A metric learning toolkit
Project description
BioEncoder
BioEncoder is a toolkit for supervised metric learning to i) learn and extract features from images, ii) enhance biological image classification, and iii) identify the features most relevant to classification. Designed for diverse and complex datasets, the package and the available metric losses can handle unbalanced classes and subtle phenotypic differences more effectively than non-metric approaches. The package includes taxon-agnostic data loaders, custom augmentation techniques, hyperparameter tuning through YAML configuration files, and rich model visualizations, providing a comprehensive solution for high-throughput analysis of biological images.
Read the paper: https://onlinelibrary.wiley.com/doi/10.1111/ele.14495
Functionality
>> Full list of available model architectures, losses, optimizers, schedulers, and augmentations <<
- Taxon-agnostic dataloaders (making it applicable to any dataset - not just biological ones)
- Support of timm models, and pytorch-optimizer
- Access to state-of-the-art metric losses, such as Supcon and Sub-center ArcFace.
- Exponential Moving Average for stable training, and Stochastic Moving Average for better generalization and performance.
- LRFinder for the second stage of the training.
- Easy customization of hyperparameters, including augmentations, through
YAMLconfigs (check the config-templates folder for examples) - Custom augmentations techniques via albumentations
- TensorBoard logs and checkpoints (soon to come: WandB integration)
- Streamlit app with rich model visualizations (e.g., Grad-CAM and timm-vis)
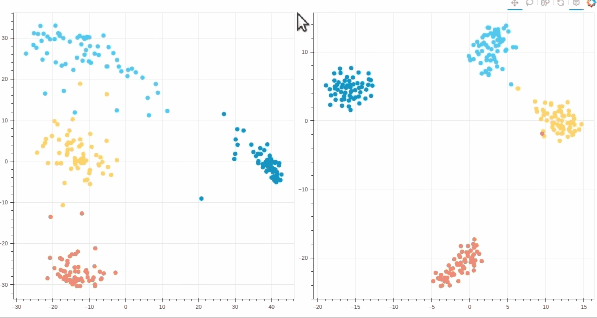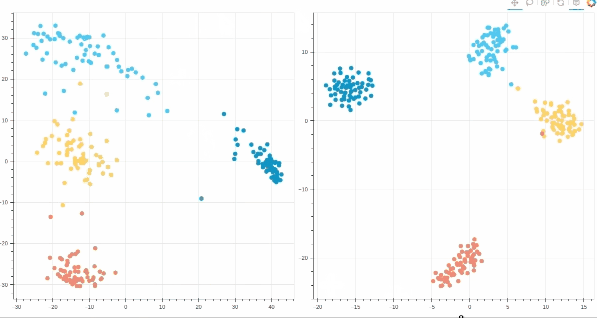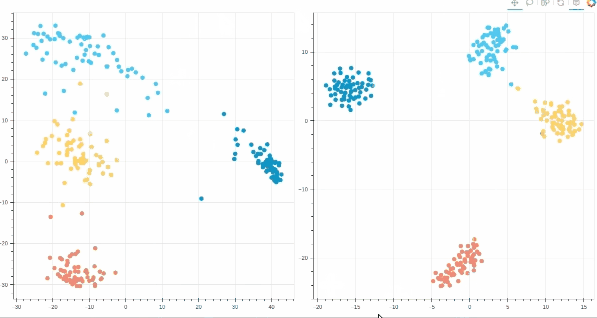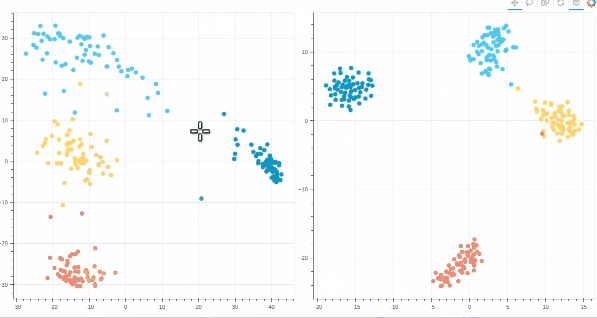
- Interactive t-SNE and PCA plots using Bokeh
Quickstart
>> Comprehensive help files <<
1. Install BioEncoder (into a virtual environment with pytorch/CUDA):
pip install bioencoder
2. Get the example dataset from the data repo and the config files by downloading the git repo, and extract both.
3. Start interactive session (e.g., in Spyder or VS code) and run the following commands one by one:
## use "overwrite=True to redo a step
import bioencoder
## global setup (pick a target directory for all output that bioencoder generates, e.g. training dataset, model weights, etc.)
bioencoder.configure(root_dir=r"bioencoder_wd", run_name="v1")
## split dataset (the dataset you downloaded)
bioencoder.split_dataset(image_dir=r"damselflies-aligned-trai_val", max_ratio=6, random_seed=42, val_percent=0.1, min_per_class=20)
## train stage 1
bioencoder.train(config_path=r"bioencoder_configs/train_stage1.yml")
bioencoder.swa(config_path=r"bioencoder_configs/swa_stage1.yml")
## explore embedding space and model from stage 1
bioencoder.interactive_plots(config_path=r"bioencoder_configs/plot_stage1.yml")
bioencoder.model_explorer(config_path=r"bioencoder_configs/explore_stage1.yml")
## (optional) learning rate finder for stage 2
bioencoder.lr_finder(config_path=r"bioencoder_configs/lr_finder.yml")
## train stage 2
bioencoder.train(config_path=r"bioencoder_configs/train_stage2.yml")
bioencoder.swa(config_path=r"bioencoder_configs/swa_stage2.yml")
## explore model from stage 2
bioencoder.model_explorer(config_path=r"bioencoder_configs/explore_stage2.yml")
## inference (stage 1 = embeddings, stage 2 = classification)
bioencoder.inference(config_path="bioencoder_configs/inference.yml", image="path/to/image.jpg" / np.array)
4. Alternatively, you can directly use the command line interface:
## use the flag "--overwrite" to redo a step
bioencoder_configure --root-dir "~/bioencoder_wd" --run-name v1
bioencoder_split_dataset --image-dir "damselflies-aligned-trai_val" --max-ratio 6 --random-seed 42
bioencoder_train --config-path "bioencoder_configs/train_stage1.yml"
bioencoder_swa --config-path "bioencoder_configs/swa_stage1.yml"
bioencoder_interactive_plots --config-path "bioencoder_configs/plot_stage1.yml"
bioencoder_model_explorer --config-path "bioencoder_configs/explore_stage1.yml"
bioencoder_lr_finder --config-path "bioencoder_configs/lr_finder.yml"
bioencoder_train --config-path "bioencoder_configs/train_stage2.yml"
bioencoder_swa --config-path "bioencoder_configs/swa_stage2.yml"
bioencoder_model_explorer --config-path "bioencoder_configs/explore_stage2.yml"
bioencoder_inference --config-path "bioencoder_configs/inference.yml" --path "path/to/image.jpg"
Citation
Please cite BioEncoder as follows:
@article{https://doi.org/10.1111/ele.14495,
author = {Lürig, Moritz D. and Di Martino, Emanuela and Porto, Arthur},
title = {BioEncoder: A metric learning toolkit for comparative organismal biology},
journal = {Ecology Letters},
volume = {27},
number = {8},
pages = {e14495},
keywords = {biodiversity, deep metric learning, feature space, machine learning, phenotypic differences, python package, species identification},
doi = {https://doi.org/10.1111/ele.14495},
url = {https://onlinelibrary.wiley.com/doi/abs/10.1111/ele.14495},
eprint = {https://onlinelibrary.wiley.com/doi/pdf/10.1111/ele.14495},
year = {2024}
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file bioencoder-1.0.2.tar.gz.
File metadata
- Download URL: bioencoder-1.0.2.tar.gz
- Upload date:
- Size: 43.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
02f4100de756c612cc8452943f7fd4bff485735d1aecdd4ea8a4c0720ec5b97a
|
|
| MD5 |
b6a618f6d659b6473ac476c03c6444ed
|
|
| BLAKE2b-256 |
f5fa7b5b9985993c09e92ef07e629004d9a53e55c2c225956c04d1c066c08f2a
|
File details
Details for the file bioencoder-1.0.2-py3-none-any.whl.
File metadata
- Download URL: bioencoder-1.0.2-py3-none-any.whl
- Upload date:
- Size: 53.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d1e5cc6c37cc4c0d5b76a795ff808256f116451a9ea5938625ee4264f91e5ce4
|
|
| MD5 |
71d284a27ba8ca527a5d9eeced2af9b3
|
|
| BLAKE2b-256 |
c7402dc828f6fe28bd527350a0a5acc3ff9e1e2ea6b58e895c4b2f0a29869c7f
|