climax is a Command Line IMAge eXplorer
Project description
CLImax (a Command Line IMAge eXplorer)
Installing CLImax
We recommend that you install CLImax as a tool using uv:
$ uv tool install climax-rfglab
A CLImax executable will be created in a bin directory in the PATH, which allows the tool to be run without uv. If the directory with the CLImax executable is not in the PATH, a warning will be displayed and the following command can be issued to add it to the PATH:
$ uv tool update-shell
CLImax can also be installed as a regular Python package using uv:
$ uv pip install climax-rfglab
or pip:
$ python3 -m pip install climax-rfglab
A note on the Python interpreter
CLImax requires that you have Python 3.10 or above installed.
Using CLImax
After installing CLImax with one of the methods above, You can invoke it with:
$ climax <filename>
You can also run CLImax without installing it, taking advantage of uvx:
$ uvx --from climax-rfglab climax <filename>
There are a few ways to open an image with CLImax:
-
You can specifiy the path to the image that you want to open (e.g. tests/cells_movie.tif) :
$ climax tests/cells_movie.tif -
Or you can indicate a folder (slices in this example) that contains an image sequence:
$ climax ./slices -
If there are image channels split into different files, you can specify a group of substrings to distinguish which files in the folder belong to which channel. For example, to open the files in the slices folder containing the substrings '488' and '561' as two different channels:
$ climax ./slices -s 488 561 -
You can use a list of paths to concatenate sideways (i.e. display side-by-side, but all the images must have the same dimensions!!):
$ climax cells_movie_1.tif cells_movie_2.tif -
You can specify the color map used to display the image. The color map defaults to 'gray'. Check here for a list of color maps.
$ climax cells_movie.tif -c viridis -
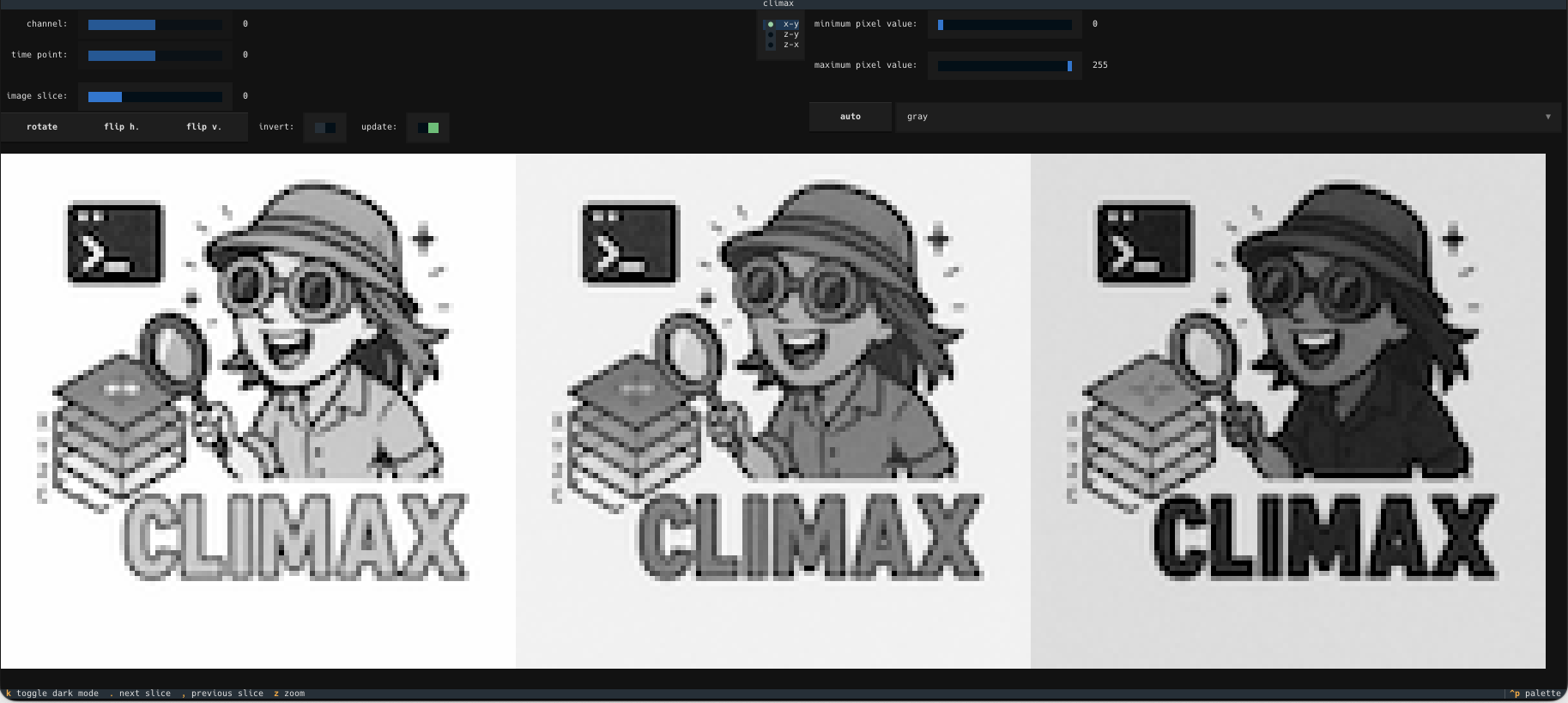
Notice that if you try to open a color image, the channels will be split and displayed side-by-side (red, green, blue). Thus, for instance, if you open the CLImax logo:
CLImax provides a dashboard with additional functionality:
| name | function |
|---|---|
| rotate | rotate 90° clockwise |
| flip h. | flip horizontally |
| flip v. | flip vertically |
| invert | invert color map |
| update | continuous/discrete update |
| x-y, z-y, z-x | slicing plane through stack |
| auto | automated contrast adjustment (min-mode, max-99th percentile) |
CLImax today
As we develop and improve CLImax, there may be small changes to the user interface. This is how CLImax looks as of today:
Citing CLImax
If you use CLImax, please cite this repository. We are working on the paper!
Sponsors
We are grateful for the generous support from the following agencies and institutions, which contribute to the development and maintenance of CLImax:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file climax_rfglab-2025.5.1.tar.gz.
File metadata
- Download URL: climax_rfglab-2025.5.1.tar.gz
- Upload date:
- Size: 25.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.5.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d00b9f241c30c7994db3a1ed8265a40ed6d558db63e5dfd1b987369924f41bfc
|
|
| MD5 |
99d6cb4916472ff219d5c6284182e77e
|
|
| BLAKE2b-256 |
9ae659cdeb61660eca77f4bbbf8f0b8557f64ca7fac284f18adca9af576f0822
|
File details
Details for the file climax_rfglab-2025.5.1-py3-none-any.whl.
File metadata
- Download URL: climax_rfglab-2025.5.1-py3-none-any.whl
- Upload date:
- Size: 24.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.5.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
82632516b3e7f121a6ff4bcb6da0d77f3bd20b9a2febb98c5dcc0631d4d0b494
|
|
| MD5 |
332fc15002dfe5f5503d196db1579204
|
|
| BLAKE2b-256 |
edd745c18066bb16cd9ba9d140b63670f0af77889070072c2f53f44935cfabad
|