A Snakemake pipeline for alternative promoter analysis
Project description
🧬 SnakeAltPromoter Facilitates Differential Alternative Promoter Analysis
SnakeAltPromoter is a Snakemake-based pipeline for streamlined, reproducible, and scalable analysis of alternative promoter usage from RNA-seq or CAGE data. It integrates all major steps—from raw read preprocessing to promoter-level quantification and differential analysis—using state-of-the-art tools.
🧭 Workflow Overview
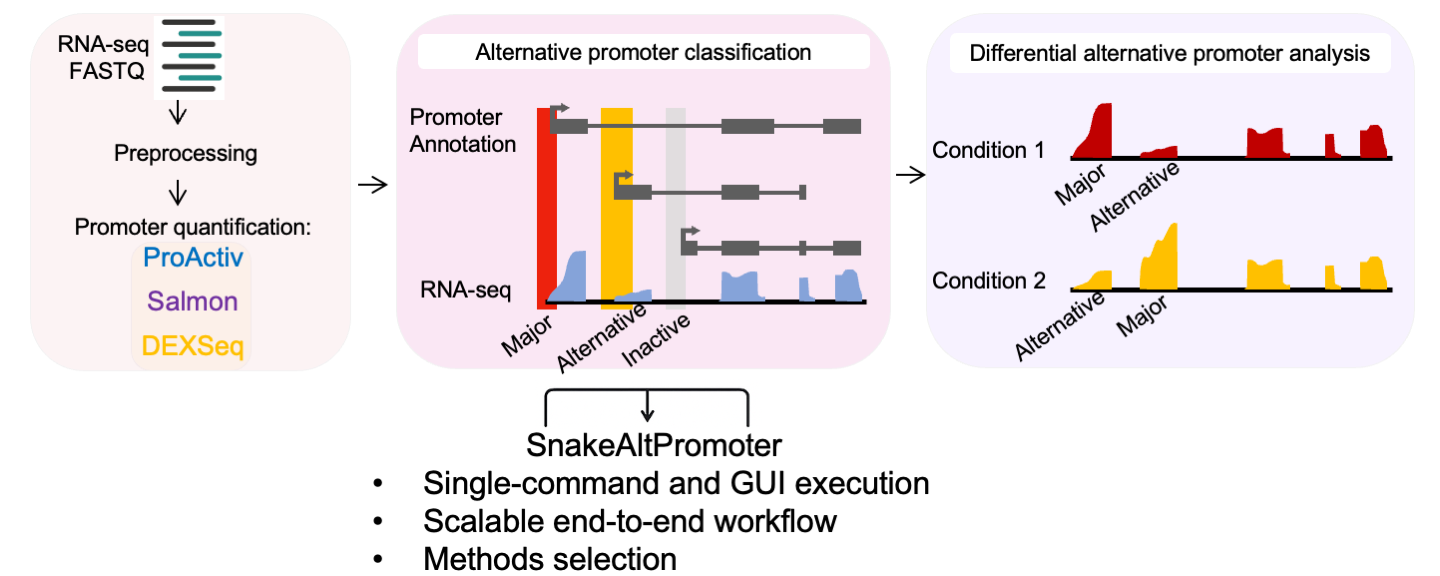
Below is the schematic overview of the SnakeAltPromoter pipeline, as shown in our paper:
Figure 1. Overview of the SnakeAltPromoter pipeline, showing genome setup, RNA-seq/CAGE processing, promoter quantification, classification, and differential promoter usage analysis.
✳️ Key Features
-
End-to-end automation of alternative promoter analysis from FASTQ to results.
-
Integrated QC and preprocessing via FastQC, TrimGalore, STAR, and MultiQC.
-
Supports multiple promoter quantification tools:
-
Built-in differential analysis using DESeq2 and ProActiv modules.
-
Fully modular, reproducible, and scalable—ideal for large multi-sample RNA-seq datasets.
🧩 Installation
1. Install via Conda (Recommended)
# Create and activate environment
conda create -n SnakeAltPromoter -c bioconda SnakeAltPromoter
conda activate SnakeAltPromoter
2. Install via Pip
pip install --user --no-cache-dir SnakeAltPromoter
3. Manual Installation (Build from Source)
git clone https://github.com/YidanSunResearchLab/SnakeAltPromoter.git
cd SnakeAltPromoter
conda create -n SnakeAltPromoter -c conda-forge python
conda activate SnakeAltPromoter
pip install .
4. Verify Installation
Snakealtpromoter --help
If successful, usage instructions and command-line options will be displayed.
🖥️ Optional: Launch GUI
For a graphical interface:
sap-ui
🧪 Minimal Test Case
Example human chr22 genome fasta and gtf files are available in the snakealtpromoter/data/ directory.
Example RNA-seq data are available in the snakealtpromoter/data/ directory.
1. Genome Setup
git clone https://github.com/YidanSunResearchLab/SnakeAltPromoter.git
cd SnakeAltPromoter
Genomesetup \
--organism hg38 \
--organism_fasta "$(pwd)/snakealtpromoter/data/hg38.fa" \
--genes_gtf "$(pwd)/snakealtpromoter/data/hg38.gtf" \
-o ./genome \
--threads 30
2. Run Test Analysis
git clone https://github.com/YidanSunResearchLab/SnakeAltPromoter.git
cd SnakeAltPromoter
Snakealtpromoter \
-i "$(pwd)/snakealtpromoter/data" \
--genome_dir "$(pwd)/genome" \
-o test_output \
--threads 30 \
--organism hg38 --trim \
--sample_sheet "$(pwd)/snakealtpromoter/data/samplesheet/samplesheet.tsv"
Output directory structure is described in the documentation:
📊 Reproduce Results from the Paper
To reproduce analyses in the SnakeAltPromoter manuscript:
- Download the GENCODE v46 genome FASTA and GTF from GENCODE.
- Retrieve heart RNA-seq and CAGE data from
GSE147236 using
fastq-dump.
1. Genome Setup
Genomesetup \
--organism hg38 \
--organism_fasta /Absolute/path/to/genome.fa \
--genes_gtf /Absolute/path/to/genes.gtf \
-o /Absolute/path/to/genome \
--threads 30
2. RNA-seq Processing
git clone https://github.com/YidanSunResearchLab/SnakeAltPromoter.git
cd SnakeAltPromoter
Snakealtpromoter \
-i /Absolute/path/to/heart_RNAseq/ \
--genome_dir /Absolute/path/to/genome \
-o heart_RNAseq_output \
--threads 30 \
--organism hg38 --trim \
--sample_sheet "$(pwd)/snakealtpromoter/data/samplesheet/Heart_RNAseq.tsv"
3. CAGE Processing
git clone https://github.com/YidanSunResearchLab/SnakeAltPromoter.git
cd SnakeAltPromoter
Snakealtpromoter \
-i /Absolute/path/to/heart_CAGE/ \
--genome_dir /Absolute/path/to/genome \
-o heart_CAGE_output \
--threads 30 \
--organism hg38 \
--sample_sheet "$(pwd)/snakealtpromoter/data/samplesheet/Heart_CAGE.tsv" \
--method cage --reads single
4. Compare with Published Results
| Supplementary Table | Description |
|---|---|
| Table 1 | Comprehensive promoter coordinates |
| Table 2 | Promoter classifications (major/minor) by ProActiv, Salmon, DEXSeq, and CAGE |
| Table 3 | Promoter counts across samples |
| Table 4 | Differential promoter activity (healthy vs. failure) across tools |
🚀 Quick Start for your own genome and sequencing data
Step 1. Genome Setup
Prepare genome indices and promoter annotations:
Genomesetup \
--organism hg38 \
--organism_fasta /Absolute/path/to/genome.fa \
--genes_gtf /Absolute/path/to/genes.gtf \
-o /Absolute/path/to/genome \
--threads 30
Step 2. Process RNA-seq Data
Run alternative promoter analysis:
Snakealtpromoter \
-i /Absolute/path/to/input_fastqs/ \
--genome_dir /Absolute/path/to/genome \
-o ./output/ \
--threads 30 \
--organism hg38 --trim \
--sample_sheet /Absolute/path/to/samplesheet.tsv \
--method cage --reads single # Add these only for CAGE data
For detailed documentation, see:
🤝 Contributing
Contributions are welcome! Please open an issue or submit a pull request via GitHub.
🧾 Citation
If you use SnakeAltPromoter, please cite:
Tan J. et al. (2025). SnakeAltPromoter Facilitates Differential Alternative Promoter Analysis. bioRxiv. https://doi.org/10.1101/2025.08.16.669128
⚖️ License
See the LICENSE file for details.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file snakealtpromoter-1.0.3.tar.gz.
File metadata
- Download URL: snakealtpromoter-1.0.3.tar.gz
- Upload date:
- Size: 19.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cb670e59cdb6f6bbf3ca92cfa8dd51f5489acee26b7e1df6248daaede52c1d74
|
|
| MD5 |
555c61db0ebec35f482fde1389d34e44
|
|
| BLAKE2b-256 |
1e06b60934d3a3e056086e2261bf211f4da8a0f40c9c7eab0be3c39f167a2c48
|
File details
Details for the file snakealtpromoter-1.0.3-py3-none-any.whl.
File metadata
- Download URL: snakealtpromoter-1.0.3-py3-none-any.whl
- Upload date:
- Size: 20.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1f16260f12453611f721dea638182e31d326deba5028bbcd984828db6e794cc9
|
|
| MD5 |
c9deaac6b268e8fdabbaf08d4548e439
|
|
| BLAKE2b-256 |
1e8b6adcb82b37de478498a49d8037c0bf13662ecdf5f22cba5e6b84dc1ebd52
|