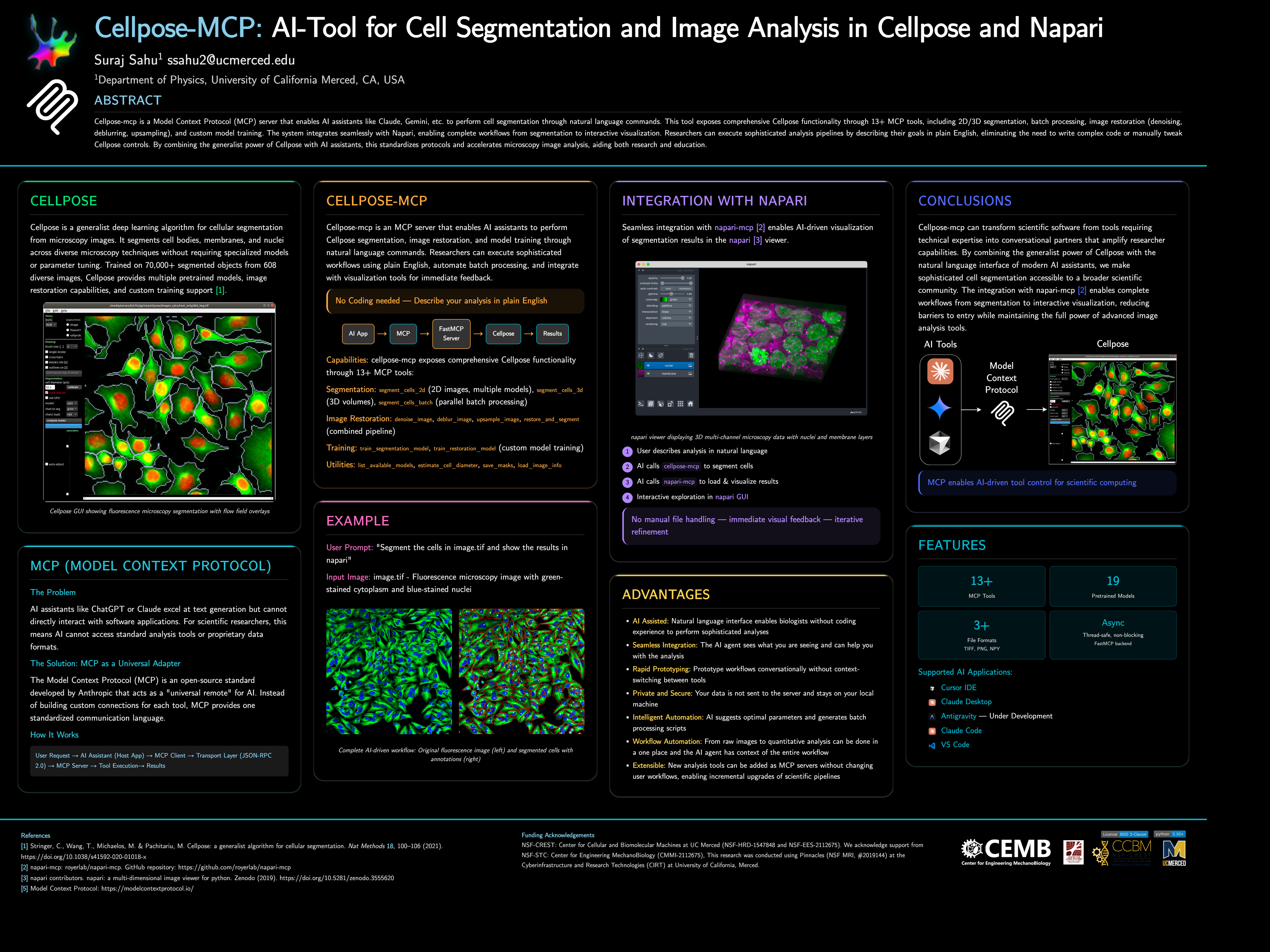
MCP server for AI-powered Cellpose cell segmentation via Model Context Protocol
Project description
Cellpose MCP Server
Cellpose-mcp is a Model Context Protocol (MCP) server that enables AI assistants like Claude, Cursor IDE, etc. to perform cell segmentation through natural language commands. This tool exposes comprehensive Cellpose functionality through 13+ MCP tools, including 2D/3D segmentation, batch processing, image restoration (denoising, deblurring, upsampling), and custom model training. The system integrates seamlessly with Napari, enabling complete workflows from segmentation to interactive visualization.
📌 Note: This project started as a fun project inspired by napari-mcp and adapted for Cellpose segmentation workflows. If you would like to contribute then please get in touch with me at ssahu2@ucmerced.edu.
🚀 Quick Start
pip install cellpose-mcp
OR for development:
git clone https://github.com/surajinacademia/cellpose_mcp.git
cd cellpose_mcp
pip install -e .
Auto-Configure Your AI Application
| Application | Installation Command | Notes |
|---|---|---|
| Cursor IDE | cellpose-mcp-install cursor |
Auto-configures MCP settings |
| Claude Desktop | cellpose-mcp-install claude-desktop |
Adds to Claude Desktop config |
| Antigravity | cellpose-mcp-install antigravity |
Configures Antigravity MCP |
| Claude Code | cellpose-mcp-install claude-code |
Or manually add .mcp.json to project root |
| VS Code | cellpose-mcp-install vscode |
Configures Cline/Roo Cline extension |
Manual Configuration for Claude Code
If you prefer manual setup, create a .mcp.json file in your project root:
{
"mcpServers": {
"cellpose": {
"command": "python",
"args": ["-m", "cellpose_mcp"],
"env": {
"KMP_DUPLICATE_LIB_OK": "TRUE",
"OMP_NUM_THREADS": "1"
}
}
}
}
After installation, restart your AI app and try asking:
"Can you list available Cellpose models?"
"Segment the cells in ./data/sample.tif using the cyto2 model"
🎯 What Can You Do?
Example: Cell Segmentation in Action
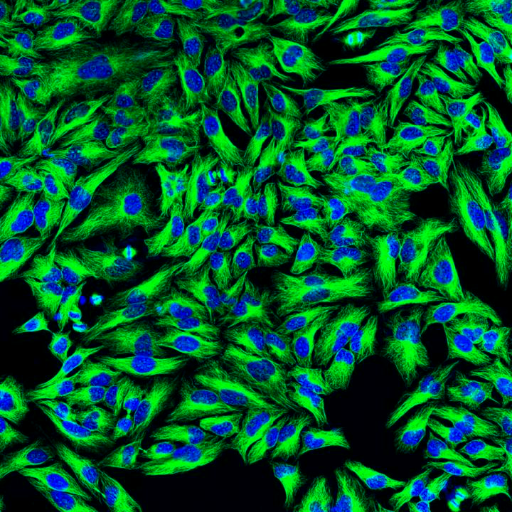
Original Image: Fluorescence microscopy with green-stained cytoplasm and blue-stained nuclei |
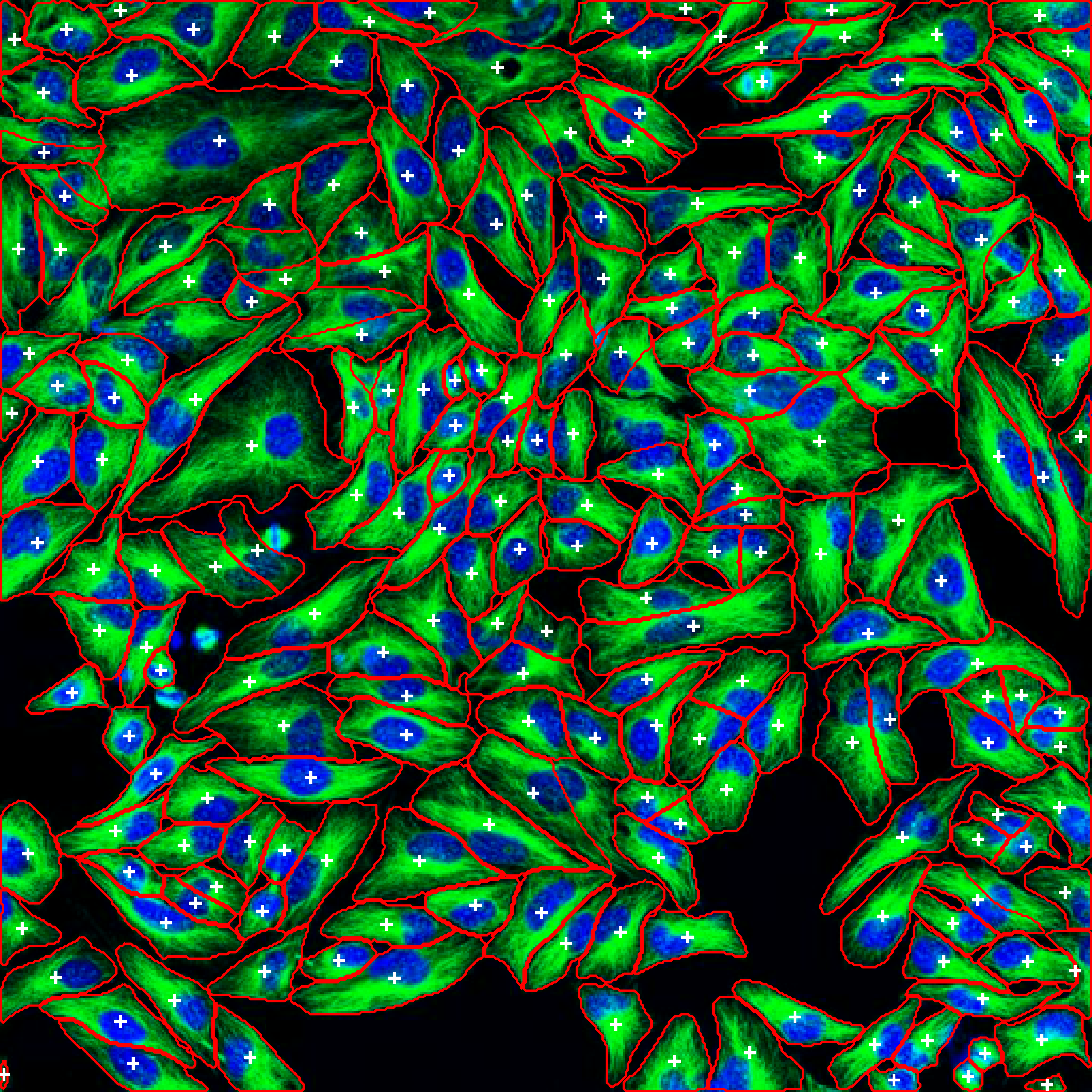
Segmented Result: Cells automatically detected with boundaries and labels |
Basic Cell Segmentation
"Segment the cells in ./data/sample.tif using the cyto2 model"
"List available Cellpose models"
"Estimate cell diameter from ./data/image.tif"
Advanced Workflows
"Segment all TIFF files in ./data/images/ and save masks to ./output/"
"Train a custom segmentation model using images in ./train/images/ and masks in ./train/masks/"
"Restore and segment the noisy image in ./data/noisy.tif using oneclick_cyto3"
Batch Processing
"Process all images in ./data/ with the cyto2 model and save results to ./output/"
🛠 Available MCP Tools
The server exposes 13+ tools for complete Cellpose functionality:
Segmentation Tools
segment_cells_2d- Segment cells in 2D imagessegment_cells_3d- Segment cells in 3D volumessegment_cells_batch- Batch process multiple images
Image Restoration Tools
denoise_image- Denoise microscopy imagesdeblur_image- Deblur microscopy imagesupsample_image- Upsample low-resolution imagesrestore_and_segment- Combined restoration + segmentation
Training Tools
train_segmentation_model- Train custom segmentation modeltrain_restoration_model- Train custom restoration model
Utility Tools
list_available_models- List all pretrained modelsestimate_cell_diameter- Estimate cell diameter from imagesave_masks- Save masks in various formatsload_image_info- Get image metadata
📖 Documentation
- Quick Start Guide - Get running in 3 steps
- Available Tools - Complete tool list
- Release Notes - Detailed v0.1.0 release information
- Changelog - Version history and changes
📋 Architecture
- FastMCP Server: Handles MCP protocol communication
- Cellpose Integration: Manages model loading and segmentation operations
- Tool Layer: Exposes Cellpose functionality as MCP tools
- File I/O: Handles image reading, writing, and mask generation
Key features:
- Thread-safe: All operations are properly serialized
- Non-blocking: Async operations for better performance
- Napari Integration: Integration with Napari for visualization and analysis
Author: Suraj Sahu
Affiliation: Department of Physics, University of California Merced, CA, USA
Email: ssahu2@ucmerced.edu
📄 License
BSD-3-Clause License - see LICENSE file for details.
🙏 Acknowledgments
- Napari MCP by royerlab
- Cellpose team for the excellent segmentation library
- FastMCP for the MCP framework
- Anthropic for Claude and MCP development
- Model Context Protocol - Open standard for AI-tool integration
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cellpose_mcp-0.1.1-py3-none-any.whl.
File metadata
- Download URL: cellpose_mcp-0.1.1-py3-none-any.whl
- Upload date:
- Size: 16.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
51e1fec31229408b3b41a437ce60a1173d11496fac30b2d11c7ed64b1b32eb7a
|
|
| MD5 |
79144cd501a0adce9aa44f03ff3f8962
|
|
| BLAKE2b-256 |
7e7dc803b1669a5b62a420d39ef7d2e31a26986ced693a718823128bb60d3a53
|