This is a reimplementation of fundamental functionalities of 'Pydistinto', a software project developed at the [TCDH](https://tcdh.uni-trier.de/).
Project description
PyDistintoX
A Python tool for contrastive text analysis using 16 statistical distinctiveness measures (based on Pydistinto, see Project Status for details).
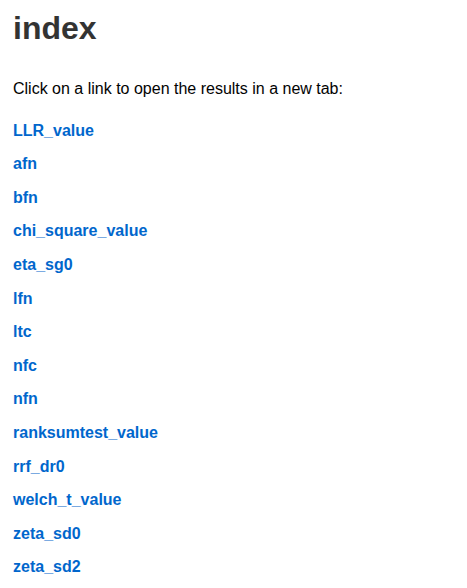
Overview |
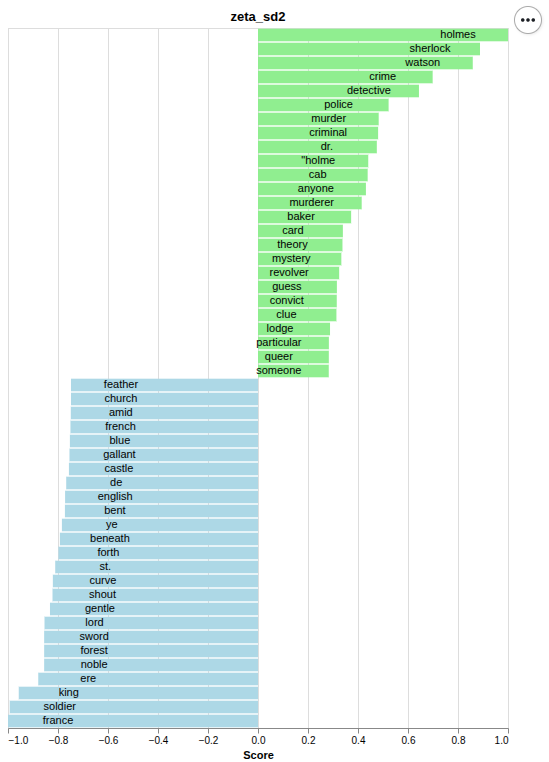
Zeta SD2 |
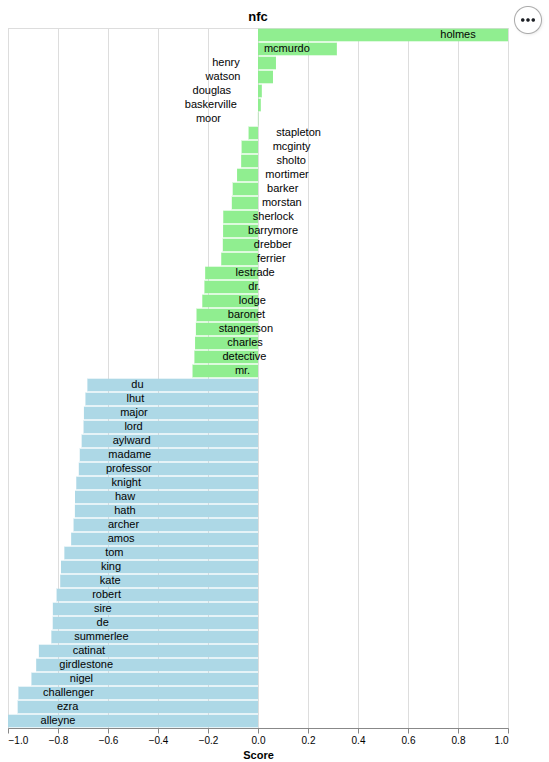
NFC |
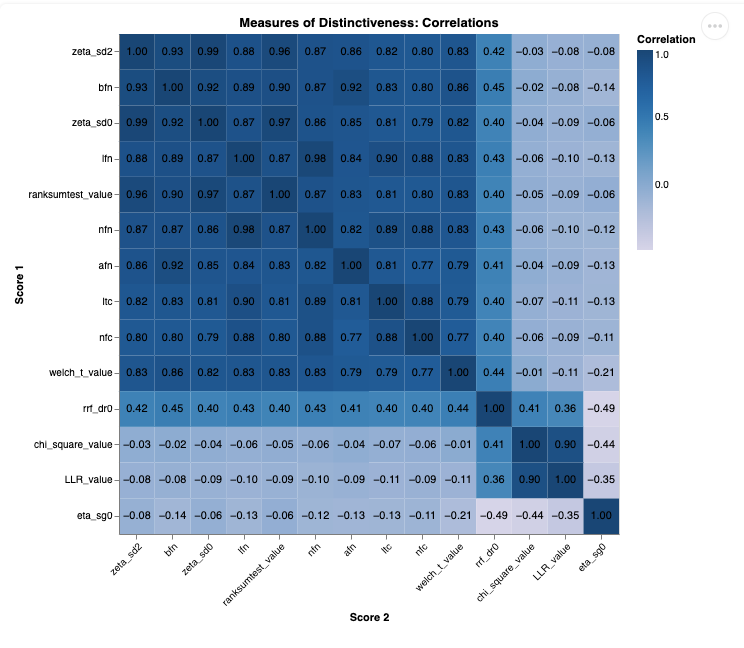
Heatmap |
||
PyDistintoX is designed to compare and analyze two text corpora using a comprehensive set of 16 statistical measures (e.g., TF-IDF, Zeta, Chi-squared, LLR, Eta). Built as a command-line interface (CLI) tool, it enables researchers, linguists, and data analysts to efficiently quantify and visualize lexical distinctiveness between corpora.
Key Features:
- 16 distinctiveness measures for robust contrastive analysis.
- CLInterface for quick, scriptable workflows.
- Python Library for custom workflows (see documentation)
- Visualization support (e.g., heatmaps) for intuitive interpretation.
- Lightweight and scalable—optimized for large corpora using Gensim and NumPy/SciPy.sparse.
Overview
PyDistintoX can be used in two ways:
- Standalone Application (CLI): Run analyses directly from the command line.
- Python library: Import functions for custom workflows.
Furthermore, we support the installation via uv as well via pip.
Prerequisites
Choose either:
| uv (recommended) | Pure Python |
|---|---|
| install here | >= 3.11.1, < 3.13 |
Note: Replace
pythonin the commands below with your specific version if needed.
The two corpora you want to compare must be stored in separate directories.
Standalone Application (CLI)
Run analyses directly from the command line.
Installation
Choose either:
uv (recommended)
uv tool install pydistintox
pip
pip install pipx
pipx install pydistintox
Setup
Download spacy model of your choice. You need to know the model name when using the CLI later as well.
Then run
pydistintox --download <MODEL_NAME>
For an english language model run for example:
pydistintox --download en_core_web_sm
An Example Script can be found here
Usage
Base Command: pydistintox [OPTIONS]. The results will be saved in the current working directory.
Command Line Options
Required
--tar path/to/target/corpusspecify the directory for target corpus. Give the directory, not a path to a file! (Escape spaces in the path if necessary)--ref path/to/reference/corpusspecify the directory for reference corpus. Give the directory, not a path to a file! (Escape spaces in the path if necessary)--model spacy_model_namespaCy model name in the format:{lang}_core_{dataset}_{size}. Example:en_core_web_sm(lang=en, dataset=web, size=sm). Available sizes: sm, md, lg, trf. Find models here.
Optional
--load-nlp path/to/json/dirskips tokenization and lemmatization, and load nlp results from path. Make surepath/to/json/dircontains the folderstarandref. If no path is specified, the defaultdata/interim/jsonis used.--raw-scoresScores will not be scaled to -1,1.--verboseor-vchanges logging mode to verbose
Example Command
pydistintox -v --model en_core_web_sm --tar ./data/example_texts/tar --ref ./data/example_texts/ref
Workflow Overview
The application processes your texts in three steps:
- NLP Processing: Texts are tokenized and lemmatized. If
--save-nlpflag is set, the results are saved as JSON in the specified directory. - Statistical Analysis: Various distinctiveness measures are calculated.
- Results Visualization: The results are automatically displayed in your default browser.
If you already created the JSON files (f.e. from a previous run), you can skip the first step by setting the --load-nlp flag (see Command Line Options above).
Python library
Import functions for custom workflows.
click to expand
Installation
-
If you are not using a virtual environment yet: Create one:
uv venvpython -m venv .venv
Activate it with:
source .venv/bin/activate(Linux/macOS).\.venv\Scripts\activate(Windows)
Then run either:
| uv (recommended) | pip | |
|---|---|---|
| Install | uv pip install pydistintox |
pip install pydistintox |
| Check Installation | You can see the place of installation via uv pip show pydistintox. |
You can see the place of installation via pip show pydistintox. |
Example Implementation:
# Assuming you do not have an active virtual environment
python3.11 -m venv .venv
source .venv/bin/activate
# and you are in the root directory of the pydistintox
# project. You may install it via
pip install pydistintox
# You can check your installation via
pip show pydistintox
Usage
from pathlib import Path
from pydistintox import *
spacy_download('en_core_web_sm')
# Caution: Keep in mind that the download command installs a Python package into your environment.
# In order for it to be found after installation, you MAY need to restart or reload your Python process so that new packages are recognized.
# Have a look at https://gitlab.com/leongluesing/PyDistintoX/#troubleshooting
config = Config(
skip_nlp = False,
save_nlp= JSON,
spacy_model_name='en_core_web_sm',
target=INPUT_TAR,
reference=INPUT_REF,
verbose = True,
source='YOUR COMMENT HERE',
scaling= True,
measures_to_calculate = ['zeta_sd2', 'afn']
)
...
For a complete example, see the script: library.py
Available Import Functions
Key functions for programmatic use (import via from pydistintox import ...).
Full documentation here
Remove PyDistintoX
| uv | pip | |
|---|---|---|
| uninstall | uv pip uninstall pydistintox |
pip uninstall pydistintox |
| clean cache | uv cache clean |
pip cache purge |
| remove model | uv pip uninstall en_core_web_sm |
pip uninstall en_core_web_sm |
You may delete the virtual environment completely. Thereby all dependencies and downloaded models will be deleted as well: First deactivate the virtual environment (deactivate) and then delete its folder (rm -r .venv or rmdir /s .venv on windows).
Troubleshooting
Installation of spaCy Model
The spaCy documentation warns against downloading and loading a model in the same script (see here):
Keep in mind that the download command installs a Python package into your environment. In order for it to be found after installation, you will need to restart or reload your Python process so that new packages are recognized.
However, in most cases it works nevertheless. If it is not working in your case, you need to preinstal the model manually with:
(uv) pip install $(spacy info <MODEL_NAME> --url)
If the installtion of a spacy model via (uv) pip install <MODEL_NAME> still fails, you may download the model by some other means and run (uv) pip install path/to/model instead.
Technical Reference
The distinctiveness measures provided are scaled to the range of [-1, 1] by default for visualization, but this scaling does not make them statistically comparable. Each measure is based on different underlying assumptions, mathematical formulations, and distributions (e.g., frequency-based vs. dispersion-based). Direct comparison of values across measures is not meaningful, unless one explicitly accounts for the differing mathematical foundations.
Non-TF-IDF Measures
Using the gensim tf-idf-model as helper function, PyDistintoX calculates absolute and binary frequencies. This is tf-idf divided by 1, i. e. disregarding document frequency, so strictly speaking tf-no-idf. Based on this data, the following distinct measures are calculated.
The following table is adapted from the foundational article on which this software is based: "Evaluation of Measures of Distinctiveness", by Keli Du, Julia Dudar, Christof Schöch (Wayback Machine). For more information see here (Wayback Machine)
| Name | Type of measure | References | Evaluated in | Implementation Key |
|---|---|---|---|---|
| TF-IDF | Term weighting | Luhn 1957; Spärck Jones 1972 | Salton and Buckley 1988 | – |
| Ratio of relative frequencies (RRF) | Frequency-based | Damerau 1993 | Gries 2010 | rrf_dr0 |
| Chi-squared test (χ²) | Frequency-based | Dunning 1993 | Lijffijt et al. 2014 | chi_square_value |
| Log-likelihood ratio test (LLR) | Frequency-based | Dunning 1993 | Egbert and Biber 2019; Paquot and Bestgen 2009; Lijffijt et al. 2014 | LLR_value |
| Welch’s t-test (Welch) | Distribution-based | Welch 1947 | Paquot and Bestgen 2009; Lijffijt et al. 2014 | welch_t_value |
| Wilcoxon rank sum test (Wilcoxon) | Dispersion-based | Wilcoxon 1945; Mann and Whitney 1947 | Paquot and Bestgen 2009; Lijffijt et al. 2014 | ranksumtest_value |
| Burrows Zeta (Zeta_orig) | Dispersion-based | Burrows 2007; Craig and Kinney 2009 | Schöch 2018 | zeta_sd0 |
| logarithmic Zeta (Zeta_log) | Dispersion-based | Schöch 2018 | Schöch 2018; Du et al. 2021 | zeta_sd2 |
| Eta | Dispersion-based | Du et al. 2021 | Du et al. 2021 | eta_sg0 |
TF-IDF Measures
PyDistintoX uses the gensim tf-idf model for calculating distinctness scores. The following parameters are used:
- 'nfn',
- 'nfc',
- 'bfn',
- 'afn',
- 'lfn',
- 'ltc'
For more information on the meaning of these parameters see wikipedia Wayback Machine, or the following paragraph:
Explanation from the Gensim Documentation
From the gensim documentation:
smartirs (str, optional) –
SMART (System for the Mechanical Analysis and Retrieval of Text) Information Retrieval System, a mnemonic scheme for denoting tf-idf weighting variants in the vector space model. The mnemonic for representing a combination of weights takes the form XYZ, for example ‘ntc’, ‘bpn’ and so on, where the letters represents the term weighting of the document vector.
Term frequency weighing:
- b - binary,
- t or n - raw,
- a - augmented,
- l - logarithm,
- d - double logarithm,
- L - log average.
Document frequency weighting:
- x or n - none,
- f - idf,
- t - zero-corrected idf,
- p - probabilistic idf.
Document normalization:
- x or n - none,
- c - cosine,
- u - pivoted unique,
- b - pivoted character length.
Default is ‘nfc’. For more information visit SMART Information Retrieval System.
SpaCy and Gensim
While gensim features its own rule-based tokenizer, we provide the opportunity to plug-in spaCy models for tokenization and lemmatization. Results of this process are saved as json files in spaCy's own format. We then extract lemmas from these data and recast them as Gensim input documents (basically list of words forming sentences which in turn are combined in a list to form a document).
Security
If you discover a security vulnerability in PyDistintoX, please report it to leongluesing@posteo.de with details (version, reproduction steps, impact) and allow 48–72 hours for a response before public disclosure.
Project Status
PyDistintoX was developed by Leon Glüsing and Stefan Heßbrüggen-Walter. Original development was funded by the Deutsche Forschungsgemeinschaft via the project "[Prodatphil: Science and Logic(https://www.uni-muenster.de/Wissenschaftstheorie/en/Forschung/PRODATPHIL/index.html)]", project no. 537184692. Since Leon Glüsing’s (my) contract has ended in March 2026, the project is now voluntarily maintained and further developed by Leon Glüsing. While no major updates are currently planned, the project’s future trajectory could evolve with increased community involvement or active contributions. So feel free to get involved!
This is a reimplementation of fundamental functionalities of "Pydistinto", a software project developed at the TCDH. The original script can be found here.
The following major changes are worth mentioning:
-
using Gensim and NumPy instead of Pandas for the construction of term-document matrices; Gensim can parse corpora that do not fit into RAM, NumPy/SciPy is used to process sparse matrices with lower memory requirements,
-
using the inbuilt tf-idf functions of Gensim for adding more tf-idf related measures of distinctiveness (see below),
This is a lite version of the original Pydistinto since we offer less features, f.e. no randomization of texts. However, if your corpus reaches a size that makes it difficult for the original Pydistinto to cope, it may be worth to try out our solution. Furthermore, the CLI Version of PyDistintoX is easier to use than the original Pydistinto.
Contribute
This project thrives on community contributions. Please feel free to submit pull requests or open issues! You will find more detailed information for developers here
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pydistintox-0.2.3.tar.gz.
File metadata
- Download URL: pydistintox-0.2.3.tar.gz
- Upload date:
- Size: 28.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.5.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9a536fec66c8f0a4f5c535c07f9482c85154b0082b2ddddbcb72e215179355a0
|
|
| MD5 |
0d2466bf94fc22fff69607a3b333b92d
|
|
| BLAKE2b-256 |
75a1ce3db73c1b754c34dd902dbabb1837d61d755ebdc1770ec8934acad049f0
|
File details
Details for the file pydistintox-0.2.3-py3-none-any.whl.
File metadata
- Download URL: pydistintox-0.2.3-py3-none-any.whl
- Upload date:
- Size: 38.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.5.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a3c6492a783df28ede2436e42b6f325b6169c6fea18100c3c3d13ae856e82e6c
|
|
| MD5 |
c2ae0b30796012b32b8130ef44c30ab8
|
|
| BLAKE2b-256 |
153706ee284ae2572a27de95a09fadc05a86e07b0a605f1ba0239f40ce969ea1
|











