CLEAR: Calibrated Learning for Epistemic and Aleatoric Risk - Uncertainty quantification for regression
Project description

CLEAR: Calibrated Learning for Epistemic and Aleatoric Risk
Ilia Azizi1*, Juraj Bodik1,2*, Jakob Heiss2*, Bin Yu2,3
1Department of Operations, HEC, University of Lausanne, Switzerland 2Department of Statistics, University of California, Berkeley, USA 3Department of Electrical Engineering and Computer Sciences, University of California, Berkeley, USA *Equal contribution
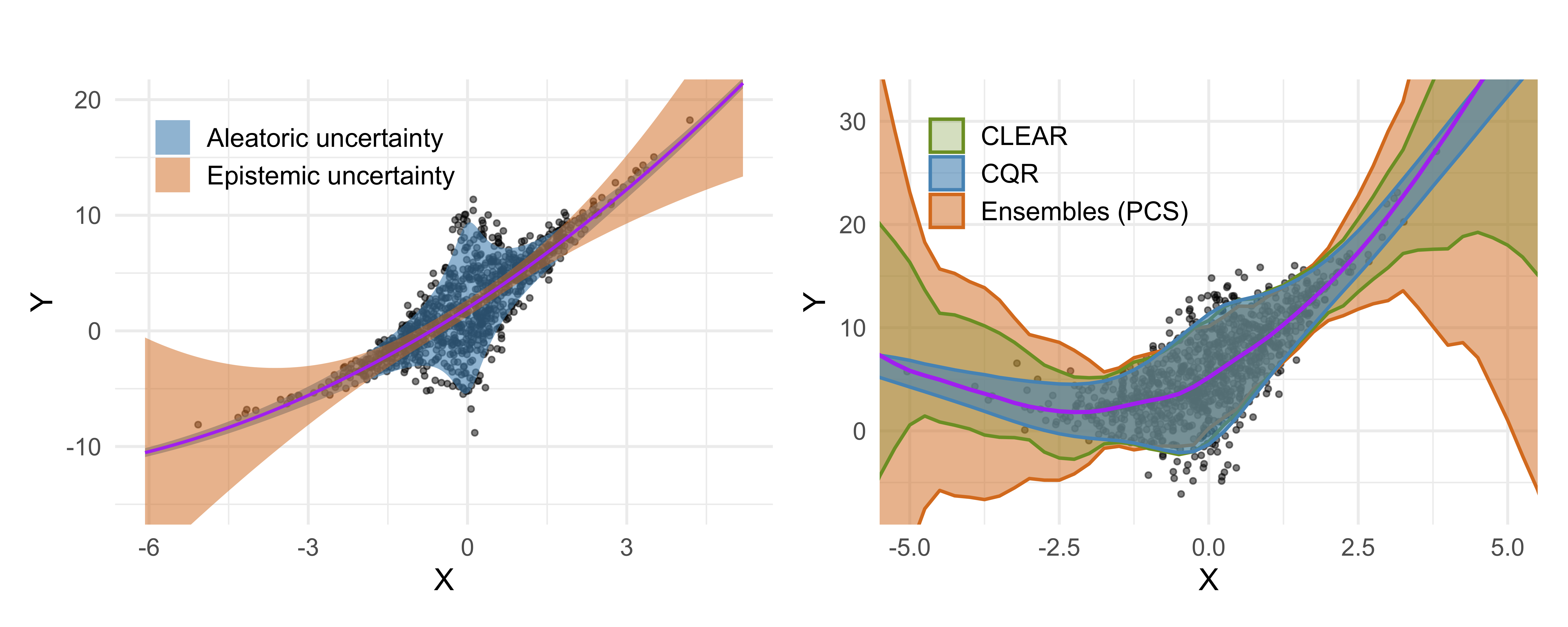
Abstract: Accurate uncertainty quantification is critical for reliable predictive modeling. Existing methods typically address either aleatoric uncertainty due to measurement noise or epistemic uncertainty resulting from limited data, but not both in a balanced manner. We propose CLEAR, a calibration method with two distinct parameters, $\gamma_1$ and $\gamma_2$, to combine the two uncertainty components and improve the conditional coverage of predictive intervals for regression tasks. CLEAR is compatible with any pair of aleatoric and epistemic estimators; we show how it can be used with (i) quantile regression for aleatoric uncertainty and (ii) ensembles drawn from the Predictability–Computability–Stability (PCS) framework for epistemic uncertainty. Across 17 diverse real-world datasets, CLEAR achieves an average improvement of 28.2% and 17.4% in the interval width compared to the two individually calibrated baselines while maintaining nominal coverage. Similar improvements are observed when applying CLEAR to Deep Ensembles (epistemic) and Simultaneous Quantile Regression (aleatoric). The benefits are especially evident in scenarios dominated by high aleatoric or epistemic uncertainty.
Installation
Quick Installation (PyPI)
Requirements: Python 3.11+
Install directly from PyPI:
pip install clear-uq
Local Development Installation
If you have the repository cloned, install in editable mode from the root directory:
pip install -e .
Or install from GitHub:
pip install git+https://github.com/Unco3892/clear.git
Development Setup
For development or to run all experiments with baseline comparisons (PCS_UQ, UACQR):
-
Ensure Python 3.11+ is installed on your system.
-
Create and activate a virtual environment using Conda:
conda create --name clear python=3.11 --yes conda activate clear
Note: If you encounter tkinter issues when running tests, run:
conda install -c conda-forge tk tcl --yes
-
Install the package (includes all dependencies and pytest):
pip install -e .
-
Run tests to verify everything works:
python -m pytest
Dependencies & Configuration:
- All dependencies are in
requirements.txtand automatically read bypyproject.toml - Dependencies include baseline comparison packages (
catboost,lightgbm) from PCS_UQ and UACQR - GPU optional installation: The default requirement installs the CPU-only build of PyTorch. If you need CUDA support (e.g., CUDA 12.6), edit
requirements.txtinside the activated Conda environment by commenting out the CPUtorchinstallation line and uncommenting the two lines directly beneath it (the--index-urloption and thetorch==2.6.0+cu126entry) before runningpip install -r requirements.txt.
Known Compatibility Notes:
pygamconflicts with latest scipy/numpy versions but CLEAR works fine with current versions- Baselines may have compatibility issues with numpy>=2.0.0 or quantile_forest>=1.4.0
- We maintain maximum compatibility where possible; please report any issues
Demo
For a high-level overview and practical demonstrations of CLEAR see demo.py & demo.ipynb which include comprehensive demonstrations on real datasets (Parkinsons, Airfoil).
Run the demo:
python demo.py
Minimal Code Snippet
Here's a quick overview of the CLEAR pipeline:
import numpy as np
from sklearn.datasets import make_regression
from sklearn.model_selection import train_test_split
from clear.clear import CLEAR
# 0. Generate Data (Minimal Example)
X, y = make_regression(n_samples=200, n_features=1, noise=10, random_state=42)
X_train, X_temp, y_train, y_temp = train_test_split(X, y, test_size=0.5, random_state=42)
X_calib, X_test, y_calib, y_test_actual = train_test_split(X_temp, y_temp, test_size=0.5, random_state=42)
# Some given external epistemic predictions
val_ep_median = X_calib[:,0] * np.mean(y_train/X_train[:,0])
val_ep_lower = val_ep_median - 2
val_ep_upper = val_ep_median + 2
test_ep_median = X_test[:,0] * np.mean(y_train/X_train[:,0])
test_ep_lower = test_ep_median - 2
test_ep_upper = test_ep_median + 2
# 1. Initialize CLEAR
clear_model = CLEAR(desired_coverage=0.95, n_bootstraps=10, random_state=777)
# 2. Fit CLEAR's Aleatoric Component (same can also be done internally for the epistemic model)
clear_model.fit_aleatoric(
X=X_train,
y=y_train,
quantile_model='rf',
fit_on_residuals=True,
epistemic_preds= (X_train[:,0] * np.mean(y_train/X_train[:,0])))
# 3. Get Aleatoric Predictions for Calibration Set
al_median_calib, al_lower_calib, al_upper_calib = clear_model.predict_aleatoric(X=X_calib, epistemic_preds=val_ep_median)
# 4. Calibrate CLEAR
clear_model.calibrate(
y_calib=y_calib,
median_epistemic=val_ep_median,
aleatoric_median=al_median_calib,
aleatoric_lower=al_lower_calib,
aleatoric_upper=al_upper_calib,
epistemic_lower=val_ep_lower,
epistemic_upper=val_ep_upper
)
print(f"Optimal Lambda: {clear_model.optimal_lambda:.3f}, Gamma: {clear_model.gamma:.3f}")
# 5. Predict with Calibrated CLEAR
clear_lower, clear_upper = clear_model.predict(
X=X_test,
external_epistemic={ 'median': test_ep_median, 'lower': test_ep_lower, 'upper': test_ep_upper }
)
print(f"Test CLEAR intervals (first 3): Lower={clear_lower[:3].round(2)}, Upper={clear_upper[:3].round(2)}")
Repo Structure
Below is a high-level overview of the important directories and files in this project:
├── data/ # Contains all datasets used for experiments and case studies.
├── models/ # Stores pre-trained models and outputs from model training runs.
├── PCS_UQ/ # Codebase and experiments for the PCS-UQ framework (referenced in paper).
├── UACQR/ # Codebase and experiments for the UACQR method (referenced in paper).
├── docs/figures/ # Directory for storing plots generated from experiment results.
│ ├── real/ # Plots related to experiments on real-world datasets.
│ └── simulations/ # Plots related to experiments on simulated datasets.
├── results/ # Stores raw and aggregated results from various experiments.
│ ├── case_study/ # Results specific to the Ames housing case study.
│ ├── standard/ # Results for standard (non-conformalized) model variants.
│ │ ├── qPCS_all_10seeds_all/ # Variant (a) against the baselines.
│ │ ├── qPCS_qxgb_10seeds_qxgb/ # Variant (b) against the baselines.
│ │ └── PCS_all_10seeds_qrf/ # Variant (c) against the baselines.
│ ├── conformalized/ # Results for conformalized model variants (same sub-structure as standard/).
│ └── de_sqr/ # Results for Deep Ensemble + SQR experiments.
├── src/ # Main source code for the CLEAR.
│ ├── case_study/ # Code for the Ames housing case study, including data preparation and analysis.
│ ├── clear/ # Core implementation of the CLEAR algorithm and associated utilities.
│ │ ├── clear.py # Implements the main CLEAR calibration logic.
│ │ ├── metrics.py # Defines metrics for evaluating prediction intervals.
│ │ └── utils.py # Contains utility functions used across the CLEAR module.
│ │ ├── models/ # Contains the general models used in the experiments
│ ├── experiments/ # Scripts to run download/process data, run benchmarks, and manage experiments.
│ │ ├── download_process_real_data.py # Downloads and preprocesses real-world datasets (has already been prvoided in the data/ directory).
│ │ ├── benchmark_real_data.py # Runs benchmark experiments on real-world datasets with PCS and CQR components.
│ │ ├── benchmark_real_data_de_sqr.py # Runs benchmark experiments on real-world datasets with DE and SQR components.
│ │ ├── benchmark_uacqr.py # Runs benchmark experiments for the UACQR method.
│ │ ├── benchmark_simulations.py # Runs benchmark experiments on simulated datasets.
│ │ └── utils.py # Utility functions for experiment scripts.
│ ├── pcs/ # Utility scripts related to training and using PCS models that are used in the experiments as well.
│ └── tests/ # Contains tests for the project's codebase.
├── demo.py # A demonstration script showcasing the CLEAR methodology.
├── demo.ipynb # A Jupyter Notebook demonstration of the CLEAR methodology (generated from demo.py).
├── README.md # This file: Project overview, setup instructions, and repository structure.
├── pytest.ini # Configuration file for pytest.
├── requirements.txt # Lists Python packages and versions required to run the project.
└── .gitignore # Specifies intentionally untracked files that Git should ignore.
Reproducibility
To reproduce the experiments from the paper:
-
Prepare Environment:
conda create --name clear python=3.11 --yes conda activate clear pip install clear-uq
-
Download Pre-trained Models (optional for running benchmarks):
python download_data.py # Uses rclone for faster, resumable downloads
-
Run Benchmark Experiments:
-
The core scripts for running experiments are located in the
src/experiments/directory. -
Optional: We provide the trained PCS ensemble models in one of the sections below, however, you can also retrain the PCS quantile and mean predictor models in
src/pcs/using thetrain_pcs_quantile.pyandtrain_pcs_mean.pyscripts.cd src/pcs python train_pcs_quantile.py python train_pcs_mean.py
-
To run benchmarks on real-world datasets, navigate to
src/experiments/and execute:python benchmark_real_data.pycd src/experiments # Standard models (variants a → b → c) python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 25 --global_log --approach both --models_dir ../../models/pcs_top1_qpcs_10_standard --csv_results_dir ../../results/standard/qPCS_all_10seeds_all python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_qxgb_10_standard --csv_results_dir ../../results/standard/qPCS_qxgb_10seeds_qxgb python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_pcs_10_standard --csv_results_dir ../../results/standard/PCS_all_10seeds_qrf # Conformalized models (variants a → b → c) python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 25 --global_log --approach both --models_dir ../../models/pcs_top1_qpcs_10_conformalized --csv_results_dir ../../results/conformalized/qPCS_all_10seeds_all python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_qxgb_10_conformalized --csv_results_dir ../../results/conformalized/qPCS_qxgb_10seeds_qxgb python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_pcs_10_conformalized --csv_results_dir ../../results/conformalized/PCS_all_10seeds_qrf
Note: The GAM in variant (a) may not converge for
data_naval_propulsiondue to apygambug: https://github.com/dswah/pyGAM/issues/357 -
To run benchmarks on simulated datasets, navigate to
src/experiments/and execute:python benchmark_simulations.pycd src/experiments # Homoscedastic noise with sigma=1, d=1, 100 simulations python benchmark_simulations.py --d 1 --num_simulations 100 --noise_type homo --use_external_pcs # Heteroscedastic noise with sigma=1+|x|, d=1, 100 simulations python benchmark_simulations.py --d 1 --num_simulations 100 --noise_type hetero1 --use_external_pcs # Heteroscedastic noise with sigma=1+1/(1+x^2), d=1, 100 simulations python benchmark_simulations.py --d 1 --num_simulations 100 --noise_type hetero2 --use_external_pcs # Multivariate: Homoscedastic noise with sigma=1, d=1, 100 simulations python benchmark_simulations.py --randomize_d --num_simulations 100 --noise_type homo --use_external_pcs
-
To run benchmarks specifically for the UACQR comparison (Table 1 and Appendix tables), navigate to
src/experiments/and execute both runs:cd src/experiments # Standard CLEAR variant (c) vs UACQR (Table 1) python benchmark_uacqr.py --data_dir ../../models/pcs_top1_pcs_10_standard --clear_results_dir ../../results/standard/PCS_all_10seeds_qrf --output_csv ../../results/uacqr/uacqr_benchmark_results_standard.csv # Conformalized CLEAR variant (c) vs UACQR (Appendix tables) python benchmark_uacqr.py --data_dir ../../models/pcs_top1_pcs_10_conformalized --clear_results_dir ../../results/conformalized/PCS_all_10seeds_qrf --output_csv ../../results/uacqr/uacqr_benchmark_results_conformalized.csv
-
To run the Deep Ensemble + SQR variant described in
benchmark_real_data_de_sqr.py, use:cd src/experiments # Standard DE+SQR models (auto-detect datasets in qxgb_10_standard) python benchmark_real_data_de_sqr.py --coverage 0.95 --models_dir ../../models/pcs_top1_qxgb_10_standard --output_dir ../../results/de_sqr --seed 42 --batch_size 64 --ensemble_epochs 1500 --sqr_lr 5e-4
The script automatically falls back to CPU mode when CUDA is unavailable; expect longer runtimes without a GPU. Omit
--max_runsto process every stored run (paper setting), and adjust--ensemble_epochs,--batch_size, or enable--fast_modeif you need faster turnaround. -
For faster overall runtime, the real-data benchmarks can be chained in order (variant b → c → a):
cd src/experiments # Fastest order one-liners: standard (variant b → c → a) python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 25 --global_log --approach both --models_dir ../../models/pcs_top1_qxgb_10_standard --csv_results_dir ../../results/standard/qPCS_qxgb_10seeds_qxgb ; python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_pcs_10_standard --csv_results_dir ../../results/standard/PCS_all_10seeds_qrf ; python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_qpcs_10_standard --csv_results_dir ../../results/standard/qPCS_all_10seeds_all # Fastest order one-liners: conformalized (variant b → c → a) python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 25 --global_log --approach both --models_dir ../../models/pcs_top1_qxgb_10_conformalized --csv_results_dir ../../results/conformalized/qPCS_qxgb_10seeds_qxgb ; python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_pcs_10_conformalized --csv_results_dir ../../results/conformalized/PCS_all_10seeds_qrf ; python benchmark_real_data.py --coverage 0.95 --generate_tables --n_jobs 30 --global_log --approach both --models_dir ../../models/pcs_top1_qpcs_10_conformalized --csv_results_dir ../../results/conformalized/qPCS_all_10seeds_all
-
Consult the respective scripts for any command-line arguments or configurations you might want to adjust (e.g., specific datasets, model parameters, number of seeds). Most scripts support
argparseto adjust parameters.
-
-
Results: Raw results are saved to
results/and plots todocs/figures/ -
Case Study:
The Ames housing case study can be run by navigating to
src/case_study/and using the script:python ames_clear_case_study.pypython ames_clear_case_study.py
The experiments were conducted on a machine with the following specifications:
- Operating System: Microsoft Windows 11 Home, Version 10.0.22631
- Processor: 13th Gen Intel(R) Core(TM) i9-13900KF
- RAM: 32 GB
- GPU: NVIDIA GeForce RTX 4090
- CUDA Version: 12.6 worked system-wide and 11.2 also worked with conda
- Python & Dependencies: As described above and listed in requirements.txt
Please note that you do not need a GPU to run the experiments, however, it is recommended to use parallelization with 20+ threads (jobs) to speed up the experiments.
Models
Pre-trained PCS ensemble models are available via Google Drive:
-
Extract contents to the
models/directory -
A demo folder is included for quick testing
Model Variants:
pcs_top1_qpcs_10_standard- Variant (a)pcs_top1_qxgb_10_standard- Variant (b)pcs_top1_pcs_10_standard- Variant (c)- Conformalized versions also available
Model Training & Calibration Time
Below, we also provide the average training time for each of the variants on a given real-world dataset computed on the machine mentioned above. The times are provided in seconds (s) and have been averaged over 10 seeds. For CLEAR, the only required computation is the calibration time (computation of $\lambda$ and $\gamma_1$). Note that in our experiments and results, we provide many other models (e.g., CQR, CQR-R, ALEATORIC-R/CQR-R+, $\lambda=1$, $\gamma_1=1$, etc.) for comparison purposes; only the times for the methods included in the paper are provided below. The experiments were run exactly as described in the paper (particularly for the 100 bootstraps).
Variant (a)
| Dataset | PCS (s) | ALEATORIC-R (s) | CLEAR (s) | Total (s) |
|---|---|---|---|---|
| ailerons | 13.46 | 6.80 | 0.27 | 20.53 |
| airfoil | 3.10 | 1.92 | 0.17 | 5.19 |
| allstate | 1124.47 | 22.93 | 0.19 | 1147.59 |
| ca_housing | 8.51 | 7.79 | 0.54 | 16.84 |
| computer | 61.92 | 76.09 | 0.16 | 138.16 |
| concrete | 1.10 | 2.16 | 0.17 | 3.43 |
| elevator | 18.79 | 122.16 | 0.60 | 141.54 |
| energy_efficiency | 0.81 | 1.65 | 0.16 | 2.62 |
| insurance | 3.48 | 17.23 | 0.11 | 20.82 |
| kin8nm | 3.25 | 5.43 | 0.21 | 8.89 |
| miami_housing | 7.31 | 7.66 | 0.72 | 15.68 |
| naval_propulsion | 19.83 | 81.32 | 0.29 | 101.44 |
| parkinsons | 39.53 | 60.99 | 0.14 | 100.66 |
| powerplant | 3.08 | 4.92 | 0.21 | 8.22 |
| qsar | 348.59 | 10.00 | 0.20 | 358.78 |
| sulfur | 21.55 | 16.56 | 0.21 | 38.32 |
| superconductor | 561.84 | 795.45 | 12.70 | 1369.99 |
| Total | 2240.60 | 1241.06 | 17.06 | 3498.71 |
Variant (b)
| Dataset | PCS (s) | ALEATORIC-R (s) | CLEAR (s) | Total (s) |
|---|---|---|---|---|
| ailerons | 4.51 | 6.89 | 0.26 | 11.66 |
| airfoil | 0.78 | 1.64 | 0.16 | 2.59 |
| allstate | 27.73 | 14.22 | 0.20 | 42.15 |
| ca_housing | 5.42 | 7.69 | 0.39 | 13.50 |
| computer | 3.46 | 5.97 | 0.21 | 9.65 |
| concrete | 0.86 | 1.82 | 0.17 | 2.85 |
| elevator | 4.43 | 6.71 | 0.79 | 11.93 |
| energy_efficiency | 0.68 | 1.40 | 0.17 | 2.25 |
| insurance | 0.80 | 1.65 | 0.17 | 2.62 |
| kin8nm | 2.61 | 4.96 | 0.21 | 7.78 |
| miami_housing | 4.61 | 7.70 | 0.70 | 13.01 |
| naval_propulsion | 3.72 | 6.38 | 0.55 | 10.64 |
| parkinsons | 2.71 | 5.37 | 0.20 | 8.28 |
| powerplant | 2.73 | 4.32 | 0.22 | 7.26 |
| qsar | 5.00 | 9.67 | 0.20 | 14.86 |
| sulfur | 2.89 | 4.64 | 0.23 | 7.76 |
| superconductor | 13.81 | 22.74 | 1.00 | 37.55 |
| Total | 86.77 | 113.76 | 5.81 | 206.34 |
Variant (c)
| Dataset | PCS (s) | ALEATORIC-R (s) | CLEAR (s) | Total (s) |
|---|---|---|---|---|
| ailerons | 10.20 | 104.32 | 0.65 | 115.17 |
| airfoil | 0.88 | 17.82 | 0.11 | 18.81 |
| allstate | 651.02 | 125.77 | 0.14 | 776.93 |
| ca_housing | 3.63 | 130.87 | 4.16 | 138.66 |
| computer | 4.55 | 65.28 | 0.16 | 69.98 |
| concrete | 1.16 | 17.49 | 0.11 | 18.76 |
| elevator | 3.25 | 98.73 | 1.97 | 103.95 |
| energy_efficiency | 0.95 | 15.42 | 0.10 | 16.47 |
| insurance | 0.90 | 19.80 | 0.11 | 20.81 |
| kin8nm | 16.89 | 69.17 | 0.16 | 86.22 |
| miami_housing | 5.34 | 123.32 | 0.92 | 129.58 |
| naval_propulsion | 1.14 | 115.41 | 0.74 | 117.28 |
| parkinsons | 3.71 | 53.43 | 0.14 | 57.27 |
| powerplant | 2.37 | 60.99 | 0.17 | 63.52 |
| qsar | 21.48 | 93.84 | 0.14 | 115.46 |
| sulfur | 2.43 | 67.41 | 0.17 | 70.01 |
| superconductor | 60.63 | 644.13 | 8.10 | 712.86 |
| Total | 790.52 | 1823.18 | 18.05 | 2631.75 |
Data
The project utilizes 18 diverse real-world regression datasets (17 for the benchmarks and 1 for the case study). All processed datasets are provided in the data/ directory. The script src/experiments/download_process_real_data.py (adapted from the original PCS-UQ paper's data processing notebook) was used to download and process these datasets.
Each dataset is stored in its own subdirectory within data/ (e.g., data/data_ailerons/) and typically includes the following files:
X.csv: The feature matrix (processed, with one-hot encoding for categorical features).y.csv: The target variable vector.importances.csv: Feature importances as determined by a Random Forest model during preprocessing.bin_df.pkl: A pickled Pandas DataFrame containing binned versions of features, used for subgroup analysis.
The datasets included are:
| Dataset | n | d |
|---|---|---|
| ailerons | 13750 | 40 |
| airfoil | 1503 | 5 |
| allstate | 5000 | 1037 |
| ca housing | 20640 | 8 |
| computer | 8192 | 21 |
| concrete | 1030 | 8 |
| diamond | 53940 | 23 |
| elevator | 16599 | 18 |
| energy efficiency | 768 | 10 |
| insurance | 1338 | 8 |
| kin8nm | 8192 | 8 |
| miami housing | 13932 | 28 |
| naval propulsion | 11934 | 24 |
| parkinsons | 5875 | 18 |
| powerplant | 9568 | 4 |
| qsar | 5742 | 500 |
| sulfur | 10081 | 5 |
| superconductor | 21263 | 79 |
For details on the original sources and specific preprocessing steps for each dataset, please refer to the comments and documentation within src/experiments/download_process_real_data.py.
Citation
If you use CLEAR in your research, please cite:
@inproceedings{
azizi2026clear,
title={{CLEAR}: Calibrated Learning for Epistemic and Aleatoric Risk},
author={Ilia Azizi and Juraj Bodik and Jakob Heiss and Bin Yu},
booktitle={The Fourteenth International Conference on Learning Representations},
year={2026},
url={https://openreview.net/forum?id=RY4IHaDLik}
}
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file clear_uq-0.3.1.tar.gz.
File metadata
- Download URL: clear_uq-0.3.1.tar.gz
- Upload date:
- Size: 164.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
279a86204b8545bb8f7e27e105252c829f7e8bfecdec8befb82647a627d8b805
|
|
| MD5 |
156763d9d200aa92a35808cb7821e27a
|
|
| BLAKE2b-256 |
12f1cf651d3843ff837e907caf6986c3ace6d16e3827279720bae5fdc262da78
|
Provenance
The following attestation bundles were made for clear_uq-0.3.1.tar.gz:
Publisher:
publish-to-pypi.yml on Unco3892/clear
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
clear_uq-0.3.1.tar.gz -
Subject digest:
279a86204b8545bb8f7e27e105252c829f7e8bfecdec8befb82647a627d8b805 - Sigstore transparency entry: 1003798331
- Sigstore integration time:
-
Permalink:
Unco3892/clear@fa4605181c101820bb662d6369abe55042bf2799 -
Branch / Tag:
refs/tags/v0.3.1 - Owner: https://github.com/Unco3892
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-to-pypi.yml@fa4605181c101820bb662d6369abe55042bf2799 -
Trigger Event:
release
-
Statement type:
File details
Details for the file clear_uq-0.3.1-py3-none-any.whl.
File metadata
- Download URL: clear_uq-0.3.1-py3-none-any.whl
- Upload date:
- Size: 171.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5d3226148ab3200bb2e3ab82ccad7b5f1082212b22d34e114aa81802bcdb72d5
|
|
| MD5 |
81b3ab2cb73971abeb0e00e9b7374e05
|
|
| BLAKE2b-256 |
4e6d6f6f94b7fca0a6f529b9b6956ff3cda226dd1e59fab6f48d263ae8815cd4
|
Provenance
The following attestation bundles were made for clear_uq-0.3.1-py3-none-any.whl:
Publisher:
publish-to-pypi.yml on Unco3892/clear
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
clear_uq-0.3.1-py3-none-any.whl -
Subject digest:
5d3226148ab3200bb2e3ab82ccad7b5f1082212b22d34e114aa81802bcdb72d5 - Sigstore transparency entry: 1003798335
- Sigstore integration time:
-
Permalink:
Unco3892/clear@fa4605181c101820bb662d6369abe55042bf2799 -
Branch / Tag:
refs/tags/v0.3.1 - Owner: https://github.com/Unco3892
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-to-pypi.yml@fa4605181c101820bb662d6369abe55042bf2799 -
Trigger Event:
release
-
Statement type: