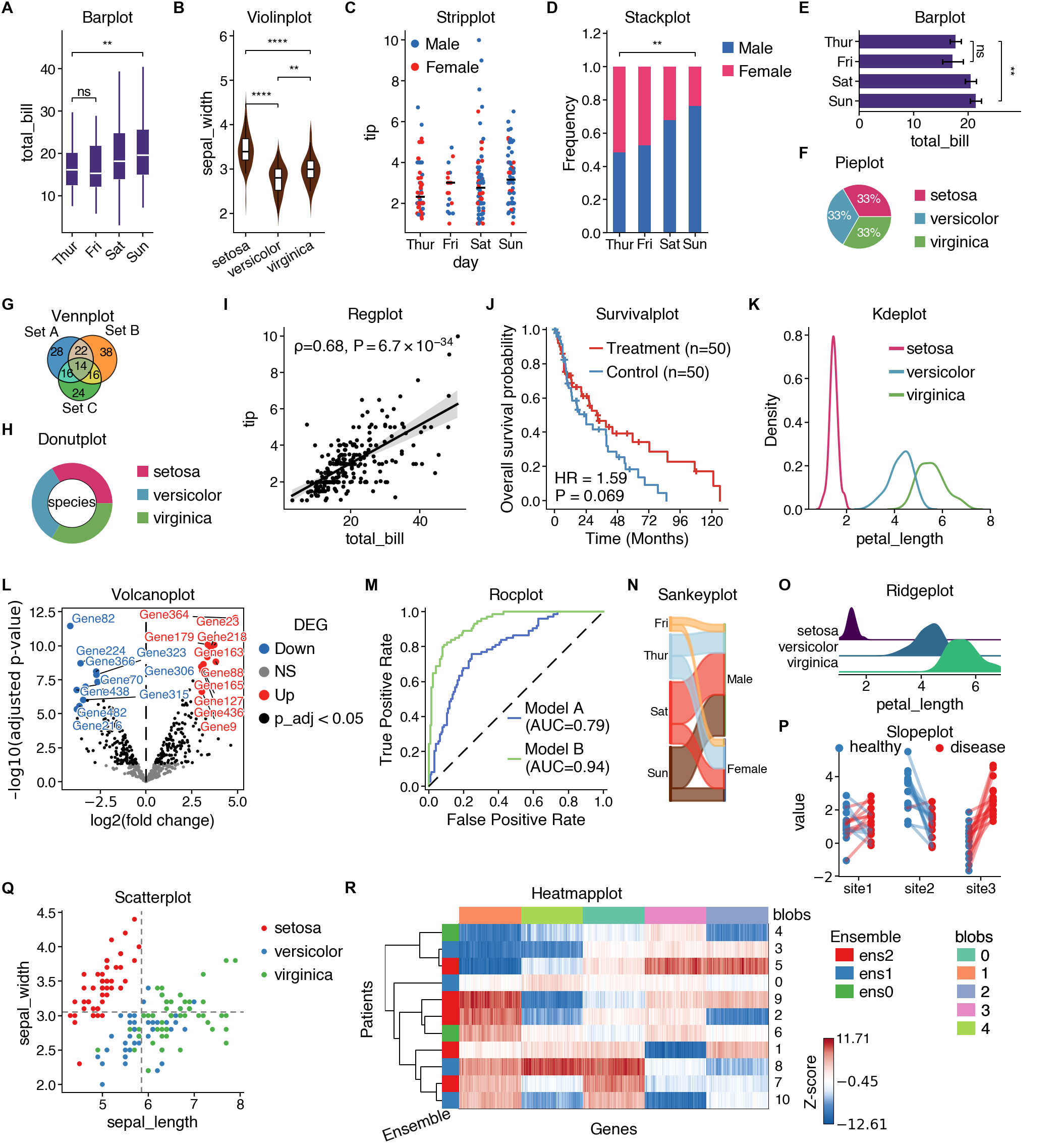
A comprehensive plotting library for bioinformatics and data visualization
Project description
cnsplots

Publication-Ready Scientific Plots for Cell, Nature, and Science Journals
Create visually stunning, journal-quality figures with minimal code. Built on matplotlib, fully compatible with seaborn, and optimized for Adobe Illustrator.
Documentation · Examples Gallery · Report Bug · Request Feature
Overview
cnsplots is a Python visualization library designed specifically for creating publication-ready scientific figures. It takes care of the tedious styling details so you can focus on your science.
Why cnsplots?
- 🎨 Publication-Ready: Pre-configured styles matching Cell, Nature, and Science journal requirements
- 🎯 Simple API: Create complex multi-panel figures with just a few lines of code
- 📐 Precise Control: Specify dimensions in pixels, perfect for journal submission guidelines
- 🖋️ Adobe Illustrator Compatible: SVG exports with editable fonts (no text-to-path conversion)
- 📊 Statistical Integration: Built-in statistical tests and annotations
- 🔧 Highly Customizable: Full control over colors, fonts, and styling
- 🌈 Rich Color Palettes: Curated color schemes optimized for scientific visualization
- 🧩 Multi-Panel Support: Easy creation of complex figure layouts
Features
📊 25+ Plot Types
Basic Plots
- Box plots, violin plots, bar plots, strip plots
- Scatter plots, line plots, regression plots
- Histograms, KDE plots, ridge plots
Scientific Plots
- Survival plots (Kaplan-Meier)
- Cumulative incidence plots
- ROC curves and forest plots
- Volcano plots and GSEA plots
- Confusion matrices
Specialized Plots
- Heatmaps with hierarchical clustering
- Dot plots for enrichment
- Venn diagrams and UpSet plots
- Sankey diagrams
- Pie and donut charts
- QQ plots and slope plots
🎨 Beautiful Color Palettes
Multiple curated palettes including:
- Qualitative: Cell, Nature, Science, Ecotyper1-6, Set1-3, Tableau, Bold
- Sequential: Parula, gnuplot, custom gradients
- Diverging: BlueRed, BuRd_custom, OrBu_custom
📐 Multi-Panel Figures
Create complex layouts with automatic panel labeling (A, B, C...):
import cnsplots as cns
mp = cns.multipanel(max_width=540)
# Panel A
mp.panel("A", height=150, width=150)
cns.boxplot(data=df1, x="group", y="value")
# Panel B
mp.panel("B", height=150, width=150)
cns.scatterplot(data=df2, x="x", y="y")
# Continues...
Installation
From PyPI
pip install cnsplots
For Development
First install uv, then:
git clone https://github.com/faridrashidi/cnsplots
cd cnsplots
make install
This installs the package with its development and documentation extras. Core
scientific integrations such as scanpy, lifelines, and gseapy remain
runtime dependencies of cnsplots itself.
Quick Start
Basic Usage
import cnsplots as cns
import seaborn as sns
# Load example data
df = sns.load_dataset("tips")
# Create a figure (height, width in pixels)
cns.figure(height=150, width=100)
# Create a publication-ready boxplot
cns.boxplot(data=df, x="day", y="total_bill")
# Save as vector graphic
cns.savefig("figure.svg")
Statistical Comparisons
# Add statistical significance annotations
cns.figure(150, 150)
cns.boxplot(
data=df,
x="day",
y="total_bill",
pairs=[("Thur", "Fri"), ("Sat", "Sun")], # Compare these pairs
)
# Prints: P-values were determined by two-sided Mann-Whitney U test.
Custom Colors
# Use custom color palette
cns.figure(150, 200, color_cycle="Ecotyper1")
cns.violinplot(data=df, x="day", y="total_bill", hue="sex")
Examples Gallery
Explore our comprehensive examples gallery featuring:
- 📦 Basic statistical plots
- 🧬 Genomics and bioinformatics visualizations
- 📈 Time-series and survival analysis
- 🎯 Machine learning results (ROC, confusion matrices)
- 🔬 Multi-omics data visualization
- 🎨 Custom color schemes and styling
Documentation
Full documentation is available at cnsplots.farid.one
Key Concepts
Figure Dimensions
Specify sizes in pixels for precise control:
cns.figure(height=150, width=100) # Starts from a 150px × 100px canvas
With cns.settings.figure_autofit=True (the default), the final rendered
canvas can expand or trim on draw to keep long titles, outside legends, and
annotations tightly in bounds.
Color Palettes
Access curated color palettes:
# Qualitative palettes (for categorical data)
cns.figure(color_cycle="Ecotyper1") # Default, optimized for journals
cns.figure(color_cycle="Cell") # Custom Cell-inspired journal palette
cns.figure(color_cycle="Nature") # Nature-inspired journal palette
cns.figure(color_cycle="Science") # Science-inspired journal palette
cns.figure(color_cycle="Set1") # ColorBrewer Set1
# Sequential palettes (for continuous data)
cns.figure(color_map="parula") # MATLAB-style
cns.figure(color_map="gnuplot") # Default sequential
# Get individual colors
red = cns.RED
blue = cns.BLUE
Statistical Tests
Many plot functions include built-in statistical testing:
# Boxplot with Mann-Whitney U test
cns.boxplot(data=df, x="group", y="value", pairs="all")
# Barplot with Welch's t-test
cns.barplot(data=df, x="group", y="value", pairs=[("A", "B")])
# Stackplot with Fisher's exact test
cns.stackplot(data=df, x="group", y="category", pairs=[("A", "B")])
Export for Publication
# SVG for vector graphics (recommended)
cns.savefig("figure.svg")
# High-resolution PNG
cns.savefig("figure.png")
# PDF with editable text
cns.savefig("figure.pdf")
For Illustrator-optimized SVG post-processing, install MuPDF's mutool.
Without it, cns.savefig("figure.svg") falls back to a standard matplotlib SVG
and emits a warning instead of failing.
Requirements
- Python ≥ 3.9
- Core: matplotlib, numpy, pandas, seaborn
- Included integrations: lifelines, gseapy, scanpy, and other plotting backends
- Optional external tool: MuPDF's
mutoolfor enhanced SVG post-processing
See pyproject.toml for complete dependency list.
Contributing
Contributions are welcome! Please feel free to submit a Pull Request. For major changes, please open an issue first to discuss what you would like to change.
Citation
If you use cnsplots in your research, please cite:
@software{cnsplots,
author = {Rashidi, Farid},
title = {cnsplots: Publication-Ready Scientific Plots},
year = {2026},
url = {https://github.com/faridrashidi/cnsplots}
}
License
This project is licensed under the BSD 3-Clause License - see the LICENSE.md file for details.
Acknowledgments
Built with:
- matplotlib - Core plotting library
- seaborn - Statistical visualizations
- lifelines - Survival analysis
- PyComplexHeatmap - Complex heatmaps
- UpSetPlot - Set intersections
Inspired by the visualization standards of Cell, Nature, and Science journals.
Support
Related Projects
- matplotlib - The foundation of Python plotting
- seaborn - Statistical data visualization
- plotnine - Grammar of graphics for Python
- altair - Declarative visualization
Made with ❤️ for the scientific community
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cnsplots-0.1.0.tar.gz.
File metadata
- Download URL: cnsplots-0.1.0.tar.gz
- Upload date:
- Size: 131.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b346a87dd917f49b8c3f42ae722b877ebd13ce935035aedf8099ed5cc2d69bd7
|
|
| MD5 |
ac5e4772244a751fac08b73bca5832e4
|
|
| BLAKE2b-256 |
885f69942807b8de96e69d830f257d09c0ce4ff74260d24fc315c1e73242aa79
|
Provenance
The following attestation bundles were made for cnsplots-0.1.0.tar.gz:
Publisher:
release-publish.yml on faridrashidi/cnsplots
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
cnsplots-0.1.0.tar.gz -
Subject digest:
b346a87dd917f49b8c3f42ae722b877ebd13ce935035aedf8099ed5cc2d69bd7 - Sigstore transparency entry: 1154692820
- Sigstore integration time:
-
Permalink:
faridrashidi/cnsplots@9d3d9e4743dfbe763e182098e44dd64193742369 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/faridrashidi
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release-publish.yml@9d3d9e4743dfbe763e182098e44dd64193742369 -
Trigger Event:
push
-
Statement type:
File details
Details for the file cnsplots-0.1.0-py3-none-any.whl.
File metadata
- Download URL: cnsplots-0.1.0-py3-none-any.whl
- Upload date:
- Size: 103.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fbfab6efbe43b8705d1ed16cad10e1a7065c4687067f78da3767636d580329a8
|
|
| MD5 |
a281abe625b7ab6995f7f38cd88e6e65
|
|
| BLAKE2b-256 |
f9366d447da5501511854b1d780fd2f3f94b4f3ff0c23376a46424fb9e16cfa0
|
Provenance
The following attestation bundles were made for cnsplots-0.1.0-py3-none-any.whl:
Publisher:
release-publish.yml on faridrashidi/cnsplots
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
cnsplots-0.1.0-py3-none-any.whl -
Subject digest:
fbfab6efbe43b8705d1ed16cad10e1a7065c4687067f78da3767636d580329a8 - Sigstore transparency entry: 1154692824
- Sigstore integration time:
-
Permalink:
faridrashidi/cnsplots@9d3d9e4743dfbe763e182098e44dd64193742369 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/faridrashidi
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release-publish.yml@9d3d9e4743dfbe763e182098e44dd64193742369 -
Trigger Event:
push
-
Statement type: