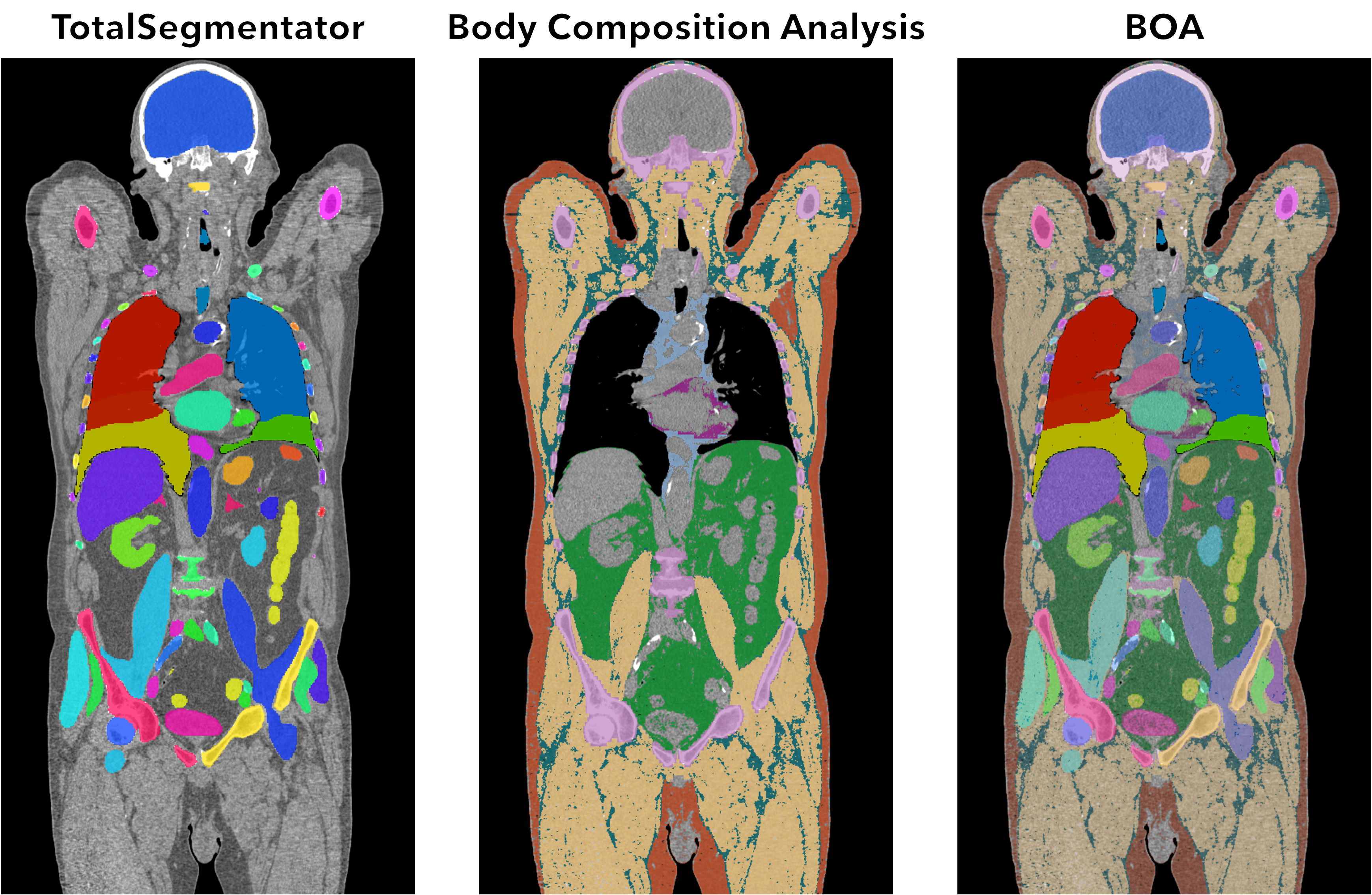
BOA is a tool for segmentation of CT scans developed by the SHIP-AI group at the Institute for Artificial Intelligence in Medicine (https://ship-ai.ikim.nrw/). Combining the TotalSegmentator and the Body Composition Analysis, this tool is capable of analyzing medical images and identifying the different structures within the human body, including bones, muscles, organs, and blood vessels.
Project description
BOA: Body and Organ Analysis
BOA is a tool for segmentation of CT scans developed by the SHIP.AI group at the Institute for Artificial Intelligence in Medicine (IKIM). Combining the TotalSegmentator and the Body Composition Analysis, this tool is capable of analyzing medical images and identifying the different structures within the human body, including bones, muscles, organs, and blood vessels.The tool also includes functionalities for the following tasks:
- Skeleton
- Organs
- Bone Mineral Density
- Contrast Media Recognition
- Cardiovascular System
- Body Parts
- Body Tissue Composition
- Body Region Detection
Example Segmentations
The BOA tool can be used to generate full body segmentations of CT scans:
Additionally, the generated segmentations can be used as input to generate realistic images using Siemens' Cinematic Rendering.
Citation
If you use this tool, please cite the following papers:
BOA:
Haubold, J., Baldini, G., Parmar, V., Schaarschmidt, B. M., Koitka, S., Kroll,
L., van Landeghem, N., Umutlu, L., Forsting, M., Nensa, F., & Hosch, R. (2023).
BOA: A CT-Based Body and Organ Analysis for Radiologists at the Point of Care.
Investigative radiology, 10.1097/RLI.0000000000001040. Advance online
publication. https://doi.org/10.1097/RLI.0000000000001040
Wasserthal J, Breit H-C, Meyer MT, et al. TotalSegmentator: Robust Segmentation
of 104 Anatomic Structures in CT Images. Radiol. Artif. Intell. 2023:e230024.
Available at: https://pubs.rsna.org/doi/10.1148/ryai.230024.
Isensee F, Jaeger PF, Kohl SAA, et al. nnU-Net: a self-configuring method for
deep learning-based biomedical image segmentation. Nat. Methods.
2021;18(2):203–211. Available at: https://www.nature.com/articles/s41592-020-01008-z.
How to run?
- Set up the environment variables.
- Either use the PACS integration or the command line tool.
Notes on Performance
To make an estimate on how much power and time is needed to process a study, we used the following table provided by the TotalSegmentator. However, for very large series (e.g. 1600 slices 1mm), the performance may be worse and more CPU power may be needed. According to our tests, 16GB of GPU should be sufficient.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file body_organ_analysis-0.1.5.tar.gz.
File metadata
- Download URL: body_organ_analysis-0.1.5.tar.gz
- Upload date:
- Size: 2.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.6.1 CPython/3.9.24 Linux/6.11.0-1018-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c5c671d732f0613ff8adf724d226c81b26e1e93d68b7c22581f05408f7a74cb7
|
|
| MD5 |
950c5addde82f9cd50ba35ef34dd16c2
|
|
| BLAKE2b-256 |
77a80756edfceafaf1315903580ff2ecd15dc7c1296a4f571a608d4fe68cd662
|
File details
Details for the file body_organ_analysis-0.1.5-py3-none-any.whl.
File metadata
- Download URL: body_organ_analysis-0.1.5-py3-none-any.whl
- Upload date:
- Size: 2.6 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.6.1 CPython/3.9.24 Linux/6.11.0-1018-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
95b193024a827621fea7c413e3f117fa54a5e6b7aba82e12942a02a4bc8df35b
|
|
| MD5 |
01ae7d74e63f000709cc05246bbff067
|
|
| BLAKE2b-256 |
32a9c5021242155cda8c4c63c4a438e6c92730ee1bce6a3f8daac37f68345cd3
|