Helper python package for ATLAS Common NTuple Analysis work.
Project description
atlas-schema v0.4.1
This is the python package containing schemas and helper functions enabling analyzers to work with ATLAS datasets (Monte Carlo and Data), using coffea.
Hello World
The simplest example is to just get started processing the file as expected:
from atlas_schema.schema import NtupleSchema
from coffea import dataset_tools
import awkward as ak
fileset = {"ttbar": {"files": {"path/to/ttbar.root": "tree_name"}}}
samples, report = dataset_tools.preprocess(fileset)
def noop(events):
return ak.fields(events)
fields = dataset_tools.apply_to_fileset(noop, samples, schemaclass=NtupleSchema)
print(fields)
which produces something similar to
{
"ttbar": [
"dataTakingYear",
"mcChannelNumber",
"runNumber",
"eventNumber",
"lumiBlock",
"actualInteractionsPerCrossing",
"averageInteractionsPerCrossing",
"truthjet",
"PileupWeight",
"RandomRunNumber",
"met",
"recojet",
"truth",
"generatorWeight",
"beamSpotWeight",
"trigPassed",
"jvt",
]
}
However, a more involved example to apply a selection and fill a histogram looks like below:
import awkward as ak
from hist import Hist
import matplotlib.pyplot as plt
from coffea import processor
from distributed import Client
from atlas_schema.schema import NtupleSchema
class MyFirstProcessor(processor.ProcessorABC):
def __init__(self):
pass
def process(self, events):
dataset = events.metadata["dataset"]
h_ph_pt = (
Hist.new.StrCat(["all", "pass", "fail"], name="isEM")
.Regular(200, 0.0, 2000.0, name="pt", label="$pt_{\gamma}$ [GeV]")
.Int64()
)
cut = ak.all(events.ph.isEM, axis=1)
h_ph_pt.fill(isEM="all", pt=ak.firsts(events.ph.pt / 1.0e3))
h_ph_pt.fill(isEM="pass", pt=ak.firsts(events[cut].ph.pt / 1.0e3))
h_ph_pt.fill(isEM="fail", pt=ak.firsts(events[~cut].ph.pt / 1.0e3))
return {
dataset: {
"entries": ak.num(events, axis=0),
"ph_pt": h_ph_pt,
}
}
def postprocess(self, accumulator):
pass
if __name__ == "__main__":
client = Client()
fileset = {"700352.Zqqgamma.mc20d.v1": {"files": {"ntuple.root": "analysis"}}}
run = processor.Runner(
executor=processor.IterativeExecutor(compression=None),
schema=NtupleSchema,
savemetrics=True,
)
out, metrics = run(fileset, processor_instance=MyFirstProcessor())
print(out)
print(metrics)
fig, ax = plt.subplots()
computed["700352.Zqqgamma.mc20d.v1"]["ph_pt"].plot1d(ax=ax)
ax.set_xscale("log")
ax.legend(title="Photon pT for Zqqgamma")
fig.savefig("ph_pt.pdf")
which produces
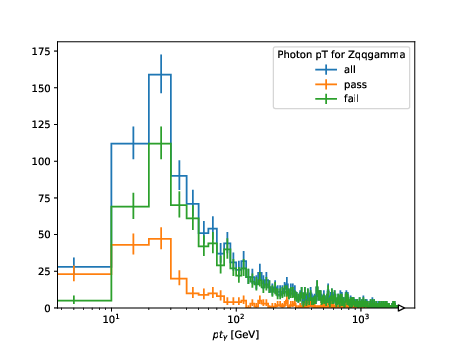
Processing with Systematic Variations
For analyses requiring systematic uncertainty evaluation, you can easily iterate
over all systematic variations using the new events["NOSYS"] alias and
systematic_names property:
import awkward as ak
from hist import Hist
from coffea import processor
from atlas_schema.schema import NtupleSchema
class SystematicsProcessor(processor.ProcessorABC):
def __init__(self):
self.h = (
Hist.new.StrCat([], name="variation", growth=True)
.Regular(50, 0.0, 500.0, name="jet_pt", label="Leading Jet $p_T$ [GeV]")
.Int64()
)
def process(self, events):
dsid = events.metadata["dataset"]
# Process all systematic variations including nominal ("NOSYS")
for variation in events.systematic_names:
event_view = events[variation]
# Fill histogram with leading jet pT for this systematic variation
leading_jet_pt = event_view.jet.pt[:, 0] / 1_000 # Convert MeV to GeV
weights = (
event_view.weight.mc
if hasattr(event_view, "weight")
else ak.ones_like(leading_jet_pt)
)
self.h.fill(variation=variation, jet_pt=leading_jet_pt, weight=weights)
return {
"hist": self.h,
"meta": {"sumw": {dsid: {(events.metadata["fileuuid"], ak.sum(weights))}}},
}
def postprocess(self, accumulator):
return accumulator
This approach allows you to seamlessly process both nominal and systematic variations in a single loop, eliminating the need for special-case handling of the nominal variation.
Developer Notes
Converting Enums from C++ to Python
This useful vim substitution helps:
%s/ \([A-Za-z]\+\)\s\+= \(\d\+\),\?/ \1: Annotated[int, "\1"] = \2
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file atlas_schema-0.4.1.tar.gz.
File metadata
- Download URL: atlas_schema-0.4.1.tar.gz
- Upload date:
- Size: 22.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
69b0e349a777b13c45d71a7d195900f764474a592bb4093f2ff771306b8f25fc
|
|
| MD5 |
89e4089be786bf4903ef7aa9c7d03a89
|
|
| BLAKE2b-256 |
d2ca4a4acf3325b412fcb0643f02b13db74e65306f624da659c8d562346a6a9d
|
Provenance
The following attestation bundles were made for atlas_schema-0.4.1.tar.gz:
Publisher:
cd.yml on scipp-atlas/atlas-schema
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
atlas_schema-0.4.1.tar.gz -
Subject digest:
69b0e349a777b13c45d71a7d195900f764474a592bb4093f2ff771306b8f25fc - Sigstore transparency entry: 591022550
- Sigstore integration time:
-
Permalink:
scipp-atlas/atlas-schema@c1e083312cabdb2a2ff9fe7b6de71a630304ea83 -
Branch / Tag:
refs/tags/v0.4.1 - Owner: https://github.com/scipp-atlas
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
cd.yml@c1e083312cabdb2a2ff9fe7b6de71a630304ea83 -
Trigger Event:
release
-
Statement type:
File details
Details for the file atlas_schema-0.4.1-py3-none-any.whl.
File metadata
- Download URL: atlas_schema-0.4.1-py3-none-any.whl
- Upload date:
- Size: 26.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
60255123ca753526486edeced16075da9711392783f794e617518c1b47cffdc2
|
|
| MD5 |
b846bfdf3fce961bca86c1186d769f6a
|
|
| BLAKE2b-256 |
4815a21d112f8b277fa0fd02fc2714c8c85874b9f0654d1eae5b7f1347f09c3f
|
Provenance
The following attestation bundles were made for atlas_schema-0.4.1-py3-none-any.whl:
Publisher:
cd.yml on scipp-atlas/atlas-schema
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
atlas_schema-0.4.1-py3-none-any.whl -
Subject digest:
60255123ca753526486edeced16075da9711392783f794e617518c1b47cffdc2 - Sigstore transparency entry: 591022555
- Sigstore integration time:
-
Permalink:
scipp-atlas/atlas-schema@c1e083312cabdb2a2ff9fe7b6de71a630304ea83 -
Branch / Tag:
refs/tags/v0.4.1 - Owner: https://github.com/scipp-atlas
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
cd.yml@c1e083312cabdb2a2ff9fe7b6de71a630304ea83 -
Trigger Event:
release
-
Statement type:

















