Python package for stochastic simulations
Project description
cayenne : Python package for stochastic simulations
Introduction
cayenne is a Python package for stochastic simulations. It offers a simple API to define models, perform stochastic simulations with them and visualize the results in a convenient manner.
Currently under active development in the develop branch.
Install
Install with pip:
$ pip install cayenne
Documentation
- General: https://cayenne.readthedocs.io.
- Benchmark repository, comparing
cayennewith other stochastic simulation packages: https://github.com/Heuro-labs/cayenne-benchmarks
Usage
A short summary follows, but a more detailed tutorial can be found here. You can define a model as a Python string (or a text file, see docs). The format of this string is loosely based on the excellent antimony library, which is used behind the scenes by cayenne.
from cayenne.simulation import Simulation
model_str = """
const compartment comp1;
comp1 = 1.0; # volume of compartment
r1: A => B; k1;
r2: B => C; k2;
k1 = 0.11;
k2 = 0.1;
chem_flag = false;
A = 100;
B = 0;
C = 0;
"""
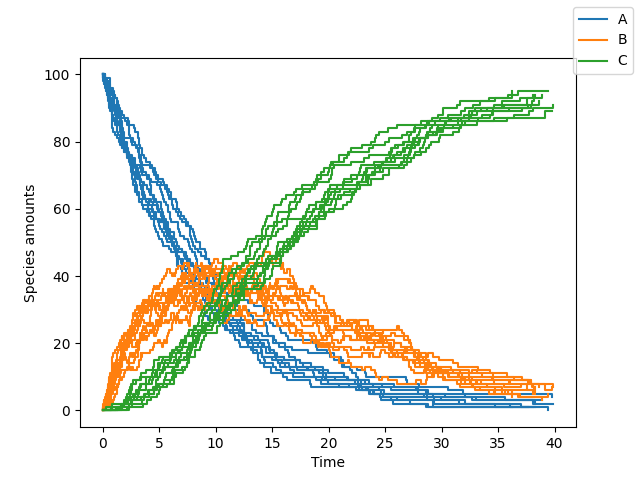
sim = Simulation.load_model(model_str, "ModelString")
# Run the simulation
sim.simulate(max_t=40, max_iter=1000, n_rep=10)
sim.plot()
Change simulation algorithm
You can change the algorithm used to perform the simulation by changing the algorithm parameter (one of "direct", "tau_leaping" or "tau_adaptive")
sim.simulate(max_t=150, max_iter=1000, n_rep=10, algorithm="tau_leaping")
Our benchmarks are summarized below, and show direct to be a good starting point. tau_leaping offers greater speed but needs specification and tuning of the tau hyperparameter. The tau_adaptive is less accurate and a work in progress.
Run simulations in parallel
You can run the simulations on multiple cores by specifying the n_procs parameter
sim.simulate(max_t=150, max_iter=1000, n_rep=10, n_procs=4)
Accessing simulation results
You can access all the results or the results for a specific list of species
# Get all the results
results = sim.results
# Get results only for one or more species
results.get_species(["A", "C"])
You can also access the final states of all the simulation runs by
# Get results at the simulation endpoints
final_times, final_states = results.final
Additionally, you can access the state a particular time point of interest $t$. cayenne will interpolate the value from nearby time points to give an accurate estimate.
# Get results at timepoint "t"
t = 10.0
states = results.get_state(t) # returns a list of numpy arrays
Benchmarks
| direct | tau_leaping | tau_adaptive | |
|---|---|---|---|
| cayenne | :heavy_check_mark: Most accurate yet | :heavy_check_mark: Very fast but may need manual tuning | Less accurate than GillespieSSA's version |
| Tellurium | :exclamation: Inaccurate for 2nd order | N/A | N/A |
| GillespieSSA | Very slow | :exclamation: Inaccurate for initial zero counts | :exclamation: Inaccurate for initial zero counts |
| BioSimulator.jl | :exclamation: Inaccurate interpolation | :exclamation: Inaccurate for initial zero counts | :exclamation: Inaccurate for initial zero counts |
License
Copyright (c) 2018-2020, Dileep Kishore, Srikiran Chandrasekaran. Released under: Apache Software License 2.0
Credits
- Cython
- antimony
- pytest
- Cookiecutter
- audreyr/cookiecutter-pypackage
- black
- Logo made with logomakr
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file cayenne-1.0.3.tar.gz.
File metadata
- Download URL: cayenne-1.0.3.tar.gz
- Upload date:
- Size: 1.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/49.2.0 requests-toolbelt/0.9.1 tqdm/4.48.0 CPython/3.6.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
897f2c6387f8ff09ae26a11128b8939e6350ff7f8794fbdf7f58015d1a176280
|
|
| MD5 |
e10bb05612e304b926a05af4b1cb74d4
|
|
| BLAKE2b-256 |
22538fdf90083da0f74c38287e380b5e2f17ed923259baf4d572de180848b5ed
|