BuDDI: Bulk Deconvolution with Domain Invariance to predict cell-type-specific perturbations from bulk
Project description
BuDDI
This repository contains all the model code used in BuDDI: Bulk Deconvolution with Domain Invariance to predict cell-type-specific perturbations from bulk
You can find notebooks that exemplify how to use the library under examples.
Overview
While single-cell experiments provide deep cellular resolution within a single sample, some are inherently more challenging than bulk experiments due to dissociation difficulties, cost, or limited tissue availability. This creates a situation where we have deep cellular profiles of one sample or condition, and bulk profiles across multiple samples and conditions.
A schematic of what BuDDi's methodological goal is shown below:
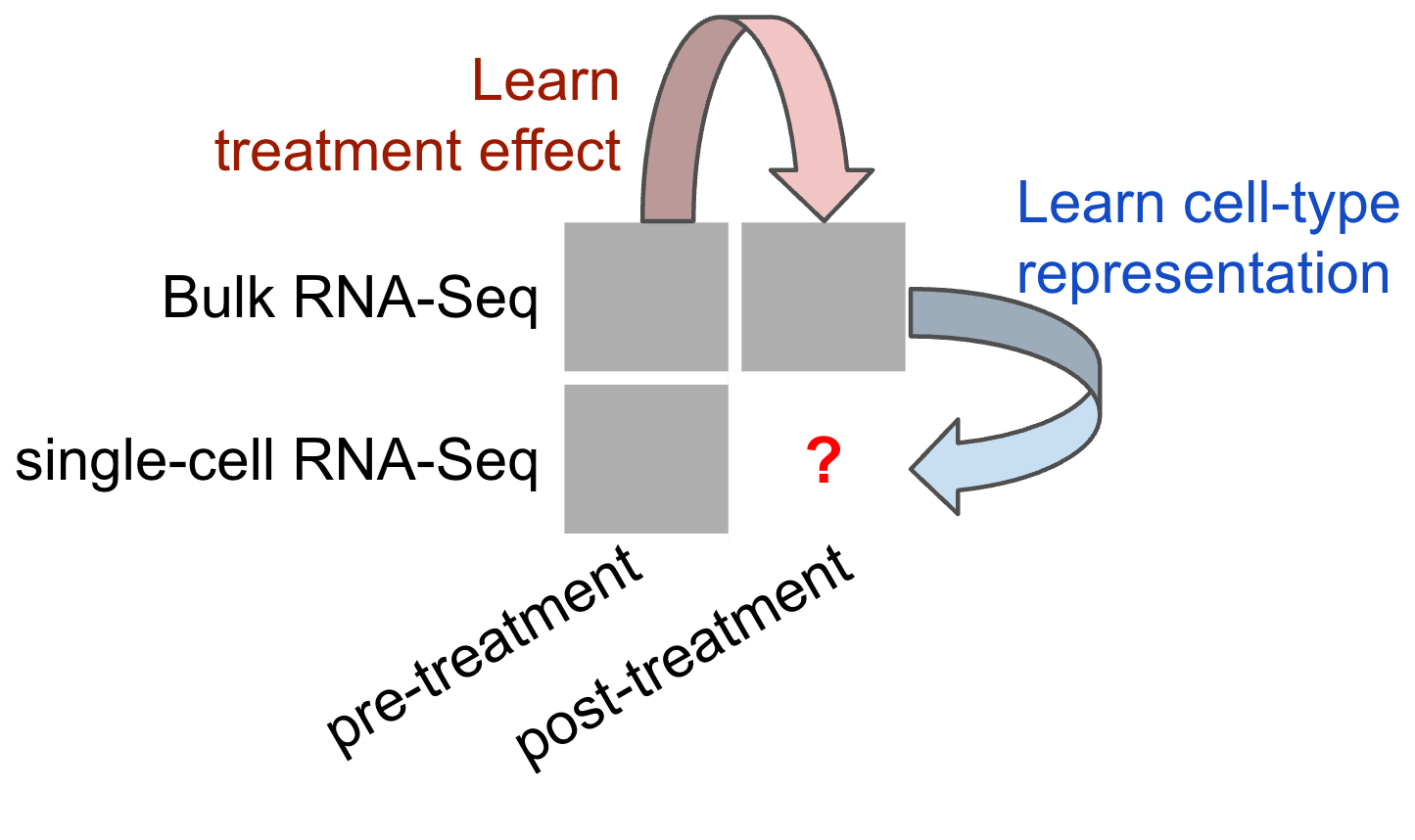
To bridge this gap, we propose BuDDI (BUlk Deconvolution with Domain Invariance).
BuDDI utilizes domain adaptation techniques to effectively integrate available corpora of case-control bulk and reference scRNA-seq observations to infer cell-type-specific perturbation effects. BuDDI achieves this by learning independent latent spaces within a single variational autoencoder (VAE) encompassing at least four sources of variability: 1) cell-type proportion, 2) perturbation effect, 3) structured experimental variability, and 4) remaining variability. Since each latent space is encouraged to be independent, we simulate perturbation responses by independently composing each latent space to simulate cell-type-specific perturbation responses.
Below is a schematic of the VAE structure of BuDDI:
Installation
First, you need to install dependencies using your package manager. Below is an example for OS X using brew.
brew install hdf5 c-blosc cython
export HDF5_DIR=/opt/homebrew/opt/hdf5
export BLOSC_DIR=/opt/homebrew/opt/c-blosc
Below is an example for Ubuntu using apt-get.
apt-get install libhdf5-serial-dev
apt-get install libblosc-dev
apt-get install pip install typing-extensions --upgrade
Second, you will install BuDDI from github using pip. It is recommended that you install this into a virtual env that you can activate before running the provided analysis scripts or your own BuDDI experiments.
pip install git+https://github.com/NDavidsonLab/buddi.git#egg=buddi[notebooks]
Using the Example Notebooks
Once you've installed BuDDI with the [notebooks] extra, you can run the example notebooks provided in the examples/ directory.
You can use the helper task buddi-lab, added via Poe the Poet, to start it.
poe buddi-lab
Alternatively, in the virtualenv in which you've installed BuDDI, you can create a kernel for the current venv and then start Jupyter Lab with the following command:
python -m ipykernel install --user --name buddi --display-name "Python (buddi)"
python -m jupyter lab examples/
Usage
See the tutorial for detailed instructions on how to use BuDDI.
To import BuDDI and its helper methods within your Python code, simply add the following
import buddi
from buddi.preprocessing import sc_preprocess
from buddi.plotting import validation_plotting as vp
Package Naming
The package is published as buddi, and the source tree now lives under src/buddi.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file buddi-0.1.0.tar.gz.
File metadata
- Download URL: buddi-0.1.0.tar.gz
- Upload date:
- Size: 57.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
54525bce1c52890ad32edb26489fb3d624b38685e43bd7c4cb1b50198191ce34
|
|
| MD5 |
ea0a3d3ca7a8af84f78d9b2e6f23f7a0
|
|
| BLAKE2b-256 |
87b3315ea83fb7a8e7890d0473f07a38584db89318371b8bc9ebec6e797838da
|
Provenance
The following attestation bundles were made for buddi-0.1.0.tar.gz:
Publisher:
pypi.yml on NDavidsonLab/buddi
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
buddi-0.1.0.tar.gz -
Subject digest:
54525bce1c52890ad32edb26489fb3d624b38685e43bd7c4cb1b50198191ce34 - Sigstore transparency entry: 1181666298
- Sigstore integration time:
-
Permalink:
NDavidsonLab/buddi@230979829f38f02a04996f24dd033a7fd7cd82c1 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/NDavidsonLab
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi.yml@230979829f38f02a04996f24dd033a7fd7cd82c1 -
Trigger Event:
release
-
Statement type:
File details
Details for the file buddi-0.1.0-py3-none-any.whl.
File metadata
- Download URL: buddi-0.1.0-py3-none-any.whl
- Upload date:
- Size: 63.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
49de7799b6ba889fd8cc0487569f401b39fe4dd81bef795e60d86fea0adba297
|
|
| MD5 |
e402a0328c0ad3bf83fa8cc8e8a56ae7
|
|
| BLAKE2b-256 |
0296aebf227f6adc2481a2d3961be478cb481829047dec9de3c3907de9d3e7e9
|
Provenance
The following attestation bundles were made for buddi-0.1.0-py3-none-any.whl:
Publisher:
pypi.yml on NDavidsonLab/buddi
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
buddi-0.1.0-py3-none-any.whl -
Subject digest:
49de7799b6ba889fd8cc0487569f401b39fe4dd81bef795e60d86fea0adba297 - Sigstore transparency entry: 1181666301
- Sigstore integration time:
-
Permalink:
NDavidsonLab/buddi@230979829f38f02a04996f24dd033a7fd7cd82c1 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/NDavidsonLab
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi.yml@230979829f38f02a04996f24dd033a7fd7cd82c1 -
Trigger Event:
release
-
Statement type:










