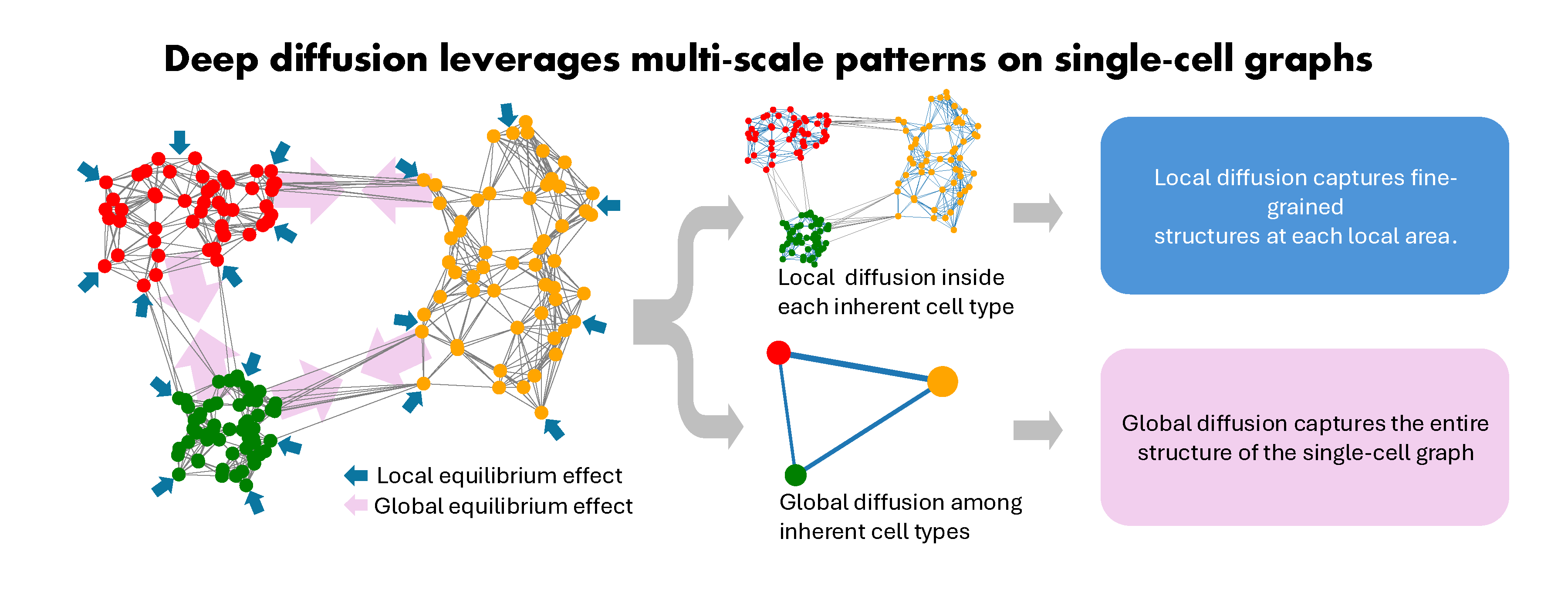
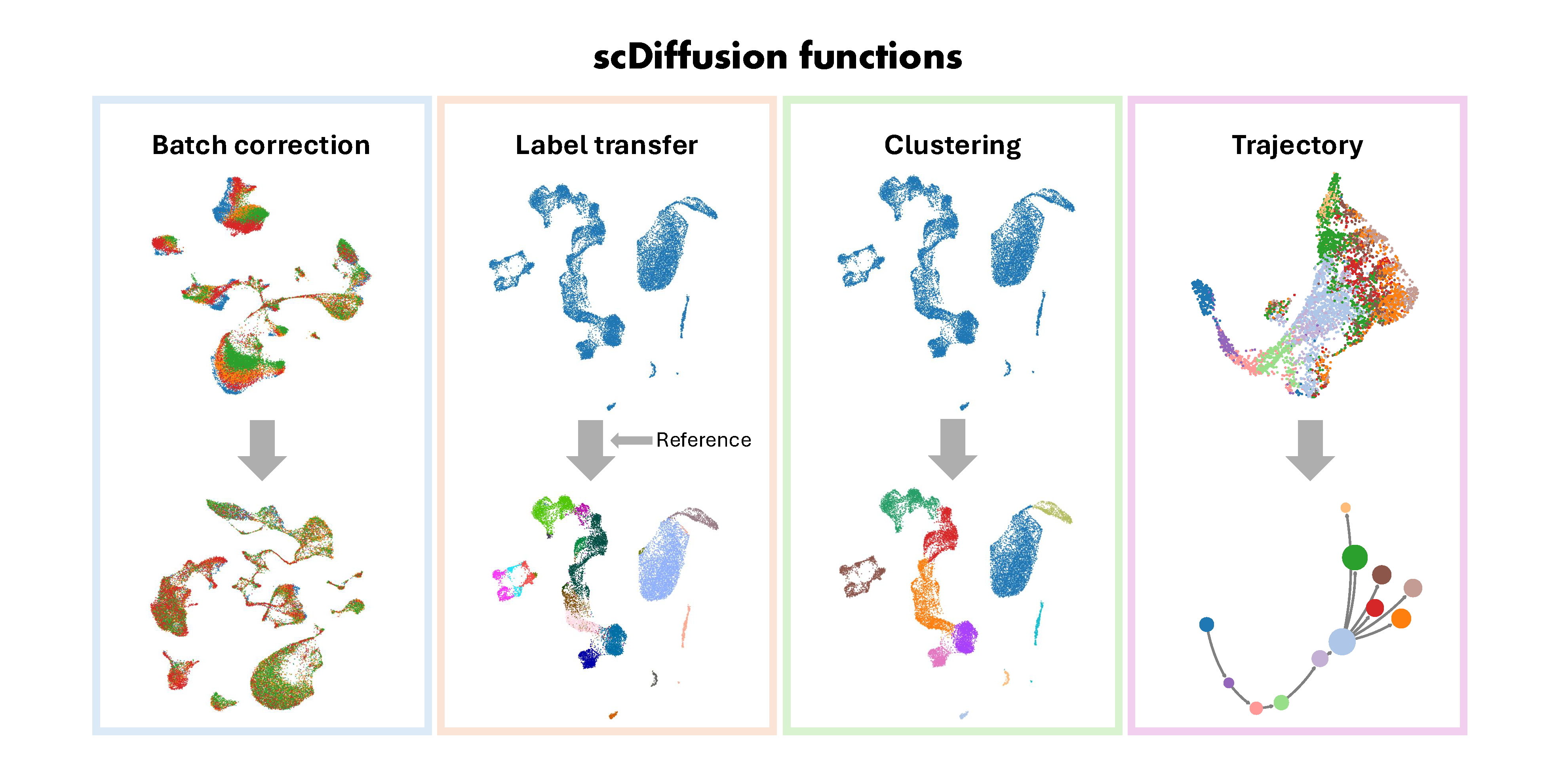
CellDiffusion(Single-Cell graph neural Diffusion) is a physics-informed graph generative model to do scRNA-seq analysis. CellDiffusion investigates cellular dynamics utilizing an attention-based neural network.
Project description
About:
CellDiffusion(Single-Cell graph neural Diffusion) is a deep diffusion model to leverage multi-scale patterns in single-cell graphs and enhance scRNA-seq analysis. Single-cell transcriptomics are typically analyzed based on gene expression within individual cells and hypothetic cell adjacencies. However, existing computational methods often suffer from a lack of leveraging and integrating multi-scale dependencies in feature space, undermining their effectiveness and robustness in downstream applications like handling of batch effects, cell type identification, and cell fate inference. To tackle this challenge, we introduce CellDiffusion to incorporate long-range information propagation among cells to uncover cellular biology from their transcriptomics. CellDiffusion integrates both local and global diffusion processes to comprehensively capture cell relationships, ranging from fine-grained structures to large-scale patterns. This approach exhibits great perception of inherent cell types and potential lineages and preserves cell identities in batch-imbalanced datasets. CellDiffusion enhances various downstream tasks, including data integration, reference-based cell type annotation, unsupervised clustering, and trajectory inference.
This repository contains the source code for the paper "CellDiffusion: graph-based deep diffusion model leverages multi-scale dependencies among single cells", Yu-Chen Liu, Lei Jiang, Simon Liang Lu, Anqi Zou, Zedong Lin, Nidhi Siddharam Loni, Heng Pan, Vijaya B. Kolachalama, Dong Xu*, Juexin Wang* & Chao Zhang*.
Installation:
CellDiffusion is available on PyPI. To install CellDiffusion, run the following command:
pip install celldiffusion
Or grab this source codes:
git clone https://github.com/CZCBLab/CellDiffusion.git
cd CellDiffusion
Python=3.9.9 is required. See other requirements in the file requirements.txt.
Run CellDiffusion in Docker:
git clone https://github.com/CZCBLab/CellDiffusion.git
cd CellDiffusion
# Build the Docker image
sudo docker build -t celldiffusion .
# Run Docker container with CPU
sudo docker run -it -p 8888:8888 --restart always celldiffusion bash
# Or run Docker container with GPU
sudo docker run -it -p 8888:8888 --restart always --gpus all celldiffusion bash
# Start Jupyter Notebook
jupyter notebook --ip="0.0.0.0" --allow-root
'celldiffusion' could be changed into your image name. Please refer to Docker and NVIDIA Container Toolkit for more details about Docker installation.
Tutorials:
For data integration, please check the notebook file "CellDiffusion_tutorial_Data_Integration.ipynb".
For reference-based cell type annotation, please check the notebook file "CellDiffusion_tutorial_Annotation_(Label_Transfer).ipynb".
For clustering tasks, please check the notebook file "CellDiffusion_tutorial_Clustering.ipynb".
For trajectory tasks, please check the notebook file "CellDiffusion_tutorial_Trajectory_Inference.ipynb".
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file celldiffusion-0.1.2.tar.gz.
File metadata
- Download URL: celldiffusion-0.1.2.tar.gz
- Upload date:
- Size: 33.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
edb9278f25a1fff4d93f30cea1d8c829b420fa3ab6c4569afed93a7b8d699c5f
|
|
| MD5 |
02a34fa7d6284565a995fa6b4385eb96
|
|
| BLAKE2b-256 |
f44e7491f63261efc592a4fde3cbb83b0aeea751b47f5dfb8dc2c4c975db5f3c
|
File details
Details for the file celldiffusion-0.1.2-py3-none-any.whl.
File metadata
- Download URL: celldiffusion-0.1.2-py3-none-any.whl
- Upload date:
- Size: 38.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e09307a34ecc71fd9d631ee02cc6c6ffc649ed96b35566500e7e22263ffc4f20
|
|
| MD5 |
7f6fdaedb7f3f1cf442674c4ca9eff25
|
|
| BLAKE2b-256 |
47f0459260069147b3cc3e4912f3ec6169115e4b0ef049a739dccc8dc2c5c731
|