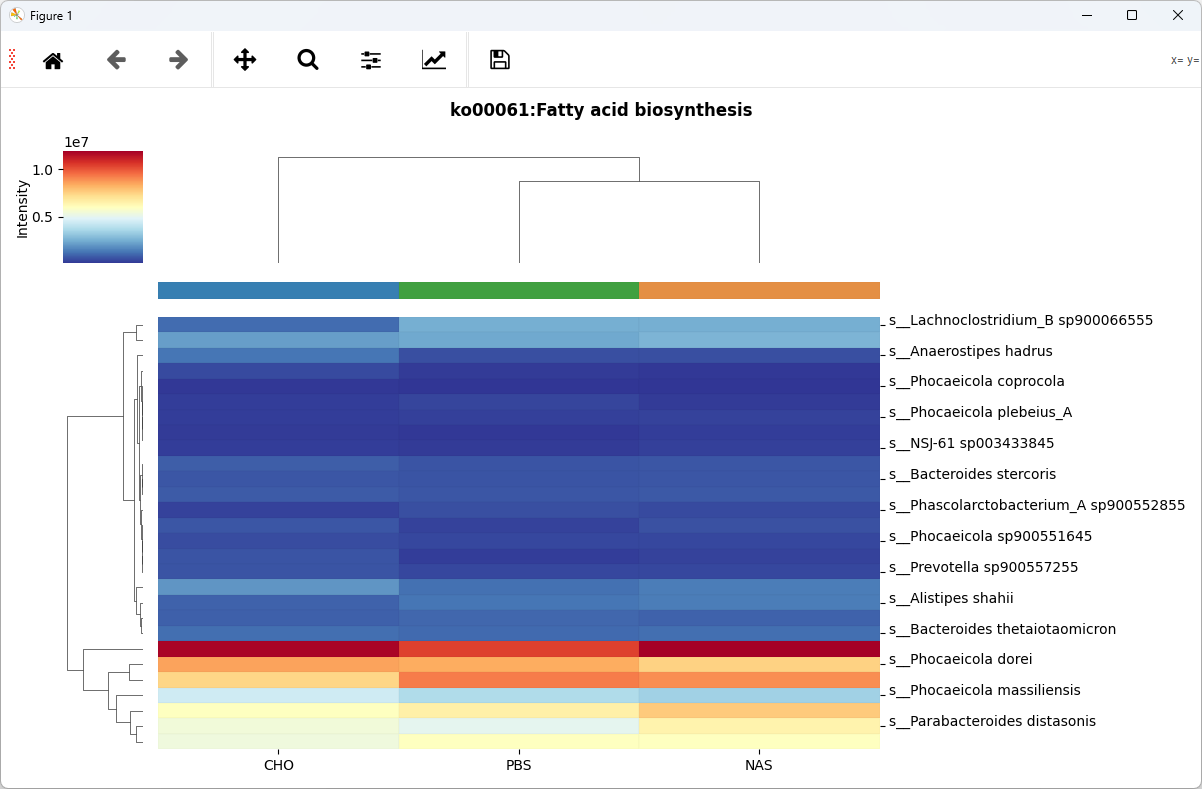
OTF Heatmap
Show OTFS intensity in groups(samples), e.g., Species-KO OTF Heatmap
Download & Installation
-
Desktop Version (Recommended)
The desktop version comes fully set up and ready to use, including all required dependencies and a graphical user interface.
-
Command-line version
Install via PyPI:
python -m pip install MetaXTools
After installation, launch the GUI by typing
metaxin your terminal.
Guidebook
Desktop Version(desktop):- Refer to the MetaX Cookbook for detailed instructions on how to use MetaX with the graphical user interface.
Command-line version:- Read the example documentation in the Notebook for detailed instructions and examples.
Citing
If you use MetaX in your research, please cite the following publication:
Operational Taxon-Function Framework in MetaX: Unveiling Taxonomic and Functional Associations in Metaproteomics. Wu, Q., Ning, Z., Zhang, A., Zhang, X., Sun, Z., & Figeys, D. (2025). Analytical Chemistry. DOI: 10.1021/acs.analchem.4c06645.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file metaxtools-1.127.1.tar.gz.
File metadata
- Download URL: metaxtools-1.127.1.tar.gz
- Upload date:
- Size: 4.6 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.10.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5c0f804e3c9a8d40e94d9d2606824933f9f30c30f27677ae659c0e4614f1d26d
|
|
| MD5 |
548040dbc4c8894bc3d61189faf53380
|
|
| BLAKE2b-256 |
5a2acdb3ece8c395a3ee0891496552cead0288857aa009aa449b801fb12069cc
|
File details
Details for the file metaxtools-1.127.1-py3-none-any.whl.
File metadata
- Download URL: metaxtools-1.127.1-py3-none-any.whl
- Upload date:
- Size: 4.7 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.10.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
debe8f2eab280e0b71f4a3206fb001f475e7de614a21f5518f31d3967a9dc367
|
|
| MD5 |
104ae98e5f011a4bad8adf7c033d091f
|
|
| BLAKE2b-256 |
602003748a0a1094c4e955fce95a6f179d984b364814dcee8f1cb6ca686e7816
|