AUDIT, Analysis & evalUation Dashboard of artIficial inTelligence
Project description
Table of Contents
Summary
AUDIT, Analysis & evalUation Dashboard of artIficial inTelligence, is a tool designed to provide researchers and developers with an interactive way to better analyze and explore MRI datasets and segmentation models. Given its functionalities to extract the most relevant features and metrics from your multiple data sources, it allows for uncovering biases both intra and inter-dataset as well as in the model predictions.
Details of our work are provided in our paper AUDIT: An open-source Python library for AI model evaluation with use cases in MRI brain tumor segmentation. We hope that users will leverage AUDIT to gain novel insights into the field of medical image segmentation.
AUDIT workflow
The diagram below illustrates the overall workflow of AUDIT, from input data to data visualization on the APP.
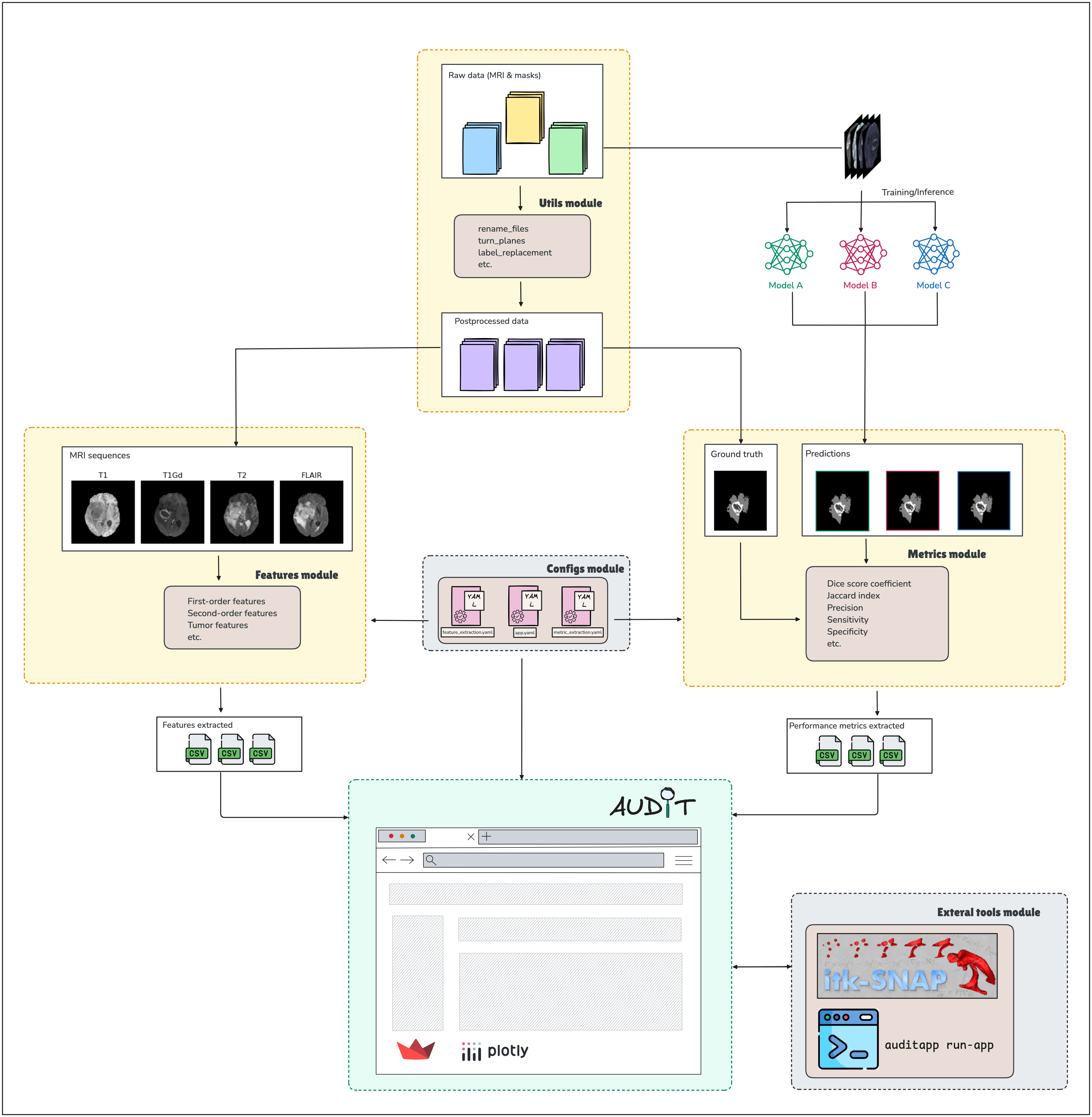
For more details, please refer to the AUDIT paper.
AUDIT analysis modes:
- Home Page: The main landing page of the tool.
- Univariate: Exploration of individual features to understand how they are distributed.
- Multivariate: Analysis of multiple features simultaneously to explore relationships and hidden patterns.
- Segmentation error matrix: Overview of disagreements between ground truth and predicted segmentation through a class-wise error matrix.
- Single model performance: Evaluation of the performance of a single model based on extracted features.
- Pairwise model performance: Perform pairwise comparisons between models to find statistically significant differences.
- Multi-model performance: Comparative analysis of performance metrics across multiple models.
- Longitudinal measurements: Analysis of data collected over time to observe trends and changes in model performance.
- Subjects' exploration: Detailed examination of individual subjects within the dataset.
Online Web application
Last released version of AUDIT is hosted at https://auditapp.streamlitapp.com for an online overview of its functionalities.
Getting Started
AUDIT can be installed either from our GitHub repository or from PyPI using the command pip install auditapp. In this guide, we will show how to install it from the GitHub repository. For a more detailed exploration of AUDIT, please check our official documentation.
1 Installation
Create an isolated Anaconda environment (recommended to avoid dependency conflicts):
conda create -n audit_env python=3.10
conda activate audit_env
Clone the repository:
git clone https://github.com/caumente/AUDIT.git
cd AUDIT
Install the required packages:
pip install -r requirements.txt
2. Configuration
Edit the config files in ./src/audit/configs/ directory to set up the paths for data loading and other configurations:
2.1 Feature extraction config file
This configuration file is used to set paths, labels, features, and longitudinal study parameters for feature extraction in AUDIT.
Show configuration
# Paths to all the datasets
data_paths:
BraTS2020: '/home/usr/AUDIT/datasets/BraTS2020/BraTS2020_images'
BraTS2024_PED: '/home/usr/AUDIT/datasets/BraTS2024_PED/BraTS2024_PED_images'
BraTS2024_SSA: '/home/usr/AUDIT/datasets/BraTS2024_SSA/BraTS2024_SSA_images'
UCSF: '/home/usr/AUDIT/datasets/UCSF/UCSF_images'
LUMIERE: '/home/usr/AUDIT/datasets/LUMIERE/LUMIERE_images'
# Sequences available
sequences:
- '_t1'
- '_t2'
- '_t1ce'
- '_flair'
# Mapping of labels to their numeric values
labels:
BKG: 0
EDE: 3
ENH: 1
NEC: 2
# List of features to extract
features:
statistical: true
texture: true
spatial: true
tumor: true
# Longitudinal study settings
longitudinal:
UCSF:
pattern: "_" # Pattern used for splitting filename
longitudinal_id: 1 # Index position for the subject ID after splitting the filename. Starting by 0
time_point: 2 # Index position for the time point after splitting the filename. Starting by 0
LUMIERE:
pattern: "-"
longitudinal_id: 1
time_point: 3
# Path where extracted features will be saved
output_path: '/home/usr/AUDIT/outputs/features'
logs_path: '/home/usr/AUDIT/logs/features'
# others
cpu_cores: 8
2.2 Metric extraction config file
This configuration file defines the paths to datasets and model predictions, the labels, metrics to compute, and output paths for metric extraction in AUDIT.
Show configuration
# Path to the raw dataset
data_path: '/home/usr/AUDIT/datasets/BraTS2024_PED/BraTS2024_PED_images'
# Paths to model predictions
model_predictions_paths:
nnUnet: '/home/usr/AUDIT/datasets/BraTS2024_PED/BraTS2024_PED_seg/nnUnet'
SegResNet: '/home/usr/AUDIT/datasets/BraTS2024_PED/BraTS2024_PED_seg/SegResNet'
# Mapping of labels to their numeric values
labels:
BKG: 0
EDE: 3
ENH: 1
NEC: 2
# List of metrics to compute
metrics:
dice: true
jacc: true
accu: true
prec: true
sens: true
spec: true
haus: true
size: true
# Library used for computing all the metrics
package: audit
# Path where output metrics will be saved
output_path: '/home/usr/AUDIT/outputs/metrics'
filename: 'BraTS2024_PED'
logs_path: '/home/usr/AUDIT/logs/metric'
# others
cpu_cores: 8
2.3 APP config file
This configuration file sets up paths for datasets, extracted features, metrics, and model predictions. It also defines available sequences and label mappings used by the AUDIT application.
Show configuration
# Sequences available. First of them will be used to compute properties like spacing
sequences:
- '_t1'
- '_t2'
- '_t1ce'
- '_flair'
# Mapping of labels to their numeric values
labels:
BKG: 0
EDE: 3
ENH: 1
NEC: 2
# Root path for datasets, features extracted, and metrics extracted
datasets_path: './datasets' # '/home/usr/AUDIT/datasets'
features_path: './outputs/features' # '/home/usr/AUDIT/outputs/features'
metrics_path: './outputs/metrics' # '/home/usr/AUDIT/outputs/metrics'
# Paths for raw datasets
raw_datasets:
BraTS2020: "${datasets_path}/BraTS2020/BraTS2020_images"
BraTS2024_SSA: "${datasets_path}/BraTS2024_SSA/BraTS2024_SSA_images"
BraTS2024_PED: "${datasets_path}/BraTS2024_PED/BraTS2024_PED_images"
UCSF: "${datasets_path}/UCSF/UCSF_images"
LUMIERE: "${datasets_path}/LUMIERE/LUMIERE_images"
# Paths for feature extraction CSV files
features:
BraTS2020: "${features_path}/extracted_information_BraTS2020.csv"
BraTS2024_SSA: "${features_path}/extracted_information_BraTS2024_SSA.csv"
BraTS2024_PED: "${features_path}/extracted_information_BraTS2024_PED.csv"
UCSF: "${features_path}/extracted_information_UCSF.csv"
LUMIERE: "${features_path}/extracted_information_LUMIERE.csv"
# Paths for metric extraction CSV files
metrics:
BraTS2024_SSA: "${metrics_path}/extracted_information_BraTS2024_SSA.csv"
BraTS2024_PED: "${metrics_path}/extracted_information_BraTS2024_PED.csv"
UCSF: "${metrics_path}/extracted_information_UCSF.csv"
LUMIERE: "${metrics_path}/extracted_information_LUMIERE.csv"
# Paths for models predictions
predictions:
BraTS2024_SSA:
nnUnet: "${datasets_path}/BraTS2024_SSA/BraTS2024_SSA_seg/nnUnet"
SegResNet: "${datasets_path}/BraTS2024_SSA/BraTS2024_SSA_seg/SegResNet"
BraTS2024_PED:
nnUnet: "${datasets_path}/BraTS2024_PED/BraTS2024_PED_seg/nnUnet"
SegResNet: "${datasets_path}/BraTS2024_PED/BraTS2024_PED_seg/SegResNet"
3. Run AUDIT backend
Use the following commands in your terminal to run the Feature extraction and Metric extraction scripts:
python src/audit/feature_extraction.py
python src/audit/metric_extraction.py
After running either script, a logs folder will be created to keep track of the execution. All output files will be stored in the folder defined in the corresponding config file (by default, in the outputs folder).
4. Run AUDIT app
AUDIT app is built on top of the Streamlit library. Use the following command to run the app and start exploring your data:
python src/audit/app/launcher.py
5. Additional configurations
5.1. ITK-Snap
AUDIT can be adjusted for opening cases with ITK-Snap while exploring the data in the different dashboards. The ITK-Snap tool must have been installed and preconfigured before. Below are placeholders for the necessary configuration for each operating system. Information will be added soon.
5.1.1. On Mac OS
show configuration
⚠️ Configuration instructions for Mac OS are not available yet. Please check back later.
5.1.2. On Linux OS
Show configuration
A brief tutorial to install ITK-SNAP 4.4
- Download ITK-SNAP 4.4
wget https://downloads.itksnap.org/itksnap-4.4.0-20250909-Linux-x86_64.tar.gz
- Extract and move to
/opt
sudo tar -xvzf itksnap-4.4.0-20250909-Linux-x86_64.tar.gz -C /opt
sudo mv /opt/itksnap-4.4.0-20250909-Linux-x86_64 /opt/itksnap
- Create a wrapper script to run from anywhere
sudo nano /usr/local/bin/itksnap
File content:
#!/bin/bash
DIR=/opt/itksnap
"$DIR/bin/itksnap" "$@"
Make it executable:
sudo chmod +x /usr/local/bin/itksnap
- Run ITK-SNAP 4.4
itksnap
Authors
Please feel free to contact us with any issues, comments, or questions.
- Carlos Aumente (UO297103@uniovi.es)
- Mauricio Reyes
- Michael Muller
- Jorge Díez
- Beatriz Remeseiro
License
Apache License 2.0
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file auditapp-0.1.2.tar.gz.
File metadata
- Download URL: auditapp-0.1.2.tar.gz
- Upload date:
- Size: 1.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.1.4 CPython/3.10.14 Darwin/24.6.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
eebeffac21340b217bc55b8808e051b90cf5047686e2c7c7ea899c6a40bfbda0
|
|
| MD5 |
c1809dfa4a5fd509e8c6543e818235be
|
|
| BLAKE2b-256 |
3cd28cde72d5a71c7b359891d0268e8e959392f2c9aa6a913868b354d9b921a2
|
File details
Details for the file auditapp-0.1.2-py3-none-any.whl.
File metadata
- Download URL: auditapp-0.1.2-py3-none-any.whl
- Upload date:
- Size: 1.1 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.1.4 CPython/3.10.14 Darwin/24.6.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dc3f7c264c9d6c31b6110beb1fc6e7cb19b0ffc91514061824588da899709334
|
|
| MD5 |
1a8344dd2b81cf4d1a9aaf646bfaa807
|
|
| BLAKE2b-256 |
54e3338e1b548c367cd5a9543198347c053fedd4a435c161240b510d93b2d872
|





















