BraTS algorithms
Project description
BraTS Orchestrator
______ _____ _____
| ___ \ |_ _/ ___|
| |_/ /_ __ __ _| | \ `--.
| ___ \ '__/ _` | | `--. \
| |_/ / | | (_| | | /\__/ /
\____/|_| \__,_\_/ \____/
_____ _ _ _
| _ | | | | | | |
| | | |_ __ ___| |__ ___ ___| |_ _ __ __ _| |_ ___ _ __
| | | | '__/ __| '_ \ / _ \/ __| __| '__/ _` | __/ _ \| '__|
\ \_/ / | | (__| | | | __/\__ \ |_| | | (_| | || (_) | |
\___/|_| \___|_| |_|\___||___/\__|_| \__,_|\__\___/|_|
Providing the top-performing algorithms from the Brain Tumor Segmentation (BraTS) challenges, through an easy-to-use Python API powered by Docker.
Features
- Access to top-performing algorithms from recent BraTS challenges
- Easy-to-use minimal API
- Extensive documentation and examples
Installation
With a Python 3.8+ environment, you can install BraTS orchestrator directly from PyPI:
# lightweight base package
pip install brats
# With preprocessing functionalities
pip install brats[preprocessing]
[!IMPORTANT]
To run BraTS orchestrator, you require a Docker installation.
Many algorithms also require GPU support (NVIDIA Docker).
In case you do not have access to a CUDA-capable GPU, the overview tables in the Available Algorithms and Usage section indicate which algorithms are CPU compatible.
Docker and NVIDIA Container Toolkit Setup
- Docker: Installation instructions on the official website
- NVIDIA Container Toolkit: Refer to the NVIDIA install guide and the official GitHub page
Singularity Support
BraTS orchestrator also supports Singularity as an alternative to Docker.
To enable Singularity, install it following the official guide and specify the backend to use as Backends.SINGULARITY when running the inference:
from brats.constants import Backends
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
output_file="path/to/segmentation.nii.gz",
backend=Backends.SINGULARITY
)
Available Algorithms and Usage
[!IMPORTANT] BraTS challenge algorithms require preprocessed brain images. See section Data preprocessing requirements
Segmentation Challenges

Note: Some legacy segmentation algorithms from BraTS challenges before 2023 are available via BraTS Toolkit.
Adult Glioma Segmentation (Pre & Post-Treatment)
Adult Glioma Segmentation on pre- and post-treatment brain MRI exams.
Usage example (code) and top 3 participants
from brats import AdultGliomaPreAndPostTreatmentSegmenter
from brats.constants import AdultGliomaPreAndPostTreatmentAlgorithms
segmenter = AdultGliomaPreAndPostTreatmentSegmenter(algorithm=AdultGliomaPreAndPostTreatmentAlgorithms.BraTS25_1, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation.nii.gz",
)
Note: If you're interested in Adult Glioma Segmentation, the BrainLes GlioMODA package may also be of interest.
Class: brats.AdultGliomaPreAndPostTreatmentSegmenter (Docs)
Challenge Paper 2023: Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Ishika Jain, et al. | N/A | ❌ | BraTS25_1 |
| 2025 | 2nd | Qu Lin, et al. | N/A | ✅ | BraTS25_2 |
| 2025 | 3rd | Liwei Jin, et al. | N/A | ✅ | BraTS25_3A |
| 2025 | 3rd | Adrian Celaya, et al. | N/A | ❌ | BraTS25_3B |
| 2024 | 1st | André Ferreira, et al. | Link | ❌ | BraTS24_1 |
| 2024 | 2nd | Heejong Kim, et al. | Link | ❌ | BraTS24_2 |
| 2024 | 3rd | Adrian Celaya | N/A | ✅ | BraTS24_3 |
Note: The MNI152 atlas, available on Zenodo, was employed for registration in the 2024 and subsequent BraTS Glioma Post-treatment Segmentation challenges.
Adult Glioma Segmentation (Pre-Treatment)
Adult Glioma Segmentation on pre-treatment brain MRI exams.
Usage example (code) and top 3 participants
from brats import AdultGliomaPreTreatmentSegmenter
from brats.constants import AdultGliomaPreTreatmentAlgorithms
segmenter = AdultGliomaPreTreatmentSegmenter(algorithm=AdultGliomaPreTreatmentAlgorithms.BraTS23_1, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation.nii.gz",
)
Note: If you're interested in Adult Glioma Segmentation, the BrainLes GlioMODA package may also be of interest.
Class: brats.AdultGliomaPreTreatmentSegmenter (Docs)
Challenge Paper 2023: Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2023 | 1st | André Ferreira, et al. | Link | ❌ | BraTS23_1 |
| 2023 | 2nd | Andriy Myronenko, et al. | Link | ❌ | BraTS23_2 |
| 2023 | 3rd | Fadillah Adamsyah Maani, et al. | Link | ❌ | BraTS23_3 |
Note: The SRI24 atlas, available on Zenodo, was employed for registration in the 2023 and prior BraTS Glioma Pre-Treatment Segmentation challenges.
BraTS-Africa Segmentation
Adult Glioma Segmentation on brain MRI exams in Sub-Sahara-Africa patient population.
Usage example (code) and top 3 participants
from brats import AfricaSegmenter
from brats.constants import AfricaAlgorithms
segmenter = AfricaSegmenter(algorithm=AfricaAlgorithms.BraTS25_1, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation.nii.gz",
)
Class: brats.AfricaSegmenter (Docs)
Challenge Paper 2023 Link
Challenge Paper 2024: N/A
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Claudia Takyi Ankomah, et al. | N/A | ✅ | BraTS25_1 |
| 2025 | 2nd | William Boonzaier, et al. | N/A | ❌ | BraTS25_2A |
| 2025 | 2nd | Mohtady Barakat, et al. | N/A | ❌ | BraTS25_2B |
| 2025 | 3rd | Ahmed Jaheen, et al. | N/A | ❌ | BraTS25_3 |
| 2024 | 1st | Abhijeet Parida, et al. | Link | ❌ | BraTS24_1 |
| 2024 | 2nd | Yanguang Zhao, et al. | Link | ✅ | BraTS24_2 |
| 2024 | 3rd | Sarim Hashmi, et al. | Link | ❌ | BraTS24_3 |
| 2023 | 1st | Andriy Myronenko, et al. | Link | ❌ | BraTS23_1 |
| 2023 | 2nd | Alyssa R Amod, et al. | Link | ❌ | BraTS23_2 |
| 2023 | 3rd | Ziyan Huang, et al. | Link | ✅ | BraTS23_3 |
Note: The SRI24 atlas, available on Zenodo, was employed for registration in BraTS Africa Segmentation challenges.
Meningioma Segmentation
Segmentation of Meningioma on brain MRI exams.
Usage example (code) and top 3 participants
from brats import MeningiomaSegmenter
from brats.constants import MeningiomaAlgorithms
### Example for 2023 algorithms
segmenter = MeningiomaSegmenter(algorithm=MeningiomaAlgorithms.BraTS25_1, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation_23.nii.gz",
)
Class: brats.MeningiomaSegmenter (Docs)
Challenge Paper 2023 Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Yu Haitao, et al. | N/A | ❌ | BraTS25_1 |
| 2025 | 2nd | Mohammad Mahdi Danesh Pajouh, et al. | N/A | ❌ | BraTS25_2 |
| 2023 | 1st | Andriy Myronenko, et al. | Link | ❌ | BraTS23_1 |
| 2023 | 2nd | Ziyan Huang, et al. | Link | ✅ | BraTS23_2 |
| 2023 | 3rd | Daniel Capellán-Martín et al. | Link | ❌ | BraTS23_3 |
Meningioma Radio Therapy Segmentation
Segmentation of Meningioma on T1C brain MRI exams.
Usage example (code) and top 3 participants
from brats import MeningiomaRTSegmenter
from brats.constants import MeningiomaRTAlgorithms
segmenter = MeningiomaRTSegmenter(algorithm=MeningiomaRTAlgorithms.BraTS25_1, cuda_devices="0")
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
output_file="segmentation_24.nii.gz",
)
Class: brats.MeningiomaRTSegmenter (Docs)
Challenge Paper 2024 Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Valeria Abramova, et al. | N/A | ❌ | BraTS25_1 |
| 2025 | 2nd | Sanskriti Srivastava, et al. | N/A | ✅ | BraTS25_2 |
| 2025 | 3rd | Nima Sadeghzadeh, et al. | N/A | ✅ | BraTS25_3 |
| 2024 | 1st | Valeria Abramova | N/A | ❌ | BraTS24_1 |
| 2024 | 2nd | Mehdi Astaraki | N/A | ❌ | BraTS24_2 |
| 2024 | 3rd | Andre Ferreira, et al. | Link | ✅ | BraTS24_3 |
Note: The MRI dataset in the Meningioma-Radiotherapy challenge was provided in native space. However, the SRI24 atlas, available on Zenodo, was employed for registration in BraTS Meningioma Pre-operative challenges.
Brain Metastases Segmentation
Segmentation on brain metastases on MRI exams for pre- and post-treatment cases.
Usage example (code) and top 3 participants
from brats import MetastasesSegmenter
from brats.constants import MetastasesAlgorithms
segmenter = MetastasesSegmenter(algorithm=MetastasesAlgorithms.BraTS25_1, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation.nii.gz",
)
Note: If you're interested in Brain Metastases Segmentation, the BrainLes AURORA package may also be of interest.
Class: brats.MetastasesSegmenter (Docs)
Challenge Paper 2023 Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Maria Bancerek, et al. | N/A | ❌ | BraTS25_1 |
| 2025 | 2nd | Wes Krikorian, et al. | N/A | ✅ | BraTS25_2 |
| 2023 | 1st | Andriy Myronenko, et al. | Link | ❌ | BraTS23_1 |
| 2023 | 2nd | Siwei Yang, et al. | Link | ❌ | BraTS23_2 |
| 2023 | 3rd | Ziyan Huang, et al. | Link | ✅ | BraTS23_3 |
Note: The SRI24 atlas, available on Zenodo, was employed for registration in BraTS Metastasis segmentation challenges.
Pediatric Segmentation
Segmentation of pediatric brain tumors on MRI exams.
Usage example (code) and top 3 participants
from brats import PediatricSegmenter
from brats.constants import PediatricAlgorithms
segmenter = PediatricSegmenter(algorithm=PediatricAlgorithms.BraTS25_1, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation.nii.gz",
)
Note: If you're interested in Pediatric Segmentation, the BrainLes PeTu package may also be of interest.
Class: brats.PediatricSegmenter (Docs)
Challenge Paper 2024 Link
Challenge Results Paper 2023 Link
Challenge Paper 2023 Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Yuxiao Yi, et al. | N/A | ❌ | BraTS25_1 |
| 2025 | 2nd | Meng-Yuan Chen, et al. | N/A | ❌ | BraTS25_2 |
| 2025 | 3rd | Haitao Yu, et al. | N/A | ❌ | BraTS25_3 |
| 2024 | 1st | Mehdi Astaraki | N/A | ❌ | BraTS24_1 |
| 2024 | 2nd | Tim Mulvany, et al. | Link | ❌ | BraTS24_2 |
| 2024 | 3rd | Sarim Hashmi, et al. | Link | ❌ | BraTS24_3 |
| 2023 | 1st | Daniel Capellán-Martín, et al. | Link | ❌ | BraTS23_1 |
| 2023 | 2nd | Andriy Myronenko, et al. | Link | ❌ | BraTS23_2 |
| 2023 | 3rd | Yubo Zhou | Link | ❌ | BraTS23_3 |
Note: The SRI24 atlas, available on Zenodo, was employed for registration in BraTS Pediatric Tumor Segmentation challenges.
Generalizability Across Tumors (BraTS-GoAT) Segmentation
Segmentation algorithm, adapting and generalizing to different brain tumors with segmentation labels of different tumor sub-regions.
Usage example (code) and top 3 participants
from brats import GoATSegmenter
from brats.constants import GoATAlgorithms
segmenter = GoATSegmenter(algorithm=GoATAlgorithms.BraTS25_1A, cuda_devices="0")
# these parameters are optional, by default the latest winning algorithm will be used on cuda:0
segmenter.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="segmentation.nii.gz",
)
Class: brats.PediatricSegmenter (Docs)
Challenge Paper 2024: N/A
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Meng-Yuan Chen, et al. | N/A | ❌ | BraTS25_1A |
| 2025 | 1st | To-Liang Hsu, et al. | N/A | ❌ | BraTS25_1B |
| 2025 | 1st | Vaidehi Satushe, et al. | N/A | ❌ | BraTS25_1C |
| 2025 | 1st | Simone Bendazzoli, et al. | N/A | ❌ | BraTS25_1D |
| 2024 | 1st | Frank Miao, Shengjie Niu | N/A | ❌ | BraTS24_1 |
Note: The datasets used in this challenge were adapted from other segmentation challenges, so the atlas type depends on the original dataset.
Inpainting Challenge
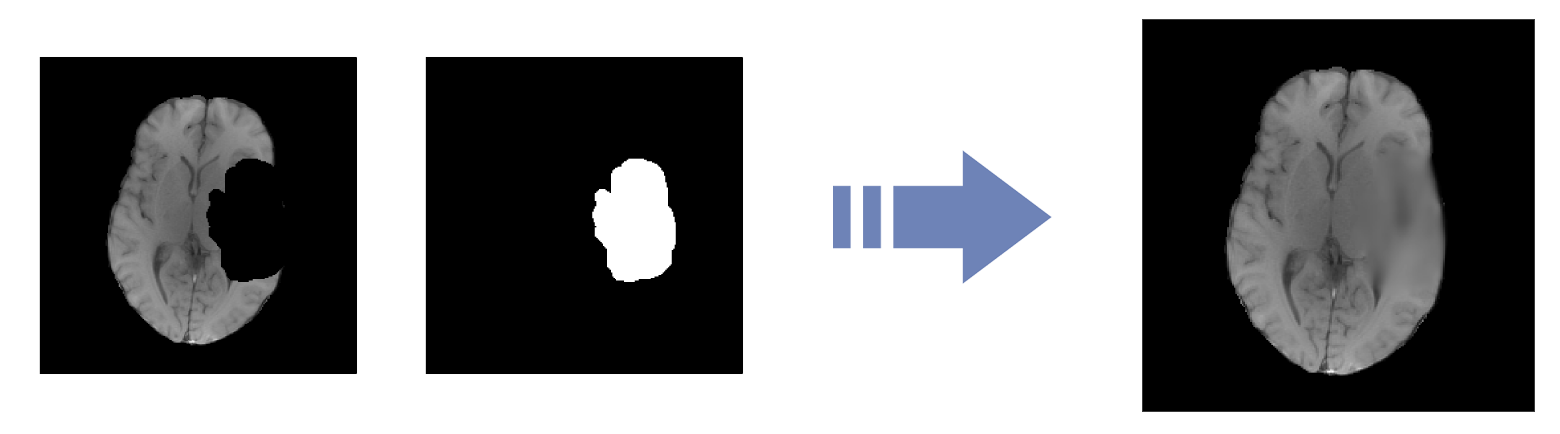
Algorithm to realistically synthesize and fill 3D healthy brain tissue in a region affected by glioma in brain MRI exams.
Usage example (code) and top 3 participants
from brats import Inpainter
from brats.constants import InpaintingAlgorithms
inpainter = Inpainter(algorithm=InpaintingAlgorithms.BraTS25_1A, cuda_devices="0")
inpainter.infer_single(
t1n="path/to/voided_t1n.nii.gz",
mask="path/to/mask.nii.gz",
output_file="inpainting.nii.gz",
)
Class: brats.Inpainter (Docs)
Challenge Paper 2023 and 2024 Link
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | Juexin Zhang, et al. | N/A | ❌ | BraTS25_1A |
| 2025 | 1st | André Ferreira, et al. | N/A | ❌ | BraTS25_1B |
| 2025 | 2nd | Juhyung Ha, et al. | N/A | ❌ | BraTS25_2 |
| 2024 | 1st | Juexin Zhang et al. | Link | ✅ | BraTS24_1 |
| 2024 | 2nd | André Ferreira, et al. | Link | ❌ | BraTS24_2 |
| 2024 | 3rd | Alicia Durrer, et al. | N/A | ❌ | BraTS24_3 |
| 2023 | 1st | Juexin Zhang, et al. | Link | ✅ | BraTS23_1 |
| 2023 | 2nd | Alicia Durrer, et al. | Link | ❌ | BraTS23_2 |
| 2023 | 3rd | Jiayu Huo, et al. | Link | ✅ | BraTS23_3 |
Note: The datasets used in this challenge were adapted from other segmentation challenges, so the atlas type depends on the original dataset.
Missing MRI Challenge
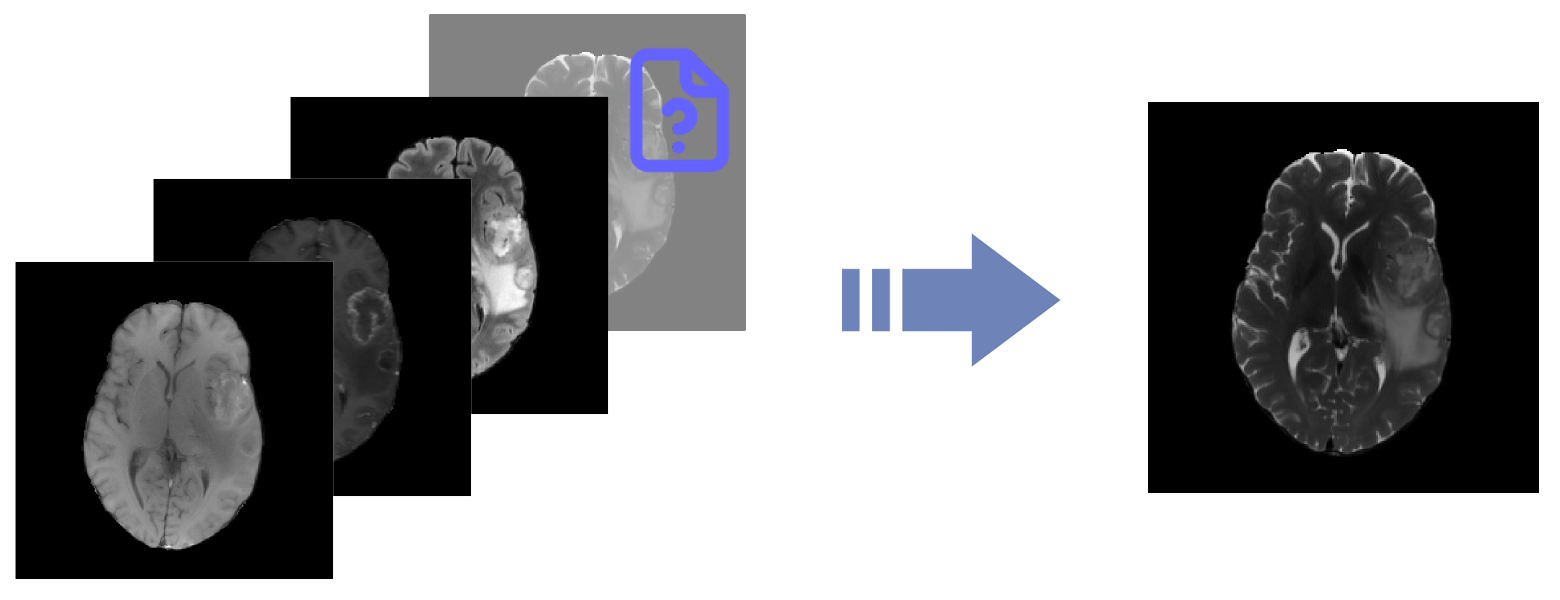

Algorithm to realistically synthesize missing MRI modalities from available sequences to enhance brain tumor segmentation.
Usage example (code) and top 3 participants
from brats import MissingMRI
from brats.constants import MissingMRIAlgorithms
missing_mri = MissingMRI(algorithm=MissingMRIAlgorithms.BraTS25_1, cuda_devices="0")
# Example to synthesize t2f modality (whichever modality is missing will be inferred)
missing_mri.infer_single(
t1c="path/to/t1c.nii.gz",
t1n="path/to/t1n.nii.gz",
# t2f="path/to/t2f.nii.gz",
t2w="path/to/t2w.nii.gz",
output_file="inferred_t2f.nii.gz",
)
Class: brats.MissingMRI (Docs)
Challenge Paper 2024: N/A
| Year | Rank | Author | Paper | CPU Support | Key Enum |
|---|---|---|---|---|---|
| 2025 | 1st | André Ferreira, et al. | N/A | ❌ | BraTS25_1 |
| 2025 | 2nd | Agustin Ujarky Cartaya Lathulerie, et al. | N/A | ❌ | BraTS25_2 |
| 2025 | 3rd | Lina Chator, et al. | N/A | ✅ | BraTS25_3 |
| 2024 | 1st | Jihoon Cho et al. | Link | ❌ | BraTS24_1 |
| 2024 | 2nd | Haowen Pang | N/A | ❌ | BraTS24_2 |
| 2024 | 3rd | Minjoo Lim et al. | Link | ❌ | BraTS24_3 |
| 2023 | 1st | Ivo Baltruschat | Link | ❌ | BraTS23_1 |
Note: The datasets used in this challenge were adapted from other segmentation challenges, so the atlas type depends on the original dataset.
[!TIP] For a full notebook example with more details, please check here:
Data Preprocessing Requirements
BraTS challenge algorithms require preprocessed brain scans.
This preprocessing typically includes co-registration, brain extraction, and registration to a challenge-specific brain atlas (template).
Please refer to each challenge’s documentation for details on which template to use.
For convenience, we provide challenge-specific wrappers around our preprocessing package, part of the BrainLesion Suite.
You can install the package with the preprocessing extra using:
pip install brats[preprocessing]
The modular architecture of the BraTS Orchestrator facilitates employing your own preprocessing routines.
Alternatively, the preprocessing package can be used independently to design custom preprocessing pipelines tailored to your specific needs.
Citation
[!IMPORTANT] If you use BraTS orchestrator in your research, please cite it to support the development!
Kofler, F., Rosier, M., Astaraki, M., Baid, U., Möller, H., Buchner, J. A., Steinbauer, F., Oswald, E., Rosa, E. de la, Ezhov, I., See, C. von, Kirschke, J., Schmick, A., Pati, S., Linardos, A., Pitarch, C., Adap, S., Rudie, J., Verdier, M. C. de, … Menze, B. (2025). BraTS orchestrator: Democratizing and Disseminating state-of-the-art brain tumor image analysis arXiv preprint arXiv:2506.13807
@misc{kofler2025bratsorchestratordemocratizing,
title={BraTS orchestrator : Democratizing and Disseminating state-of-the-art brain tumor image analysis},
author={Florian Kofler and Marcel Rosier and Mehdi Astaraki and Ujjwal Baid and Hendrik Möller and Josef A. Buchner and Felix Steinbauer and Eva Oswald and Ezequiel de la Rosa and Ivan Ezhov and Constantin von See and Jan Kirschke and Anton Schmick and Sarthak Pati and Akis Linardos and Carla Pitarch and Sanyukta Adap and Jeffrey Rudie and Maria Correia de Verdier and Rachit Saluja and Evan Calabrese and Dominic LaBella and Mariam Aboian and Ahmed W. Moawad and Nazanin Maleki and Udunna Anazodo and Maruf Adewole and Marius George Linguraru and Anahita Fathi Kazerooni and Zhifan Jiang and Gian Marco Conte and Hongwei Li and Juan Eugenio Iglesias and Spyridon Bakas and Benedikt Wiestler and Marie Piraud and Bjoern Menze},
year={2025},
eprint={2506.13807},
archivePrefix={arXiv},
primaryClass={eess.IV},
url={https://arxiv.org/abs/2506.13807},
}
Contributing
We welcome all kinds of contributions from the community!
Reporting Bugs, Feature Requests, and Questions
Please open a new issue here.
Code Contributions
Nice to have you on board! Please have a look at our CONTRIBUTING.md file.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file brats-0.1.8.tar.gz.
File metadata
- Download URL: brats-0.1.8.tar.gz
- Upload date:
- Size: 46.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
954a3f90c6adf5e98c59ce3dc6e9cd99bd15a512b379183c06a31c77386b53e0
|
|
| MD5 |
3cb9e508d8827c84f8e2e192d352005b
|
|
| BLAKE2b-256 |
ee5b8c9a408011ec85e82bd52a870664b48f20257811ac316fd3fb1fb73dcd2e
|
File details
Details for the file brats-0.1.8-py3-none-any.whl.
File metadata
- Download URL: brats-0.1.8-py3-none-any.whl
- Upload date:
- Size: 55.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
78dde085aab54f718b50c4033fed733922407bbca01c8a3e98247b828d5c68a4
|
|
| MD5 |
dc370828ea4fbd0a4fe6a4b911eecc9e
|
|
| BLAKE2b-256 |
6a13e086d08d75079639c155b640edf185dbadd680191c1e9ba51f5bf28aa372
|



















