Mutational spectra analysis package
Project description
PyMutSpec
Python library for mutational spectra analysis
Installation
pip3 install pymutspec
Example code
from Bio import SeqIO
from pymutspec.annotation import calculate_mutspec, CodonAnnotation
from pymutspec.draw import plot_mutspec12, plot_mutspec192
coda = CodonAnnotation(gencode=2) # mitochondrial genetic code
path_to_observed_mutations = ...
path_to_reference_seq = ...
# load data (mutations and sequence)
gene = SeqIO.parse(path_to_reference_seq, format='fasta')
observed_mutations = pd.read_csv(path_to_observed_mutations, sep='\t')
for col in ['Mut', 'MutType']:
assert col in observed_mutations.columns
# sample only syn mutations
mut_syn = observed_mutations[observed_mutations.MutType >= 1] # 0 for all mutations, 1 for syn, 2 for fourfold syn (syn4f)
# derive expected mutations from reference gene
sbs12_freqs, sbs192_freqs = coda.collect_exp_mut_freqs(gene, labels['all', 'syn', 'syn4f'])
sbs12_freqs_syn = sbs12_freqs['syn']
sbs192_freqs_syn = sbs192_freqs['syn']
# calculate mutation spectra
spectra12 = calculate_mutspec(mut_syn, sbs12_freqs_syn, use_context=False)
spectra192 = calculate_mutspec(mut_syn, sbs192_freqs_syn, use_context=True)
# plot mutation spectra
plot_mutspec12(spectra12)
plot_mutspec192(spectra192)
Example spectra barplots
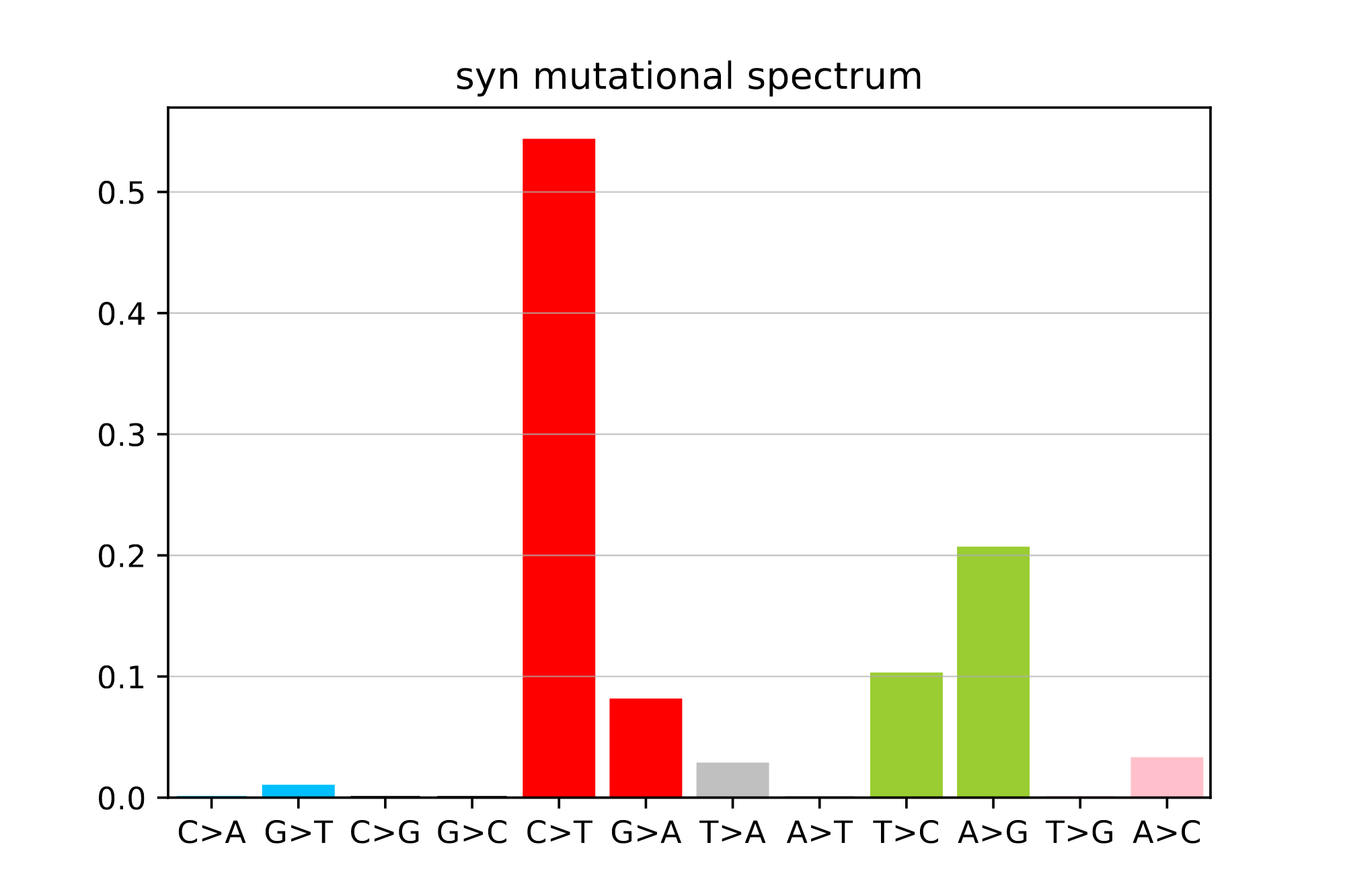
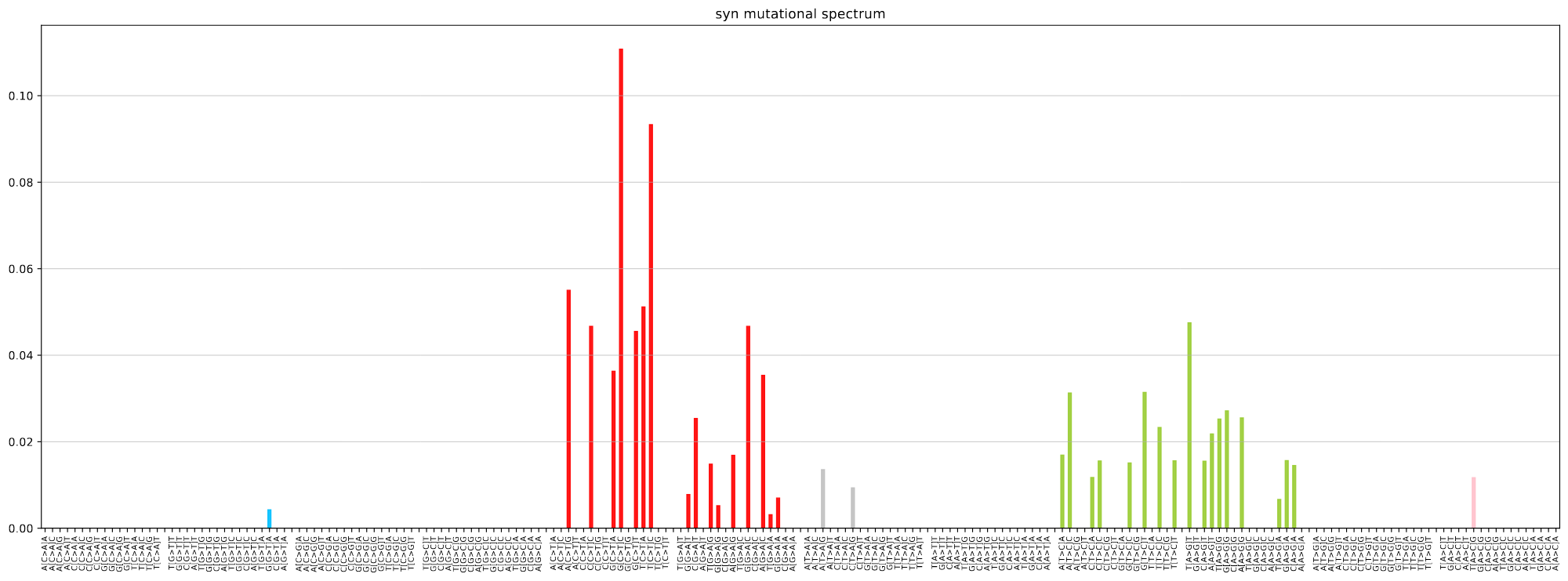
Links
- IQ-Tree2 - efficient software for phylogenomic inference
- Genetic codes
How to cite?
Bogdan Efimenko, Konstantin Popadin, Konstantin Gunbin, NeMu: a comprehensive pipeline for accurate reconstruction of neutral mutation spectra from evolutionary data, Nucleic Acids Research, Volume 52, Issue W1, 5 July 2024, Pages W108–W115, https://doi.org/10.1093/nar/gkae438
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
pymutspec-0.0.14.tar.gz
(44.3 kB
view details)
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pymutspec-0.0.14.tar.gz.
File metadata
- Download URL: pymutspec-0.0.14.tar.gz
- Upload date:
- Size: 44.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5c115e96d957fff77e56bc7659d9e66042dc91f4f45b6976ad42075d38e65043
|
|
| MD5 |
c10e44acc1f3d33dbdd651e443cbd229
|
|
| BLAKE2b-256 |
a34e8b99295daa44438d7b089a42771f6e710f4a05d2407b9b268bd38d55ac8b
|
File details
Details for the file pymutspec-0.0.14-py3-none-any.whl.
File metadata
- Download URL: pymutspec-0.0.14-py3-none-any.whl
- Upload date:
- Size: 94.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ba331144d9d8c315016d1ed4a6a93194cbe255a9570bf7d1368f3ede5cebd9e2
|
|
| MD5 |
53b164ef5c78899bbfa90103f9d636c7
|
|
| BLAKE2b-256 |
57da670b0e0cd4cd40eed833216ba54e37424c8ea81eb3b642cda82e54052a5e
|

















