A user-friendly package for analyzing gene interactions and transcriptional bursting.
Project description

Brief introduction
BurstLink is a Python package to infer the coupled dynamics of gene regulatory interactions and transcriptional bursting from single-cell transcriptomics or multi-omics data. It contains many intuitive visualization and downstream analysis tools, providing a great practical toolbox for biomedical researchers.
The coupled dynamics betweem gene regulatory interactions and transcriptional bursting
Transcriptional bursts are inherently dynamic and stochastic processes, which are influenced by gene-gene regulatory interactions through transcription factors from other genes, ultimately driving highly heterogeneous gene expression within cells. A question is to explore how dynamic gene-gene interaction-constructed regulatory network dictates bursting on a genome-wide scale.
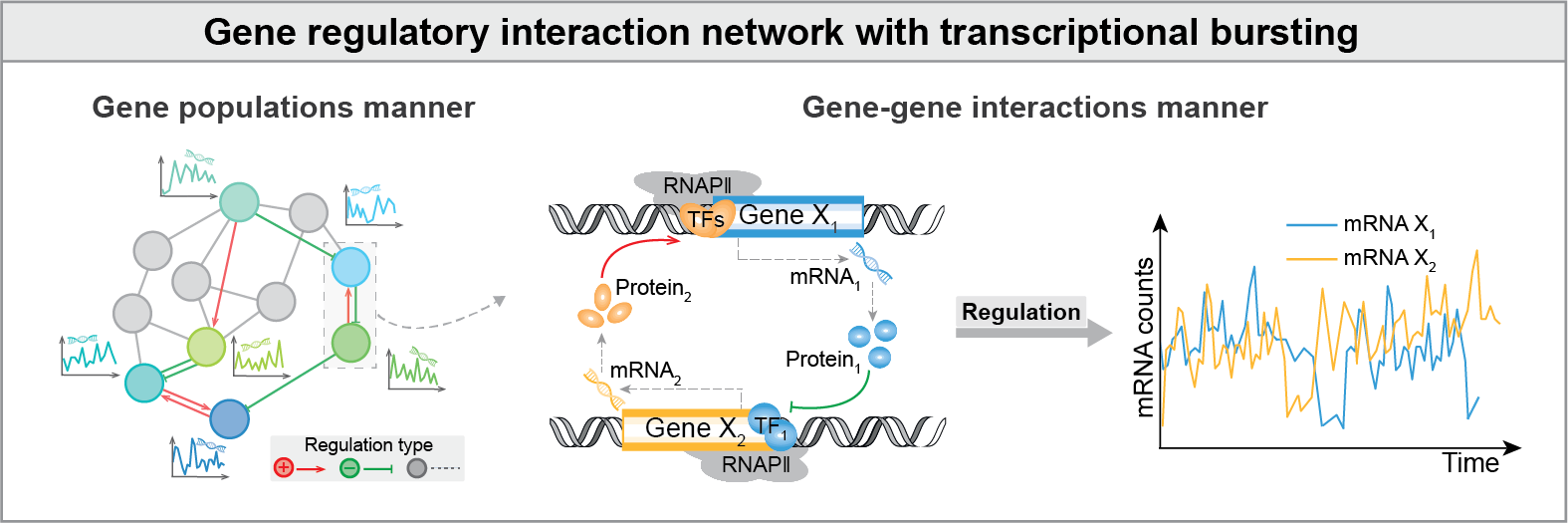
Inference workflow
BurstLink is a user-friendly package without the hyperparameter tuning, which is mainly utilized to infer coupled dynamics of gene regulatory interactions and transcriptional bursting, given scRNA-seq data of any two genes (screened by scATAC-seq data, optional). BurstLink enables gene-pair inference and supports rapid whole-genome inference and a series of downstream analyses.
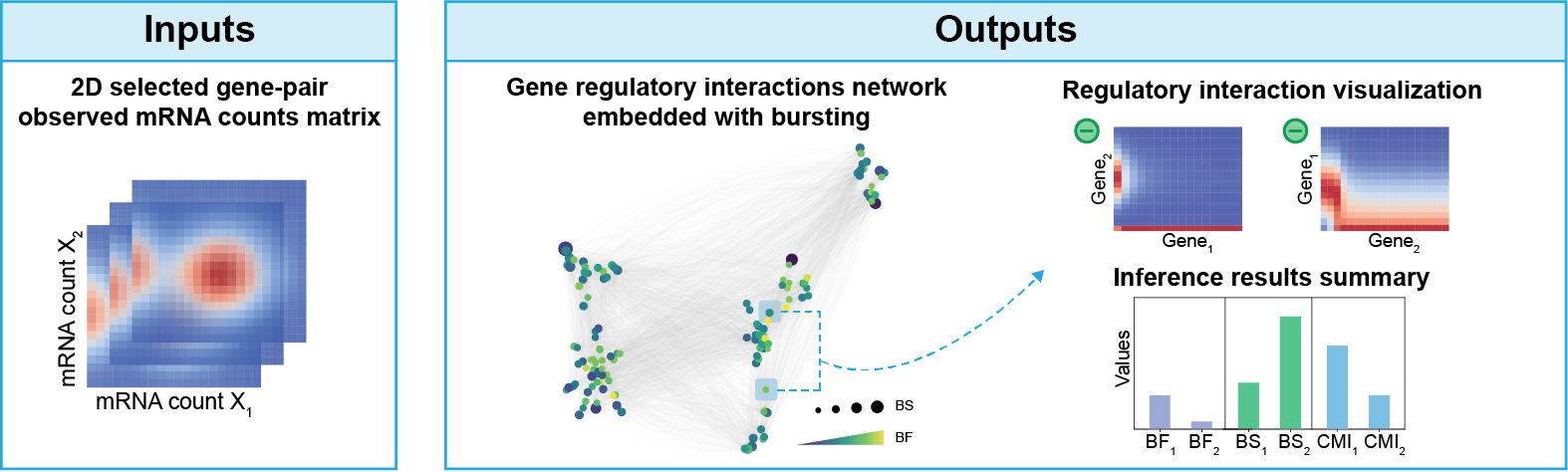
Installation
System requirements
Recommended operating systems: macOS or Linux. BurstLink was developed and tested on Linux and macOS.
Python requirements
BurstLink was developed using python 3.8.
Installation using pip
We suggest setting up BurstLink in a separate mamba or conda environment to prevent conflicts with other software dependencies. Create a new Python environment specifically for BurstLink and install the required libraries within it.
mamba create -n burstlink_env python=3.8 r-base=4.3.2
mamba activate burstlink_env
pip install burstlink
if you use conda, r-base=4.3.2 may not included in the channels. Instead, you can r-base=4.3.1 in conda.
Documentation, and Tutorials
For more realistic and simulation examples, please see BurstLink documentation that is available through the link https://burstlink.readthedocs.io/en/latest/.
Quick start
Let's get a quick start on using burstlink to infer gene regulatory interactions and transcriptional bursting by using simulation data generated from the genetic toggle switch dynamic model.
Import packages
import os, math
import numpy as np
import pandas as pd
import burstlink as bl
Setting work dictionary
To run the examples, you'll need to download the some pre-existing files in docs/tutorials/simulated_data folder and change your working directory to the simulated_data folder.
os.chdir("your_path/simulated_data")
scRNA-seq data from simulation
simul_data = np.asarray(pd.read_csv(os.path.abspath('data/simul_data_example1.csv')))[:, 1::]
You can also generate new simulation data with _synthetic_data :
params = [2, 2, 1, 1, 15, 15, 1, 0.2, 1, 3, 0.01, 2000]
S_stable = bl._utils._synthetic_data.SSA_coexpression(params, verbose = 'no_burst', fig = False)
simul_data = S_stable.astype(int)
Run the inference procedure
Import the scRNA-seq data and obatin the inference results, including (1) regulatory interactions: regulation types, regulation strengths, and regulation visualization; and (2) transcriptional bursting: burst frequencies, burst sizes, and expression variabilities.
params = [2, 2, 1, 1, 15, 15, 1, 0.2, 1, 3, 0.01, 2000]
geneinfo = np.array(['X1', 'X2', -math.copysign(1, params[9]), -math.copysign(1, params[8])]).reshape([1, 4])
infer_results = bl.tools._burst_interactions.genepair_inference(simul_data, geneinfo, figflag = 1, verbose1 = True, verbose2 = False, verbose3 = 2, test = False)

|

|

|
Reference
BurstLink: a statistical mechanistic model to reveal global transcriptional dynamics with gene regulatory interactions from single-cell data.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file burstlink-1.1.2.tar.gz.
File metadata
- Download URL: burstlink-1.1.2.tar.gz
- Upload date:
- Size: 29.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
546eceaeca18735bd90dacf49c30900210e7eafbbfbfe35096f8afd1db58c115
|
|
| MD5 |
60aa46f4e6f4e5e5724e7faab7398351
|
|
| BLAKE2b-256 |
098927cecc8b9ce50c1d1b35a91dd15a17b5c39571d3f1eb6aa4a4bf70d4f9a0
|
File details
Details for the file burstlink-1.1.2-py3-none-any.whl.
File metadata
- Download URL: burstlink-1.1.2-py3-none-any.whl
- Upload date:
- Size: 30.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8bb4181899852542556e8c6419431abfbb4f50a39bcb02c63937371c287c6de0
|
|
| MD5 |
ad2b37077752737d09a21cb294b55657
|
|
| BLAKE2b-256 |
8d79043b0afdd9f7adad1ed543222a680c7f084b89413a49dd8cc1108da4a5a4
|











