Toolkit for analyses of amino acid sequences, optimized for GlobDB compatibility
Project description
Amino acid sequence toolkit
Table of contents
- AASTK synopsis
- Installation
- Brief descriptions of the tools in AASTK
- Usage examples
- Example plots
- Citation
AASTK synopsis
The amino acid sequence toolkit (AASTK) is a suite of tools that enables construction and analyses of protein sequence datasets from the GlobDB genomes. The GlobDB is the most comprehensive genomic resource of species-representative microbial genomes of Bacteria and Archaea, and contains the largest phylogenetic sequence diversity currently available in a single resource. AASTK is designed to leverage the GlobDB to create and analyze protein sequence datasets for evolutionary and functional studies.
AASTK uses a precomputed SQL database that includes the protein sequences, their genomic context, functional annotations from KEGG, COG, and PFAM, and metadata on the genomes the protein sequences. This SQL database is available for download on the GlobDB website.
AASTK currently consists of four tools:
pasr - Creating and updating comprehensive datasets of proteins of interest.
casm - Clustering datasets of complex functionally diverse protein superfamilies.
cugo - Assessing consensus genomic context of a protein dataset.
meta - Retrieving metadata such as taxonomy, environment, and annotations from the AASTK SQL database
Installation instructions, usage examples, and short descriptions of the tools in AASTK can be found below. A more extended description can be found in the documentation. A full description of all command line tools and arguments is available on command line reference pages for PASR, CASM, CUGO, and Meta, also accessible via the dropdown menu on the AASTK documentation page.
AASTK is actively developed and supported, so do not hesitate to submit an issue if you encounter a bug or if there's a feature you would be interested in.
Installation
Using conda (recommended)
We recommend installing AASTK using conda, in a dedicated environment created for the software.
conda create aastk
conda activate aastk
conda install -c bioconda aastk
In addition to the software and dependencies, AASTK requires a SQL database for full functionality, and is best used with the GlobDB protein complement. A fastA file of the GlobDB proteins can be exported from the AASTK SQL database using aastk export_fasta so only a single download is required.
To set up the AASTK SQL database and GlobDB protein dataset, run the following commands:
wget https://fileshare.lisc.univie.ac.at/globdb/globdb_r226/globdb_r226_aastk.db.gz
gunzip globdb_r226_aastk.db.gz
aastk export_fasta -d globdb_r226_aastk.db -n 4 -o protein_fasta_dir
Using pip
pip install aastk
Dependencies:
DIAMOND and SeqKit need to be available in your PATH.
After installation of the AASTK software and dependencies, the AASTK SQL database and GlobDB protein fastA file can be set up as described under 'Using conda' above.
From source
wget https://github.com/dspeth/aastk/archive/refs/tags/v0.1.0.tar.gz
tar -xzf v0.1.0.tar.gz
cd aastk-0.1.0
pip install .
Dependencies:
DIAMOND and SeqKit need to be available in your PATH.
After installation of the AASTK software and dependencies, the AASTK SQL database and GlobDB protein fastA file can be set up as described under 'Using conda' above.
Brief descriptions of the tools in AASTK
Protein alignment score ratio (PASR)
PASR is intended for the creation of comprehensive sequence datasets of homologous proteins, using alignment of all sequences in a query dataset to a (small) seed dataset of sequences of interest using DIAMOND. Thus, the query file is the large dataset containing all proteins to be searched, and the seed is the (preliminary) dataset of homologous proteins to be found in the query data.
The PASR workflow can be run using aastk pasr, and the helper tool pasr_select can be used to select sequences to include in the final dataset, based on the PASR plot.
More extensive documentation for PASR is available on the documentation page and all command line options for PASR are available on the PASR command line reference page.
Clustering alignment score matrix (CASM)
CASM is designed to investigate the structure of a dataset of homologous proteins, such as a protein superfamily. Many protein (super)families contain members with distinct biochemical or physiological functions. Understanding the structure of a protein (super)family is an essential first step in understanding the functional landscape of a protein (super)family. CASM clusters sequences by generating an N x n alignment score matrix, by aligning all N sequences in a dataset against a subset of n sequences. T-distributed stochastic neighbourhood embedding (t-SNE) is then used to reduce this matrix to from n to 2 dimensions. Clusters are called using DBSCAN, and the t-SNE results can be annotated with metadata and visualized.
The CASM workflow can be run using aastk casm, and the helper tool pasr_select can be used to select sequences to include in the final dataset, based on the PASR plot.
More extensive documentation for CASM is available on the documentation page and all command line options for CASM are available on the CASM command line reference page.
Colocated unidirectional gene organization (CUGO)
CUGO is intended to retrieve and visualise the consensus genomic context of a dataset of homologous proteins. To do so, it uses information from the AASTK SQL database of the proteins in the GlobDB genomes. Each GlobDB genome is partitioned into CUGO units, that are limited by a strand change of encoded proteins, or when a contig ends. For each sequence in the input file, aastk cugo determines it's CUGO unit, and then extracts all sequences belonging to this CUGO unit, and optionally adjacent CUGOs. These sequences are then used for visualisation.
Consensus genomic context is visualised in three plots combined into one overview figure. The first plot shows the three most common annotations per genomic position, the second the density of amino acid sequence length per position, and the third the density of number of transmembrane helices per position (see figure below). This design ensures that there is no upper limit to the size of the query dataset size from a visualisation perspective, although data retrieval time scales with query size. CUGO uses the AASTK SQL database, which as of release 226 is 310 Gb.
Metadata retrieval (Meta)
Meta allows for retrieval of sequence metadata from the AASTK SQL database based on a protein fasta file or a list of protein identifiers. There are two types of metadata in the SQL database, those linked to protein sequences directly, and those linked to the genomes encoding the protein sequences. Available metadata categories are annotation (protein linked), taxonomy, culture collection availability, and two levels of environmental metadata (all genome linked). The selected metadata are written to a tsv file.
Usage examples
PASR
aastk pasr -m BLOSUM45 -q query.fasta -s seed.fasta -o output_dir
Where:
-m specifies the scoring matrix to be used, can be BLOSUM62 or BLOSUM45
-q specifies the query dataset in fasta format
-s specifies the seed database of homologous proteins, in fasta format
-o specifies the output directory
All command line options for PASR are available on the PASR command line reference page.
CASM
aastk casm --fasta input.faa -o output_directory --subset_size 1000 -n 4 -p 1000
Where:
--fasta and -o control the input and output
--subset_size determines the number of randomly selected proteins in the subset
-n specifies the number of threads to use
-p is the t-SNE perplexiity
All command line options for CASM are available on the CASM command line reference page.
CUGO
aastk cugo -r 0 -l -3 -u 6 -d aastk_sql_database.db -f fasta_file.faa -o cugo_output_dir
Where:
-r is the range of CUGO numbers to be considered
-l specifies the number of positions upstream of the gene of interest
-u specifies the number of positions downstream of the gene of interest
-d is the path to the AASTK SQL database
-f and -o control the input and output
All command line options for CASM are available on the CUGO command line reference page.
Meta
aastk meta --all_metadata -d aastk_sql_database.db -f fasta_file.faa -o output_directory
Where:
--all_metadata controls which metadata is added to the output tsv file
-d is the path to the AASTK SQL database
-f and -o control the input and output
All command line options for CASM are available on the Meta command line reference page.
Example plots
Examples of the graphical output of PASR, CUGO, and CASM. For a full overview of the output files generated by these tools, consult the AASTK documentation page.
Example PASR plot
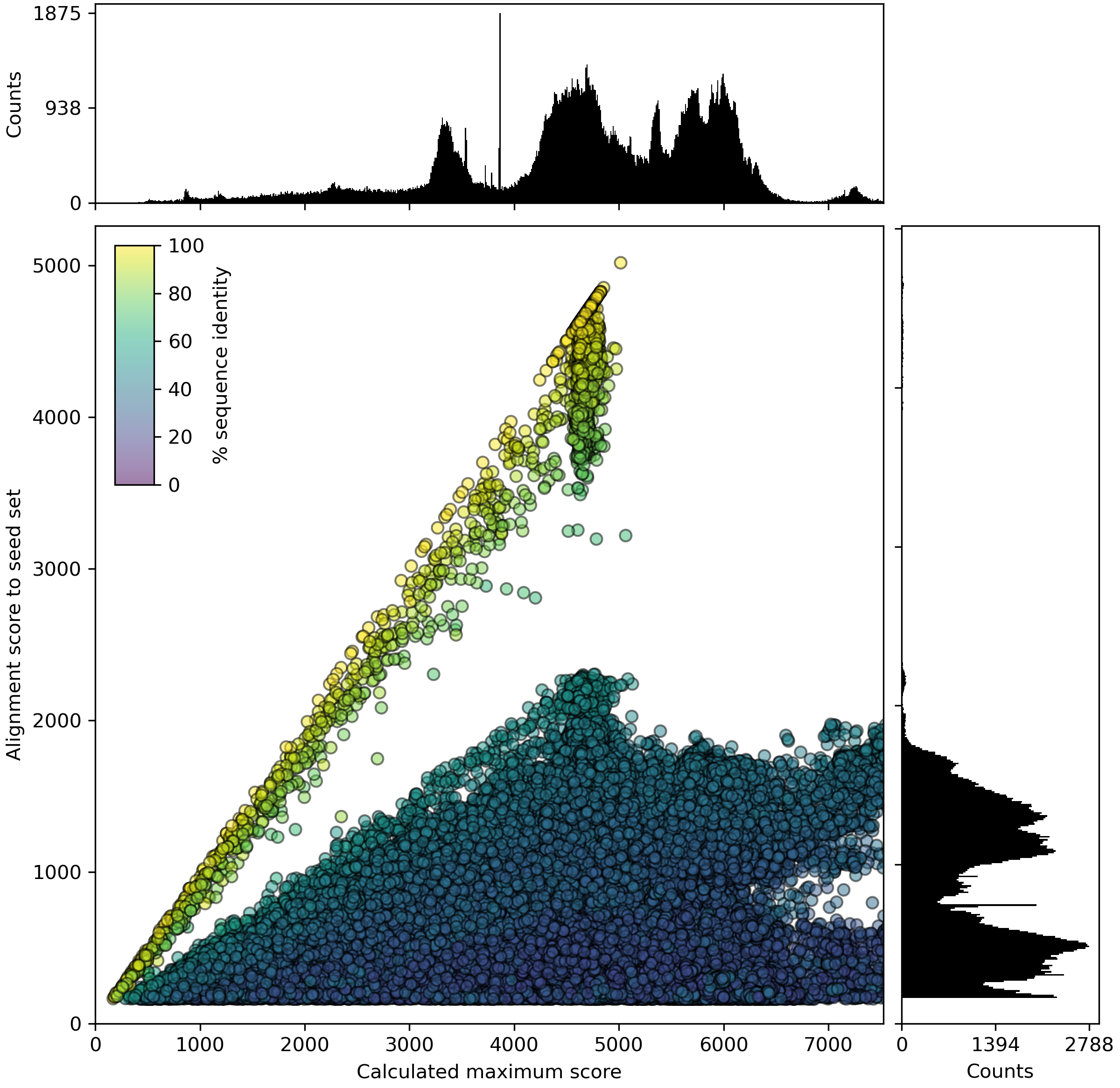
Example CUGO plot
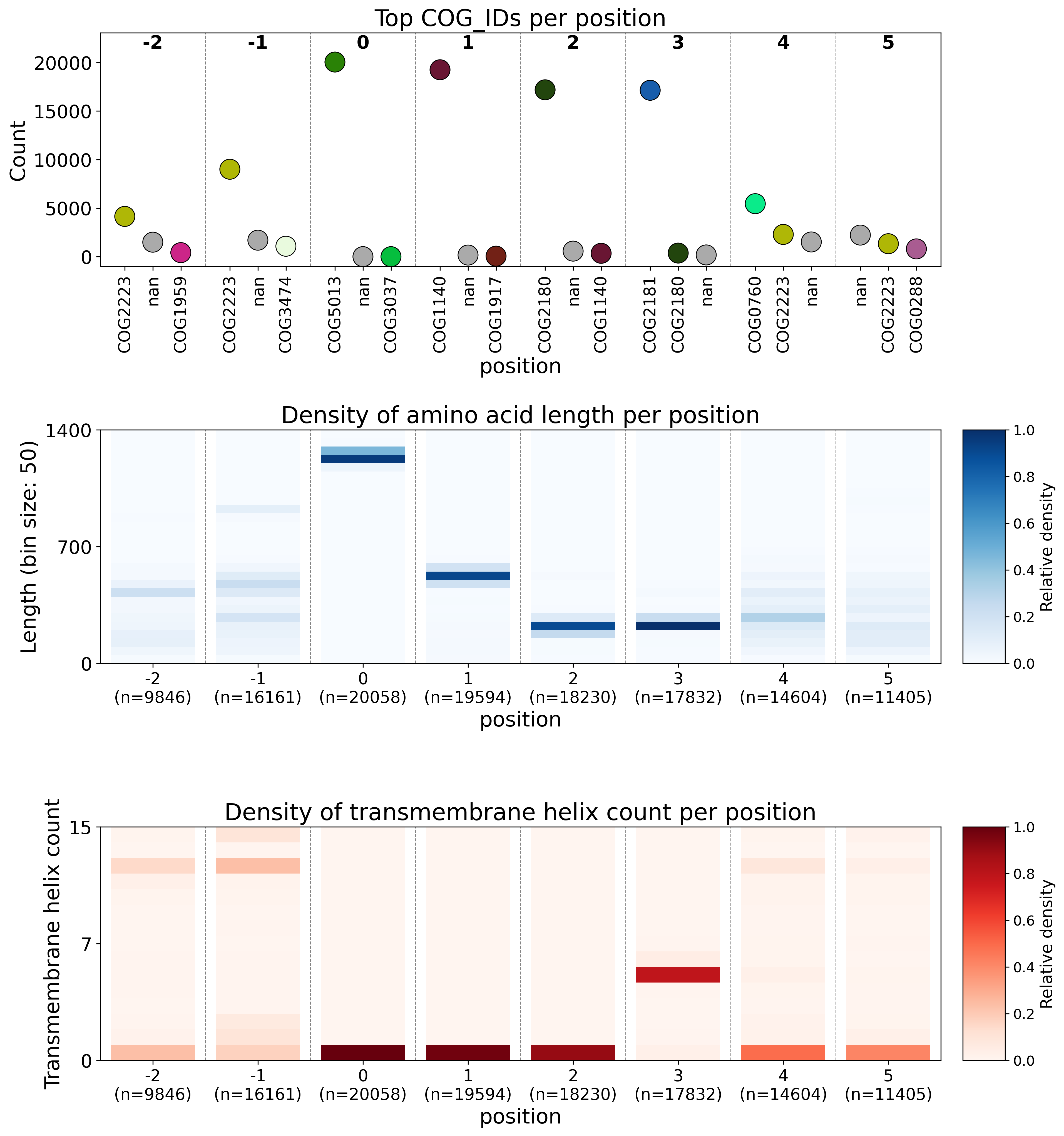
Example CASM plot after early exaggeration phase.
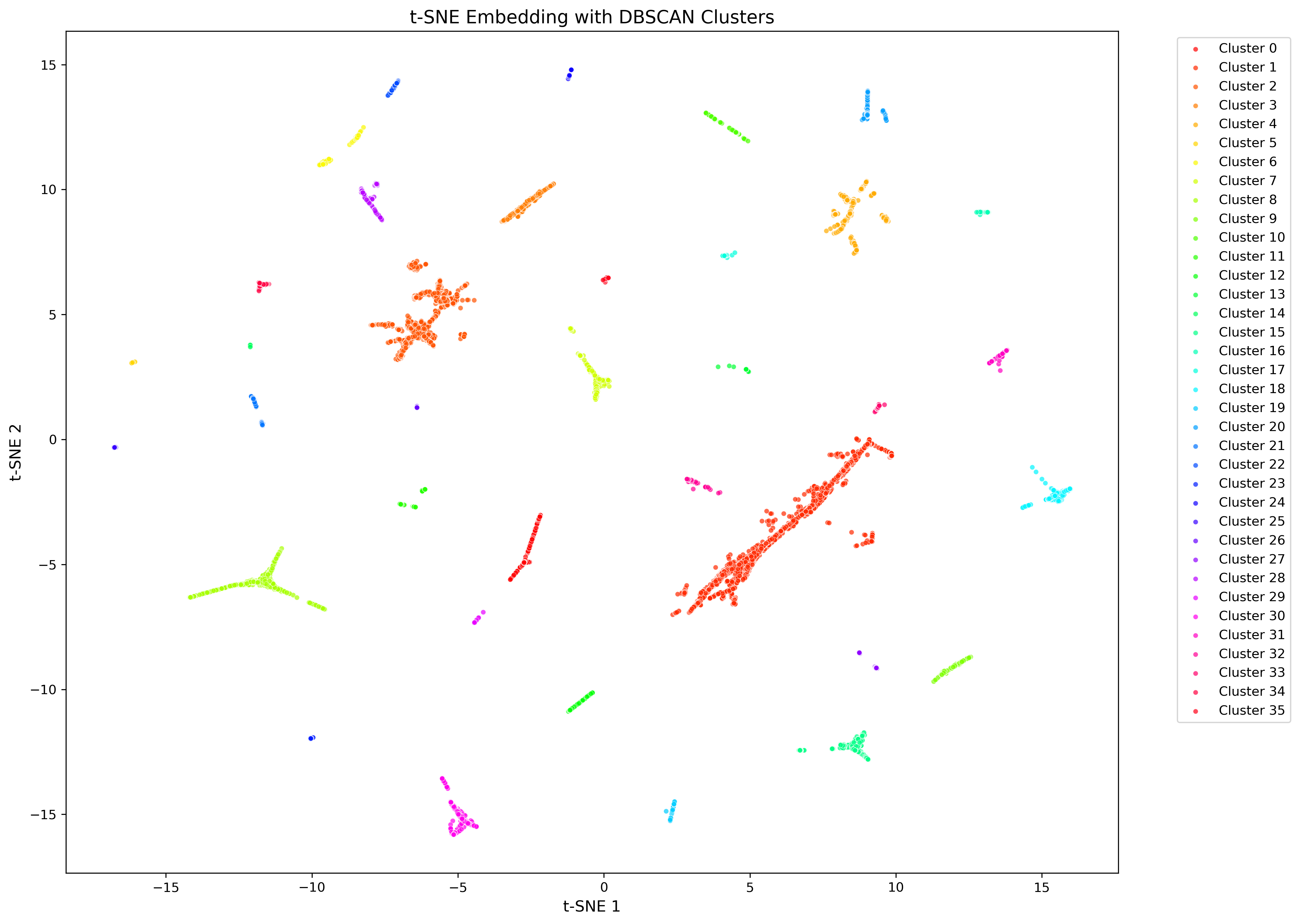
Example of final CASM plot
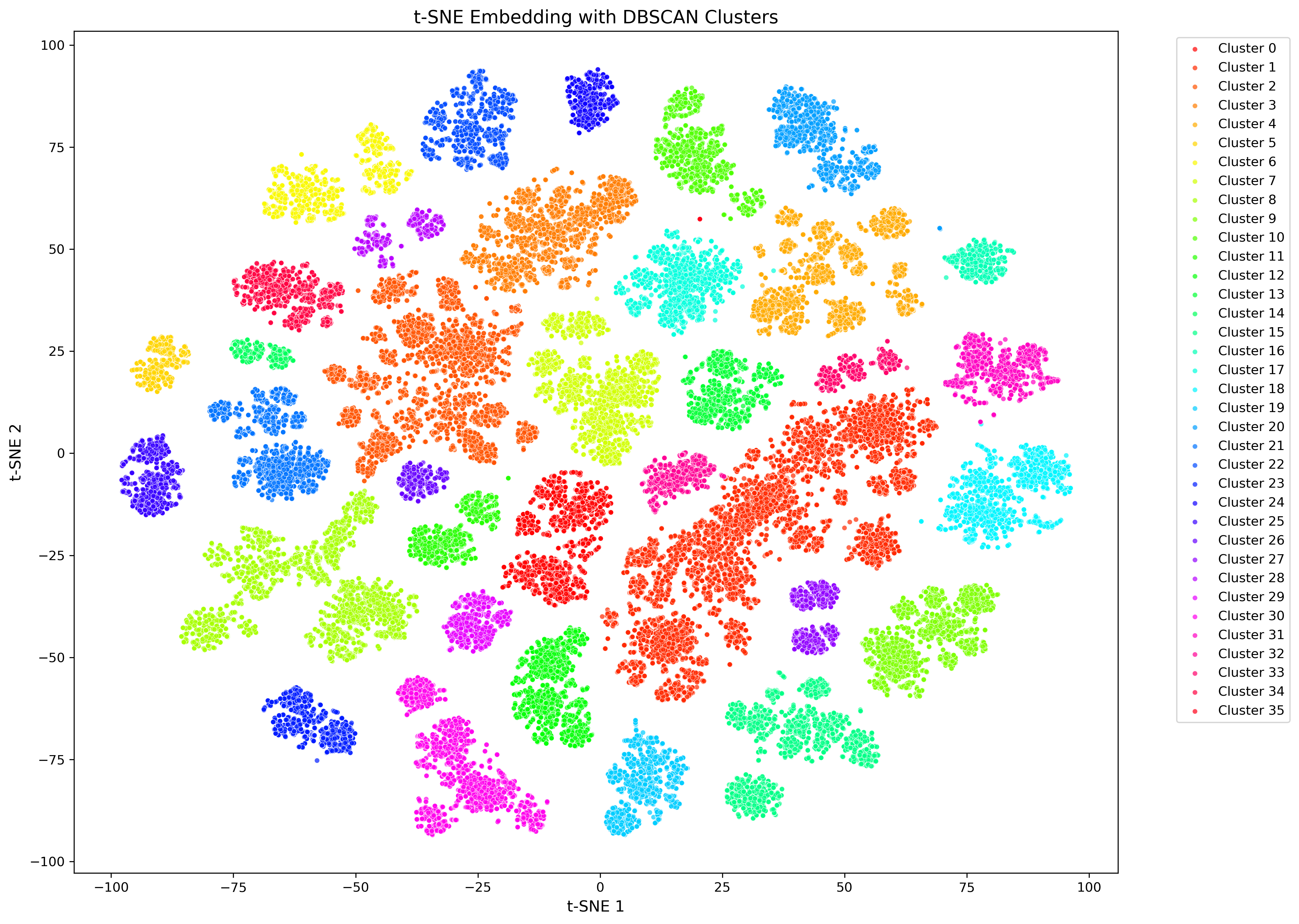
Citation
There is no publication describing AASTK yet, so please cite this repository when you use AASTK.
In addition, several parts of the software were developed independently and should be credited.
-
If you use AASTK with the GlobDB protein dataset, please cite:
Speth et al. (2025) GlobDB: a comprehensive species-dereplicated microbial genome resource
https://doi.org/10.1093/bioadv/vbaf280 -
If you use
aastk pasr, please cite:
Speth and Orphan (2018) Metabolic marker gene mining provides insight in global mcrA diversity and, coupled with targeted genome reconstruction, sheds further light on metabolic potential of the Methanomassiliicoccales
https://doi.org/10.7717/peerj.5614 -
The environmental data from
aastk metais derived from the MetaCoOc software. A manuscript is in preparation, but in the meantime please cite:
https://github.com/bcoltman/metacooc
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file aastk-0.1.1.tar.gz.
File metadata
- Download URL: aastk-0.1.1.tar.gz
- Upload date:
- Size: 69.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
239c811419c812fbc433fd690e7276b7b1d3b91438e3c6072c4938d2b8ac65da
|
|
| MD5 |
cc5bacee420d9e18a4d391e6519d48fa
|
|
| BLAKE2b-256 |
3f1ef3cfb8e707bdcf30fe486c8d56326a10898e7902924304a152872ea0d3e2
|
Provenance
The following attestation bundles were made for aastk-0.1.1.tar.gz:
Publisher:
release.yml on dspeth/aastk
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
aastk-0.1.1.tar.gz -
Subject digest:
239c811419c812fbc433fd690e7276b7b1d3b91438e3c6072c4938d2b8ac65da - Sigstore transparency entry: 1199437996
- Sigstore integration time:
-
Permalink:
dspeth/aastk@0a1e5cdfb4e0d6273599ce7d15f0d8eb7438c8fa -
Branch / Tag:
refs/tags/v0.1.1 - Owner: https://github.com/dspeth
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@0a1e5cdfb4e0d6273599ce7d15f0d8eb7438c8fa -
Trigger Event:
release
-
Statement type:
File details
Details for the file aastk-0.1.1-py3-none-any.whl.
File metadata
- Download URL: aastk-0.1.1-py3-none-any.whl
- Upload date:
- Size: 68.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d00bd1de8112373e21059431546a81a6eea3b0bb23dd3881f1f14f31a51a8302
|
|
| MD5 |
7c12879ea45f5b8693d538d47ba48f26
|
|
| BLAKE2b-256 |
14000d5e322533ee6777c3ba6975c894d55b5be2be07ea69b96efb877d1b761e
|
Provenance
The following attestation bundles were made for aastk-0.1.1-py3-none-any.whl:
Publisher:
release.yml on dspeth/aastk
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
aastk-0.1.1-py3-none-any.whl -
Subject digest:
d00bd1de8112373e21059431546a81a6eea3b0bb23dd3881f1f14f31a51a8302 - Sigstore transparency entry: 1199438011
- Sigstore integration time:
-
Permalink:
dspeth/aastk@0a1e5cdfb4e0d6273599ce7d15f0d8eb7438c8fa -
Branch / Tag:
refs/tags/v0.1.1 - Owner: https://github.com/dspeth
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@0a1e5cdfb4e0d6273599ce7d15f0d8eb7438c8fa -
Trigger Event:
release
-
Statement type:
















