Efficient and accurate virtual screening via docking-guided binding prediction with Boltz-2
Project description
Boltzina
Quick Start
Installation
# Using uv (recommended)
uv venv
uv sync
# Or using pip
pip install .
Tool setup (Vina, MAXIT, Boltz-2 model weights)
boltzina setup --all
For Uni-Dock2 (GPU-accelerated docking, requires pixi and CUDA 12):
# Clone Uni-Dock2 and build using the provided pixi.toml
git clone https://github.com/dptech-corp/Uni-Dock2 /path/to/Uni-Dock2
cp pixi.toml /path/to/Uni-Dock2/
cd /path/to/Uni-Dock2 && pixi install && pixi run build
boltzina setup --register-unidock2 /path/to/Uni-Dock2
Usage
With Boltz-2 structure prediction (sequence → dock → score)
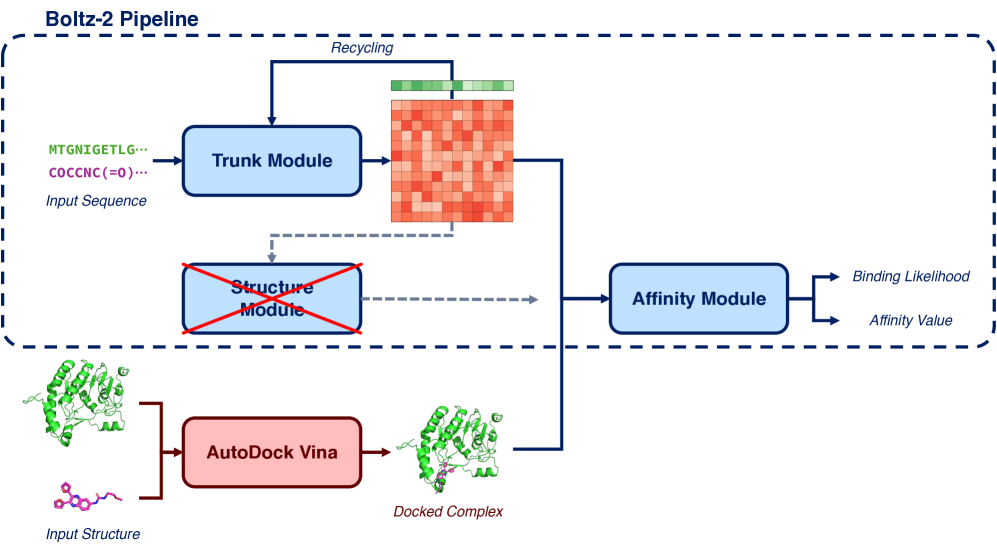
Provide a protein sequence and a SMILES/SDF file. Boltzina will:
- Run Boltz-2 structure + affinity prediction (complex with first/reference ligand)
- Determine the docking grid automatically from the predicted binding pose
- Run AutoDock Vina docking
- Score all poses with Boltz-2
# From a FASTA file (CDK2 example)
boltzina run sample/CDK2/ligands.smi \
--sequence-file sample/CDK2/cdk2.fasta \
--output-dir ./results
# From a sequence string directly
boltzina run sample/CDK2/ligands.smi \
--sequence "MENFQKVEKIGEGTYGVVYKARNKLTGEVVALKKIRLDTETEGVPSTAIREISLLKELNHPNIVKLLDVIHTENKLYLVFEFLHQDLKKFMDASALTGIPLPLIKSYLFQLLQGLAFCHSHRVLHRDLKPQNLLINTEGAIKLADFGLARAFGVPVRTYTHEVVTLWYRAPEILLGCKYYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVWPGVTSMPDYKPSFPKWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFFQDVTKPVPHLRL" \
--output-dir ./results
# Multi-chain protein: colon-separated sequences
boltzina run sample/CDK2/ligands.smi \
--sequence "MENFQKVEKIGEGTYGVVYK...:AKLSILPWGHC..." \
--output-dir ./results
# Multi-chain protein: multi-entry FASTA
boltzina run sample/CDK2/ligands.smi \
--sequence-file complex.fasta \ # >chain1 / seq / >chain2 / seq
--output-dir ./results
# Use a specific reference ligand for prediction and grid center
boltzina run sample/CDK2/ligands.smi \
--sequence-file sample/CDK2/cdk2.fasta \
--reference-ligand "CC(C)[C@H](CO)Nc1nc(Nc2ccc(C(=O)O)c(Cl)c2)c2ncn(C(C)C)c2n1" \
--output-dir ./results
# With more diffusion samples for better accuracy
boltzina run sample/CDK2/ligands.smi \
--sequence-file sample/CDK2/cdk2.fasta \
--use-msa-server \
--diffusion-samples 5 \
--output-dir ./results
With a Boltz-2 YAML input
For full control over multi-chain proteins, ligand definitions, and Boltz-2 settings,
use a boltz-compatible YAML file (see sample/CDK2/1ckp_cdk2.yaml for an example):
boltzina run sample/CDK2/ligands.smi \
--yaml sample/CDK2/1ckp_cdk2.yaml \
--output-dir ./results
The YAML format:
version: 1
sequences:
- protein:
id: A
sequence: MENFQKVEKIGEGTYGVVYK... # CDK2 sequence
- ligand:
id: B
smiles: 'CC(C)[C@H](CO)Nc1nc(Nc2ccc(C(=O)O)c(Cl)c2)c2ncn(C(C)C)c2n1'
properties:
- affinity:
binder: B
Multiple protein chains are supported (add more - protein: entries).
The properties.affinity.binder identifies the reference ligand for grid center determination.
From precomputed Boltz-2 results
If you have already run boltz predict, pass the output directory directly:
boltzina run sample/CDK2/ligands.smi \
--work-dir sample/CDK2/boltz_results_base \
--output-dir ./results
The grid center is determined automatically from the Boltz-2 predicted ligand position. You can override it explicitly:
boltzina run sample/CDK2/ligands.smi \
--work-dir sample/CDK2/boltz_results_base \
--grid-center "7.0,-4.9,7.5" \
--output-dir ./results
CLI Reference
boltzina run <INPUT> [OPTIONS]
INPUT can be a .smi/.txt file (SMILES list), .sdf file, or a directory.
Protein input (choose one; required):
| Option | Description |
|---|---|
--sequence / -s |
Protein sequence (single chain, or SEQ1:SEQ2 for multi-chain) |
--sequence-file |
FASTA file (one >entry per chain for multi-chain) |
--yaml |
Boltz-2 compatible YAML (protein + ligand + affinity) |
--work-dir |
Existing Boltz-2 output directory (docking + scoring only) |
Structure prediction options (with --sequence / --sequence-file):
| Option | Default | Description |
|---|---|---|
--reference-ligand |
first in INPUT | SMILES string or SDF file for Boltz-2 complex prediction and grid center |
Docking:
| Option | Default | Description |
|---|---|---|
--grid-center |
auto | Docking box center x,y,z |
--grid-size |
20.0 |
Docking box size (Å) |
--ligand-chain-id |
B |
Ligand chain in Boltz-2 prediction (rescore mode) |
--docking-engine |
vina |
vina or unidock2 |
--num-workers |
1 |
Parallel Vina workers |
--skip-docking |
off | Score existing poses only |
--regenerate-conformer |
off | Force 3D conformer regeneration for SDF |
Boltz-2 prediction:
| Option | Default | Description |
|---|---|---|
--use-msa-server |
off | Use online MMseqs2 MSA server |
--recycling-steps |
3 |
Boltz-2 recycling steps |
--sampling-steps |
200 |
Boltz-2 sampling steps |
--diffusion-samples |
1 |
Boltz-2 diffusion samples |
--use-potentials |
off | Boltz-2 inference-time potentials |
--subsample-msa |
off | Subsample MSA sequences |
--no-kernels |
off | Disable trifast kernels (older GPUs) |
--affinity-mw-correction |
off | MW correction to affinity |
Output:
| Option | Default | Description |
|---|---|---|
--output-dir / -o |
./boltzina_results |
Output directory |
--batch-size |
1 |
Boltz-2 scoring batch size |
--seed |
— | Random seed |
--vina-override |
off | Rerun Vina even if results exist |
--boltz-override |
off | Rerun Boltz-2 scoring even if results exist |
--keep-intermediate-files |
off | Keep intermediate docking files |
boltzina prepare <INPUT> [OPTIONS]
Convert SMILES/SDF to PDB + prepared_mols.pkl for use with run.py.
boltzina prepare ligands.smi --output-dir ./prepared
boltzina prepare ligands.sdf --output-dir ./prepared --regenerate-conformer
boltzina grid <STRUCTURE_FILE> [OPTIONS]
Compute the docking grid center from a ligand or complex file.
boltzina grid ligand.pdb --output vina_config.txt
boltzina grid complex.cif --chain B --output vina_config.txt
boltzina setup [OPTIONS]
Install and register external tools.
boltzina setup --all # Vina + MAXIT + Boltz-2 weights
boltzina setup --install-vina # Vina only
boltzina setup --install-maxit # MAXIT only
boltzina setup --register-unidock2 /path/to/Uni-Dock2
boltzina setup --show # Show current config
Legacy usage (run.py)
The original run.py interface is fully supported:
python run.py sample/CDK2/config.json
python run.py sample/CDK2/config.json --use_kernels --num_workers 4
See sample/CDK2/config.json for the configuration file format.
Benchmark Dataset
The MF-PCBA benchmark dataset used in the paper is included in mf-pcba_test.zip.
See the paper for details on the evaluation protocol.
Running Tests
# Unit tests (no GPU required)
uv run pytest tests/ --ignore=tests/test_integration.py -v
# Integration tests (requires GPU + Boltz-2 weights)
uv run pytest tests/test_integration.py -m gpu -v
Reference
Furui, K, & Ohue, M. Boltzina: Efficient and Accurate Virtual Screening via Docking-Guided Binding Prediction with Boltz-2. AI for Accelerated Materials Design - NeurIPS 2025. https://openreview.net/forum?id=OwtEQsd2hN
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file boltzina-1.0.0.tar.gz.
File metadata
- Download URL: boltzina-1.0.0.tar.gz
- Upload date:
- Size: 64.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f5b40f977eb62b5f02a92fba5bf9723da4ddaacd7825af97cdd0dd12626e3752
|
|
| MD5 |
b2121e76de932c12b9d68b38f8d59f29
|
|
| BLAKE2b-256 |
86413dc3c50317a83ce4958e37acb1ce3b7a7b238ed553f601f7eecafedb8427
|
Provenance
The following attestation bundles were made for boltzina-1.0.0.tar.gz:
Publisher:
release.yml on ohuelab/boltzina
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
boltzina-1.0.0.tar.gz -
Subject digest:
f5b40f977eb62b5f02a92fba5bf9723da4ddaacd7825af97cdd0dd12626e3752 - Sigstore transparency entry: 1351966429
- Sigstore integration time:
-
Permalink:
ohuelab/boltzina@cefaf78b48769a677c458ed08a0cdd95b6bb96e7 -
Branch / Tag:
refs/tags/v1.0.0 - Owner: https://github.com/ohuelab
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@cefaf78b48769a677c458ed08a0cdd95b6bb96e7 -
Trigger Event:
push
-
Statement type:
File details
Details for the file boltzina-1.0.0-py3-none-any.whl.
File metadata
- Download URL: boltzina-1.0.0-py3-none-any.whl
- Upload date:
- Size: 70.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8b3f6f4d933c8d59eed6ca9ba2b07fa2755b661093ccd47ff27c5b15e904df8e
|
|
| MD5 |
5a776ef56474a186559a2718302eb57c
|
|
| BLAKE2b-256 |
ef6267ede3ab7c68640290a55b5ca82d46ab02c892ad2f5bbde373647c194f12
|
Provenance
The following attestation bundles were made for boltzina-1.0.0-py3-none-any.whl:
Publisher:
release.yml on ohuelab/boltzina
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
boltzina-1.0.0-py3-none-any.whl -
Subject digest:
8b3f6f4d933c8d59eed6ca9ba2b07fa2755b661093ccd47ff27c5b15e904df8e - Sigstore transparency entry: 1351966505
- Sigstore integration time:
-
Permalink:
ohuelab/boltzina@cefaf78b48769a677c458ed08a0cdd95b6bb96e7 -
Branch / Tag:
refs/tags/v1.0.0 - Owner: https://github.com/ohuelab
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@cefaf78b48769a677c458ed08a0cdd95b6bb96e7 -
Trigger Event:
push
-
Statement type:










