Connectome Mapper 3: A Flexible and Open-Source Pipeline Software for Multiscale Multimodal Human Connectome Mapping
Project description
Connectome Mapper 3
This neuroimaging processing pipeline software is developed by the Connectomics Lab at the University Hospital of Lausanne (CHUV) for use within the SNF Sinergia Project 170873, as well as for open-source software distribution.
Description
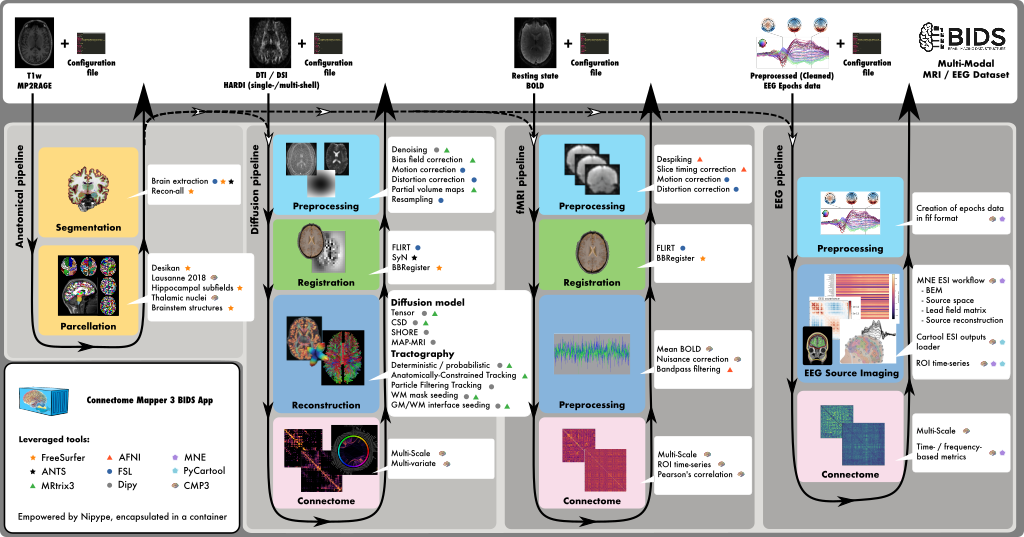
Connectome Mapper 3 is an open-source Python3 image processing pipeline software, with a Graphical User Interface, that implements full anatomical, diffusion, resting-state MRI, and EEG processing pipelines, from raw Diffusion / T1 / BOLD / preprocessed EEG to multi-resolution connection matrices, based on a new version of the Lausanne parcellation atlas, aka Lausanne2018.
Connectome Mapper 3 pipelines use a combination of tools from well-known software packages, including FSL, FreeSurfer, ANTs, MRtrix3, Dipy, AFNI, MNE, MNE_Connectivity, and Cartool (PyCartool) orchestrated by the Nipype dataflow library. These pipelines were designed to provide the best software implementation for each state of processing at the time conceptualization, and can be updated as newer and better neuroimaging software become available.
To enhance reproducibility and replicatibility, the processing pipelines with all dependencies are encapsulated in a Docker image container, which handles datasets organized following the BIDS standard and is distributed as a BIDS App @ Docker Hub. For execution on high-performance computing cluster, a Singularity image is also made freely available @ Sylabs Cloud.
To reduce the risk of misconfiguration and improve accessibility, Connectome Mapper 3 comes with an interactive GUI, aka cmpbidsappmanager, which supports the user in all the steps involved in the configuration of the pipelines, the configuration and execution of the BIDS App, and the control of the output quality. In addition, to facilitate the use by users not familiar with Docker and Singularity containers, Connectome Mapper 3 provides two Python commandline wrappers (connectomemapper3_docker and connectomemapper3_singularity) that will generate and run the appropriate command.
Since v3.1.0, CMP3 provides full support to EEG. Please check this notebook for a demonstration of the newly implemented pipeline, using the “VEPCON” dataset, available at https://openneuro.org/datasets/ds003505/versions/1.1.1.
How to install the python wrappers and the GUI?
You need to have first either Docker or Singularity engine and miniconda installed. We refer to the dedicated documentation page for more instruction details.
Then, download the appropriate environment.yml / environment_macosx.yml and create a conda environment py39cmp-gui with the following command:
$ conda env create -f /path/to/environment[_macosx].yml
Once the environment is created, activate it and install Connectome Mapper 3 with PyPI as follows:
$ conda activate py39cmp-gui
(py39cmp-gui)$ pip install connectomemapper
You are ready to use Connectome Mapper 3!
Resources
- JOSS paper: https://joss.theoj.org/papers/10.21105/joss.04248
- Documentation: https://connectome-mapper-3.readthedocs.io
- Mailing list: https://groups.google.com/forum/#!forum/cmtk-users
- Source: https://github.com/connectomicslab/connectomemapper3
- Bug reports: https://github.com/connectomicslab/connectomemapper3/issues
Carbon footprint estimation of BIDS App run 🌍🌳✨
In support to the Organisation for Human Brain Mapping (OHBM)
Sustainability and Environmental Action (OHBM-SEA) group, CMP3 enables you
since v3.0.3 to be more aware about the adverse impact of your processing
on the environment!
With the new --track_carbon_footprint option of the connectomemapper3_docker and connectomemapper3_singularity
BIDS App python wrappers, and the new "Track carbon footprint" option of the cmpbidsappmanager BIDS Interface Window,
you can estimate the carbon footprint incurred by the execution of the BIDS App.
Estimations are conducted using codecarbon to estimate the amount of carbon dioxide (CO2)
produced to execute the code by the computing resources and save the results in <bids_dir>/code/emissions.csv.
Then, to visualize, interpret and track the evolution of the emitted CO2 emissions, you can use the visualization
tool of codecarbon aka carbonboard that takes as input the .csv created::
$ carbonboard --filepath="<bids_dir>/code/emissions.csv" --port=xxxx
Please check https://ohbm-environment.org to learn more about OHBM-SEA!
Usage
Having the py39cmp-gui conda environment previously installed activated, the BIDS App can easily be run using connectomemapper3_docker, the python wrapper for Docker, as follows:
usage: connectomemapper3_docker [-h]
[--participant_label PARTICIPANT_LABEL [PARTICIPANT_LABEL ...]]
[--session_label SESSION_LABEL [SESSION_LABEL ...]]
[--anat_pipeline_config ANAT_PIPELINE_CONFIG]
[--dwi_pipeline_config DWI_PIPELINE_CONFIG]
[--func_pipeline_config FUNC_PIPELINE_CONFIG]
[--eeg_pipeline_config EEG_PIPELINE_CONFIG]
[--number_of_threads NUMBER_OF_THREADS]
[--number_of_participants_processed_in_parallel NUMBER_OF_PARTICIPANTS_PROCESSED_IN_PARALLEL]
[--mrtrix_random_seed MRTRIX_RANDOM_SEED]
[--ants_random_seed ANTS_RANDOM_SEED]
[--ants_number_of_threads ANTS_NUMBER_OF_THREADS]
[--fs_license FS_LICENSE] [--coverage]
[--notrack] [-v] [--track_carbon_footprint]
[--docker_image DOCKER_IMAGE]
[--config_dir CONFIG_DIR]
bids_dir output_dir {participant,group}
Entrypoint script of the Connectome Mapper BIDS-App version v3.1.0 via Docker.
positional arguments:
bids_dir The directory with the input dataset formatted
according to the BIDS standard.
output_dir The directory where the output files should be stored.
If you are running group level analysis this folder
should be prepopulated with the results of the
participant level analysis.
{participant,group} Level of the analysis that will be performed. Multiple
participant level analyses can be run independently
(in parallel) using the same output_dir.
optional arguments:
-h, --help show this help message and exit
--participant_label PARTICIPANT_LABEL [PARTICIPANT_LABEL ...]
The label(s) of the participant(s) that should be
analyzed. The label corresponds to
sub-<participant_label> from the BIDS spec (so it does
not include "sub-"). If this parameter is not provided
all subjects should be analyzed. Multiple participants
can be specified with a space separated list.
--session_label SESSION_LABEL [SESSION_LABEL ...]
The label(s) of the session that should be analyzed.
The label corresponds to ses-<session_label> from the
BIDS spec (so it does not include "ses-"). If this
parameter is not provided all sessions should be
analyzed. Multiple sessions can be specified with a
space separated list.
--anat_pipeline_config ANAT_PIPELINE_CONFIG
Configuration .json file for processing stages of the
anatomical MRI processing pipeline
--dwi_pipeline_config DWI_PIPELINE_CONFIG
Configuration .json file for processing stages of the
diffusion MRI processing pipeline
--func_pipeline_config FUNC_PIPELINE_CONFIG
Configuration .json file for processing stages of the
fMRI processing pipeline
--eeg_pipeline_config EEG_PIPELINE_CONFIG
Configuration .json file for processing stages of the
EEG processing pipeline
--number_of_threads NUMBER_OF_THREADS
The number of OpenMP threads used for multi-threading
by Freesurfer (Set to [Number of available CPUs -1] by
default).
--number_of_participants_processed_in_parallel NUMBER_OF_PARTICIPANTS_PROCESSED_IN_PARALLEL
The number of subjects to be processed in parallel
(One by default).
--mrtrix_random_seed MRTRIX_RANDOM_SEED
Fix MRtrix3 random number generator seed to the
specified value
--ants_random_seed ANTS_RANDOM_SEED
Fix ANTS random number generator seed to the specified
value
--ants_number_of_threads ANTS_NUMBER_OF_THREADS
Fix number of threads in ANTs. If not specified ANTs
will use the same number as the number of OpenMP
threads (see `----number_of_threads` option flag)
--fs_license FS_LICENSE
Freesurfer license.txt
--coverage Run connectomemapper3 with coverage
--notrack Do not send event to Google analytics to report BIDS
App execution, which is enabled by default.
-v, --version show program's version number and exit
--track_carbon_footprint
Track carbon footprint with `codecarbon
<https://codecarbon.io/>`_ and save results in a CSV
file called ``emissions.csv`` in the
``<bids_dir>/code`` directory.
--docker_image DOCKER_IMAGE
The path to the docker image.
--config_dir CONFIG_DIR
The path to the directory containing the configuration
files.
Contributors ✨
Thanks goes to these wonderful people (emoji key):
Thanks also goes to all these wonderful people that contributed to Connectome Mapper 1 and 2:
-
Collaborators from Signal Processing Laboratory (LTS5), EPFL, Lausanne:
- Jean-Philippe Thiran
- Leila Cammoun
- Adrien Birbaumer (abirba)
- Alessandro Daducci (daducci)
- Stephan Gerhard (unidesigner)
- Christophe Chênes (Cwis)
- Oscar Esteban (oesteban)
- David Romascano (davidrs06)
- Alia Lemkaddem (allem)
- Xavier Gigandet
-
Collaborators from Children's Hospital, Boston:
- Ellen Grant
- Daniel Ginsburg (danginsburg)
- Rudolph Pienaar (rudolphpienaar)
- Nicolas Rannou (NicolasRannou)
This project follows the all-contributors specification. Contributions of any kind welcome!
How to cite?
Please consult our Citing documentation page.
How to contribute?
Please consult our Contributing to Connectome Mapper 3 guidelines.
Funding
Work supported by the Sinergia SNFNS-170873 Grant.
License
This software is distributed under the open-source 3-Clause BSD License. See license for more details.
All trademarks referenced herein are property of their respective holders.
Copyright (C) 2009-2022, Hospital Center and University of Lausanne (UNIL-CHUV), Ecole Polytechnique Fédérale de Lausanne (EPFL), Switzerland & Contributors.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file connectomemapper-3.2.0.tar.gz.
File metadata
- Download URL: connectomemapper-3.2.0.tar.gz
- Upload date:
- Size: 261.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.7.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8ed5fcab4db7e9eabc8688d6bb6ed1f63aee520e7208544f6f84ab99b4a053d1
|
|
| MD5 |
678c3fdc30e953866589a1ff000324d8
|
|
| BLAKE2b-256 |
4b4ecd9f0b8f818318fe4bc687bccd9f2ae7470f85bbe9eba6a35ca51caa5331
|
File details
Details for the file connectomemapper-3.2.0-py3-none-any.whl.
File metadata
- Download URL: connectomemapper-3.2.0-py3-none-any.whl
- Upload date:
- Size: 5.2 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.7.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
191405313c01a11a62dbda26783431473c122eb8f6d6966c9cbdebf4f7d4231e
|
|
| MD5 |
70ec30bc49620a151e128cb2649d193e
|
|
| BLAKE2b-256 |
e44b2ac5286a02fb7c7993c3bb9ef0c6a1afed64b5ff8415d54b110178189c04
|