STHD: probabilistic cell typing of single Spots in whole Transcriptome spatial data with High Definition
Project description
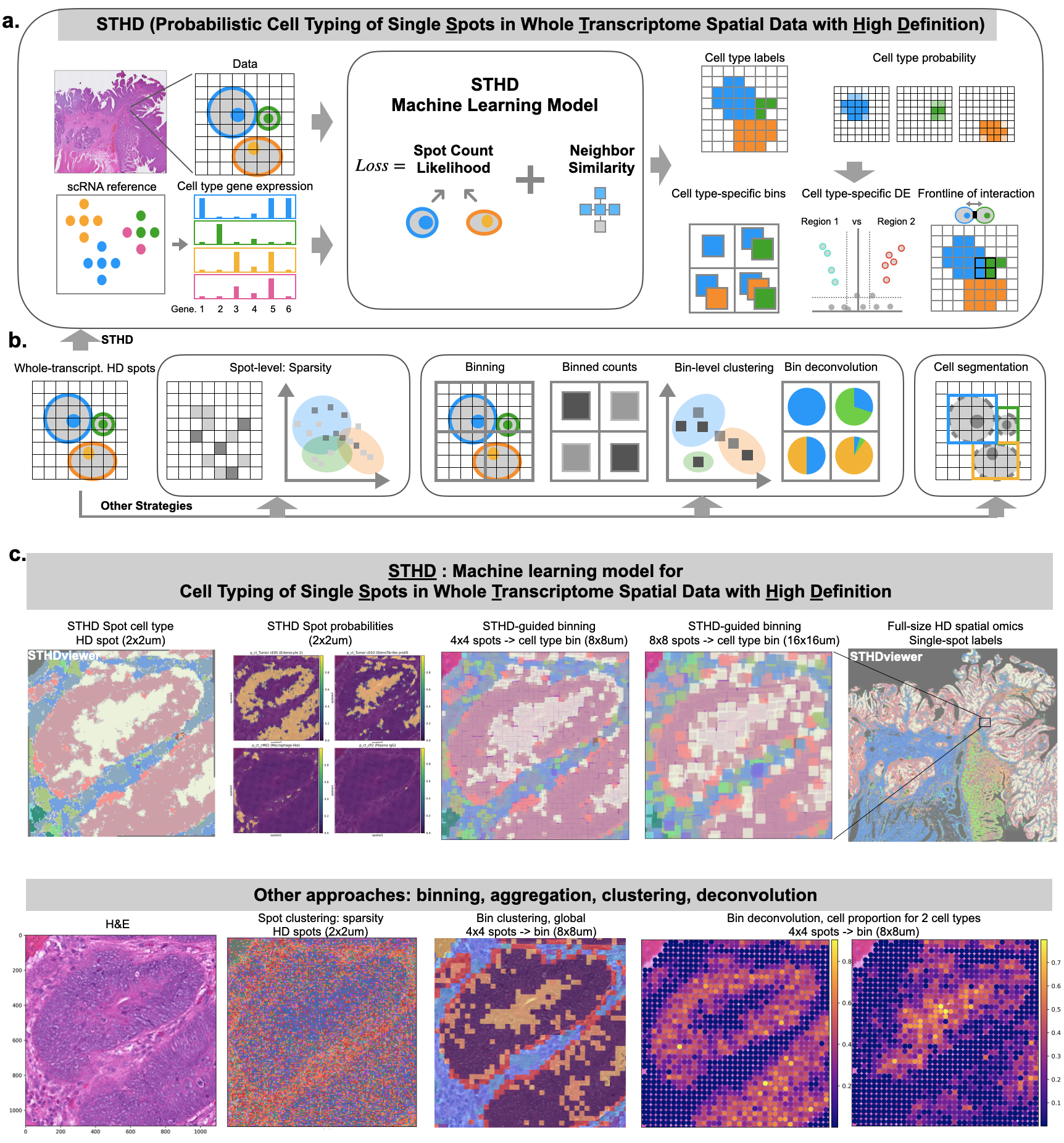
STHD: probabilistic cell typing of Single spots in whole Transcriptome spatial data with High Definition
 ---
---
- Quick start:
notebooks/tutorial.ipynb - Generates single-spot (2um) cell type labels and probabilities for VisiumHD data using a machine learning model.
- Input: VisiumHD data and reference scRNA-seq dataset with cell type annotation.
- Output: cell type labels and probabilities at 2um spot level.
- Visualization - STHDviewer: interactive, scalable, and fast spatial plot of spot cell type labels, in a HTML.
- Author: Yi Zhang, PhD, yi.zhang@duke.edu
- Website: Yi Zhang Lab at Duke
- STHDviewer of VisiumHD colon cancer sample with near 9 million spots: STHDviewer_colon_cancer_HD:https://yi-zhang-compbio-lab.github.io/STHDviewer_colon_cancer_hd
- We provided test data. Download this folder and put as
./testdata/
Install
- python version requirement: >=3.8.0
- How to use
- create new python venv
python3.8 -m venv sthd_env - activate the venv
source sthd_env/bin/activate - Install STHD from pip:
pip install STHD - Or:
- download repo:
git clone git@github.com:yi-zhang/STHD.git - install dependencies:
pip install -r STHD/requirements.txt - making sure
./STHDis in python path, e.g adding viasys.path.append('./STHD') - then in script:
from STHD import {the module you need}
- download repo:
- create new python venv
STHD Quickstart using a colon cancer VisiumHD patch:
- See
notebooks/tutorial.ipynb - The test data includes a patch crop from the VisiumHD file in
testdata/crop10
STHD pipeline on a larger VisiumHD region, or the full VisiumHD sample:
Step 1: prepare normalized gene expression profile (lambda) by cell type from reference scRNA-seq data.
- This step will generate the reference file. Details are in
notebooks/s01_build_ref_scrna.ipynb - We provided the processed file
./testdata/crc_average_expr_genenorm_lambda_98ct_4618gs.txt
Step 2: pre-processing of VisiumHD data files
- The test data includes a larger region from the VisiumHD file in
testdata/crop10large/
Preparing the VisiumHD sample.
- 10X Genomics colon cancer sample can be downloaded from: https://www.10xgenomics.com/datasets/visium-hd-cytassist-gene-expression-libraries-of-human-crc
- Required input includes 2um level spatial expression:
square_002um, which usually contains filtered_feature_bc_matrix.h5 and spatial/tissue_positions.csv . It is often from the downloaded folder "Binned outputs (all bin levels)". tissue positions in .parquet format can be converted using STHD/hdpp.py. - Required input also includes full-resolution H&E image: Visium_HD_Human_Colon_Cancer_tissue_image.btf. It is often from the "Microscope image".
- The scale factor number will also be useful, which is usually in square_002um/spatial/scalefactors_json.json
- Our processed data files are available as in:
testdata/VisiumHD/
Step 3: Patchify the large region
- This step will take a large region and split into patches. Details are in
notebooks/s11_patchify.ipynb - Or, use example command line:
# Spliting patches from a test large cropped data:
python3 -m STHD.patchify \
--spatial_path ./testdata/crop10large/all_region/adata.h5ad.gzip \
--full_res_image_path ./testdata/crop10large/all_region/fullresimg_path.json \
--load_type crop \
--dx 1500 \
--dy 1500 \
--scale_factor 0.07973422 \
--refile ./testdata/crc_average_expr_genenorm_lambda_98ct_4618gs.txt \
--save_path ./testdata/crop10large_patchify \
--mode split
- For full sample, example command line below (will take some space and time)
# Spliting patches from the full-size VisiumHD sample:
python3 -m STHD.patchify \
--spatial_path ./testdata/VisiumHD/square_002um/ \
--counts_data filtered_feature_bc_matrix.h5 \
--full_res_image_path ./testdata/VisiumHD/Visium_HD_Human_Colon_Cancer_tissue_image.btf \
--load_type original \
--dx 6000 \
--dy 6000 \
--scale_factor 0.07973422 \
--refile ./testdata/crc_average_expr_genenorm_lambda_98ct_4618gs.txt \
--save_path ./analysis/full_patchify \
--mode split
Step 4: Obtain training command line for the patch list
- This step trains STHD on each patch. The command can be flexibly modified to submit to different slurm jobs on a HPC. Details are in
notebooks/s12_per_patch_train.ipynb,Or, - Example command is:
python3 -m STHD.train --refile ./testdata/crc_average_expr_genenorm_lambda_98ct_4618gs.txt \
--patch_list ./testdata/crop10large/patches/52979_9480 ./testdata/crop10large/patches/57479_9480 ./testdata/crop10large/patches/52979_7980 ./testdata/crop10large/patches/55979_7980 ./testdata/crop10large/patches/57479_7980 ./testdata/crop10large/patches/54479_9480 ./testdata/crop10large/patches/55979_9480 ./testdata/crop10large/patches/54479_7980
Step 5: Combine the patch results
- This step combines STHD patch-wise results together. Details are in
notebooks/s13_combine_patch.ipynb, Or
#Combine predictions
python3 -m STHD.patchify \
--refile ./testdata/crc_average_expr_genenorm_lambda_98ct_4618gs.txt \
--save_path ./testdata/crop10large_patchify \
--mode combine
Step 6: Visualize!
- This step takes STHD results on a large region and generate STHDviewer for interactive exploration. Details are in
notebooks/s21_visualize.ipynb
Step 7: Downstream analyses
- One example is STHD-guided binning using a size of choice for —nspot
- Details are in
notebooks/s04_STHD_cell_type_guided_binning.ipynb; Or.
python -m STHD.binning_fast --patch_path ./testdata/crop10/ --nspot 4 --outfile ./testdata/crop10_STHDbin_nspot4.h5ad
Dependencies
requirements.txt
Reference
Sun C*, Yi Zhang*#. "STHD: probabilistic cell typing of single Spots in whole Transcriptome spatial data with High Definition". (2024) bioRxiv 2024.06.20.599803. Preprint link
Issues
Please contact Dr. Yi Zhang (yi.zhang[at]duke.edu) for desired addon features. You are welcome to follow our work by checking the Zhang lab at Duke!
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sthd-1.0.1.tar.gz.
File metadata
- Download URL: sthd-1.0.1.tar.gz
- Upload date:
- Size: 43.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
52b6632f17c765f347ac3b0c6d311071a56206743c476a3ef2454514a1288a9a
|
|
| MD5 |
8ead26d99632e5b680a508abd9cc3e10
|
|
| BLAKE2b-256 |
56cb64941eb68f747773e4cb216a3129b2f13d8c3e0677222a7c20bb9c092785
|
File details
Details for the file sthd-1.0.1-py3-none-any.whl.
File metadata
- Download URL: sthd-1.0.1-py3-none-any.whl
- Upload date:
- Size: 51.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
524a40d7cf6796ae87ed74e1429c5e540945d7a79154473cb2260bfc686f57a6
|
|
| MD5 |
a7081d17202463902778ff817cddd0f4
|
|
| BLAKE2b-256 |
a97c18c374c175f4d273e3f4d3d7b10c091929e360b9b45efc25dc1c2e1d22d7
|











