File system providing reproducibility through provenance and automation
Project description
RepeatFS
RepeatFS: a file system providing reproducibility through provenance and automation
RepeatFS is a Python, FUSE-based file system with the goal of promoting scientific informatics reproducibility by recording all file and IO operations during analysis. This provenance record can then be exported and used to replicate the analysis on other systems. During replication, RepeatFS will verify that all software versions, command line parameters, and other relevant attributes match, and will report out any deviation from the original record. While other provenance software often involves learning scripting languages or migrating your workflow to a confined platform, RepeatFS operates invisibily at the file system level, and is compatible with virtually all Linux command-line software.
In addition to replication and verification, RepeatFS also provides provenance visualization. It is capable of generating a webpage visualizing the complete provenance history, including all programs that wrote to that file, all files read by those programs, all programs that wrote to those programs, etc.
Lastly, RepeatFS provides Virtual Dynamic Files (VDFs). These VDFs automatically execute commonly performed tasks (such as converting file types) in a systematic and uniform fashion. Each supported file will have a corresponding VDF presented on disk. Upon accessing this file, RepeatFS will run the appropriate task, and populate the file on the fly with the correct information. These files are then cached in memory, so subsequently accessing them does not require the process to run a second time. VDFs may be chained together for combining operations, and may be copied to turn them into normal files.
INSTALLATION
Dependencies
- Python 3 (and pip)
- FUSE 2 (and libfuse)
- GraphViz
Dependencies are available in all popular system package managers. Python 3, Libfuse, and GraphViz are also available within Anaconda (you may be able to find a channel with FUSE available as well):
- Debian/Ubuntu:
sudo apt install python3 python3-pip fuse2 libfuse2 graphviz libgraphviz-dev - RHEL/Fedora:
sudo yum install python3 python3-pip fuse2 fuse2-libs graphviz graphviz-devel - Anaconda:
conda install -c conda-forge python=3 pip libfuse=2 graphviz
After the dependencies have been installed, RepeatFS can be installed from PyPI using pip:
pip3 install repeatfs
If installing using a normal user account, pip will likely install RepeatFS into your ~/.local/bin directory. If this directory is in your PATH variable, you can simply run repeatfs. If not, you'll need to add this directory to your PATH variable, or run RepeatFS using the full path ~/.local/bin/repeatfs.
USAGE
RepeatFS functions as a transparent layer between you and your files, recording all IO activity. In order to use RepeatFS, you'll mount the target directory you want to monitor (which includes subdirectories and files). Then, anytime you wish to access any files within the monitored directory, you'll instead use the path to the RepeatFS mount.
Mount and monitor a directory:
repeatfs mount <directory to monitor> <RepeatFS mount directory>
Stop monitoring a directory:
fusermount -u <RepeatFS mount directory>
The most powerful feature of RepeatFS is the ability to record provenance and replicate the creation of the file on a different system. To ensure all operations are successfully recorded, be sure to perform the entirety of your analysis using a RepeatFS mount.
Path to a file's provenance record - this is a VDF, and is populated automatically when accessed, and may be copied to any location. Note the plus sign next to the file name below - all VDFs are available using a plus sign next to the filename:
<RepeatFS mount directory>/<any sub directories>/<file name>+/<file name>.provenance.json
Replicate a file (replication destination must be within an active RepeatFS mount:
repeatfs replicate -r <replication destination> <provenance file>
Path to a file's provenance graph - like the provenance record, this is also a VDF. RepeatFS visualizes provenance by generating an HTML file that can be vieweed in any browser:
<RepeatFS mount directory>/<any sub directories>/<file name>+/<file name>.provenance.html
REPLICATION EXAMPLES
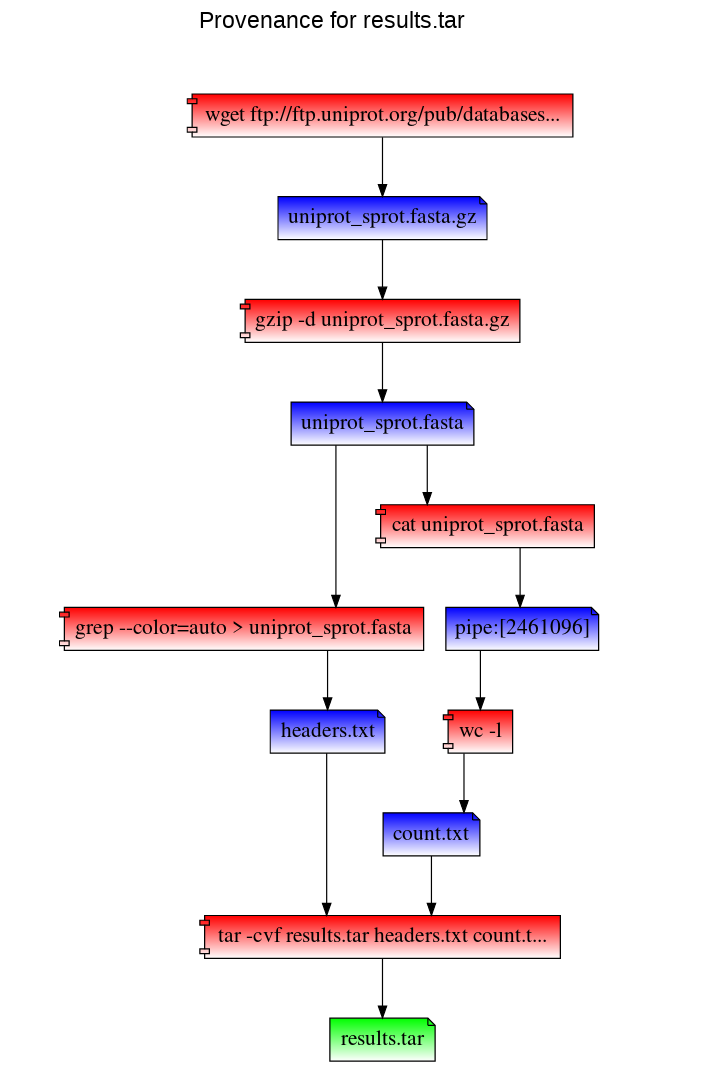
In this example, we first mount our work directory using RepeatFS. Then we download a copy of the UniProt SwissProt database, decompress it, and perform two simple tasks: extract fasta header sequences, and count the number of lines in the fasta file. We take both results, and add it to a new archive.
repeatfs mount ~/work ~/mnt
cd ~/mnt
wget ftp://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/complete/uniprot_sprot.fasta.gz
gzip -d uniprot_sprot.fasta.gz
grep ">" uniprot_sprot.fasta > ~/mnt/headers.txt
cat uniprot_sprot.fasta | wc -l > ~/mnt/count.txt
tar -cvf results.tar headers.txt count.txt
To visualize the provenance of the results.tar file within our browser:
cd results.txt+
firefox results.txt.provenance.html
RepeatFS can also replicate these steps to recreate results.tar using the results.tar.provenance.json file. You can use this file (or distribute it to others) to reproduce your work. In the following example, we've copied the provenance record into our home directory. We then mount a directory with RepeatFS and replicate the work (and save stdout and stderr into log files):
repeatfs mount ~/replicate ~/mnt
cd ~/mnt
repeatfs replicate ~/results.txt.provenance.json --stdout stdout.log --stderr stderr.log
RepeatFS will execute and verify each step. Version mismatches or other errors will be reported:
[info] Starting replication
[info] Replication complete
[info] Starting verification
[ok] Process 16056 (wget ftp://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/complete/uniprot_sprot.fasta.gz) executed
[ok] Process 16091 (gzip -d uniprot_sprot.fasta.gz) executed
[ok] Process 16107 (grep --color=auto > uniprot_sprot.fasta) executed
[ok] Process 16110 (cat uniprot_sprot.fasta) executed
[ok] Process 16113 (wc -l) executed
[ok] Process 16118 (tar -cvf results.tar headers.txt count.txt) executed
[info] Verification complete
RepeatFS can also simply list the commands that would be used during replication (using the -l argument):
repeatfs replicate ~/results.txt.provenance.json -l
This will list each command that will be run, in order. It will also list ID(s) next to each command, which can be used during replication (using the -e argument) to reconstruct a missing shell script.
[turing|1591668563.97|1652] wget ftp://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/complete/uniprot_sprot.fasta.gz
[turing|1591668614.24|1695] gzip -d uniprot_sprot.fasta.gz
[turing|1591668694.23|1756] grep --color=auto > uniprot_sprot.fasta > /tmp/mnt/headers.txt
[turing|1591669111.77|2165, turing|1591669111.77|2166] cat uniprot_sprot.fasta | wc -l > /tmp/mnt/count.txt
[turing|1591669123.5|2175] tar -cvf results.tar headers.txt count.txt
VIRTUAL DYNAMIC FILES
Provenance and replication are only a part of RepeatFS's capabilities. RepeatFS can also automate commonly performed tasks using VDFs. VDFs are special files that represent the output of some operation, such as converting file formats, extracting text, indexing a reference, etc. Whenver RepeatFS detects a file that is a valid input for one of these types of operations, it will also show a corresponding output file. When this output file is accessed (opened, copied, read), RepeatFS will automatically run the program necessary to perform the action, and populate the output file in realtime. These VDFs look and act just like normal files, though they are stored in memory. VDFs may be converted into normal files simply by copying them to another directory.
VDFs are configured within the RepeatFS configuration file as follows:
# Comments start with #
# The following entry creates a VDF that shows a corresponding FASTA for any FASTQ.
# Accessing the FASTA will automatically run seqtk
[entry] # Each VDF entry starts with [entry]
match=\.fastq$ # This regular expression controls which files are valid input
ext=.fasta # This extension will be appended to the end of the VDF
cmd=seqtk seq -A {input} # This is the command that will be run when accessing the VDF
All VDF files, including system-provided ones like provenance, or user-defined ones like the FASTQ->FASTA example above, can be found by placing a plus sign + after the input filename. So if we had a FASTQ file located at ~/mnt/example.fastq, listing the contents of ~/mnt/example.fastq+ would show the following files:
example.fastq.fasta
example.fastq.provenance.html
example.fastq.provenance.json
Note that since VDFs are treated as normal files, they also can be used as inputs to other VDFs. In this way, multiple VDFs can be chained together to perform automated operations in a modular fashion. Using the above example, if we define the following VDF:
[entry]
match=.*
ext=.count
cmd=wc -l {input}
This would create a corresponding .count for every file in the RepeatFS mount, containing the number of lines the original file contained (even if that original file was a VDF). In this way, we can view the number of lines in a FASTA converted from the FASTQ file above:
less ~/mnt/example.fastq+/example.fastq.fasta+/example.fastq.fasta.count
While VDFs are not required to record provenance in RepeatFS, they do represent a powerful tool to aid in improving reproducibilty by performing common tasks in a uniform, documented fashion.
QUESTIONS
RepeatFS is actively being developed and improved all the time. If you encounter any issue at all, please don't hesitate to reach out and file a bug report here: https://github.com/ToniWestbrook/repeatfs/issues
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file RepeatFS-0.10.0.tar.gz.
File metadata
- Download URL: RepeatFS-0.10.0.tar.gz
- Upload date:
- Size: 59.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.6.1 requests/2.23.0 setuptools/50.3.2 requests-toolbelt/0.8.0 tqdm/4.51.0 CPython/3.8.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cbd99aa66b56605feb9f5b654021053c9e28a58fdc1637815d29f6dd7e73fdf6
|
|
| MD5 |
4657a258b6265faf30c8534df20f79c5
|
|
| BLAKE2b-256 |
c1a27a49441e4b6227fbadb24e7fcd607667a10db8638f24d06678a736afabb8
|
File details
Details for the file RepeatFS-0.10.0-py3-none-any.whl.
File metadata
- Download URL: RepeatFS-0.10.0-py3-none-any.whl
- Upload date:
- Size: 84.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.6.1 requests/2.23.0 setuptools/50.3.2 requests-toolbelt/0.8.0 tqdm/4.51.0 CPython/3.8.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2eea5cf126b64ddfeb227523a4623864da84e97a2de80a4e3748251de37d3899
|
|
| MD5 |
545d10d95be1cfd5af81a81d6e849b5c
|
|
| BLAKE2b-256 |
ec4a8962741a794241105b318d70eaa3397221d6250c24e042c119649f530d20
|