A toolkit for dynamic inference of cell fate by integrating state and lineage information.
Project description
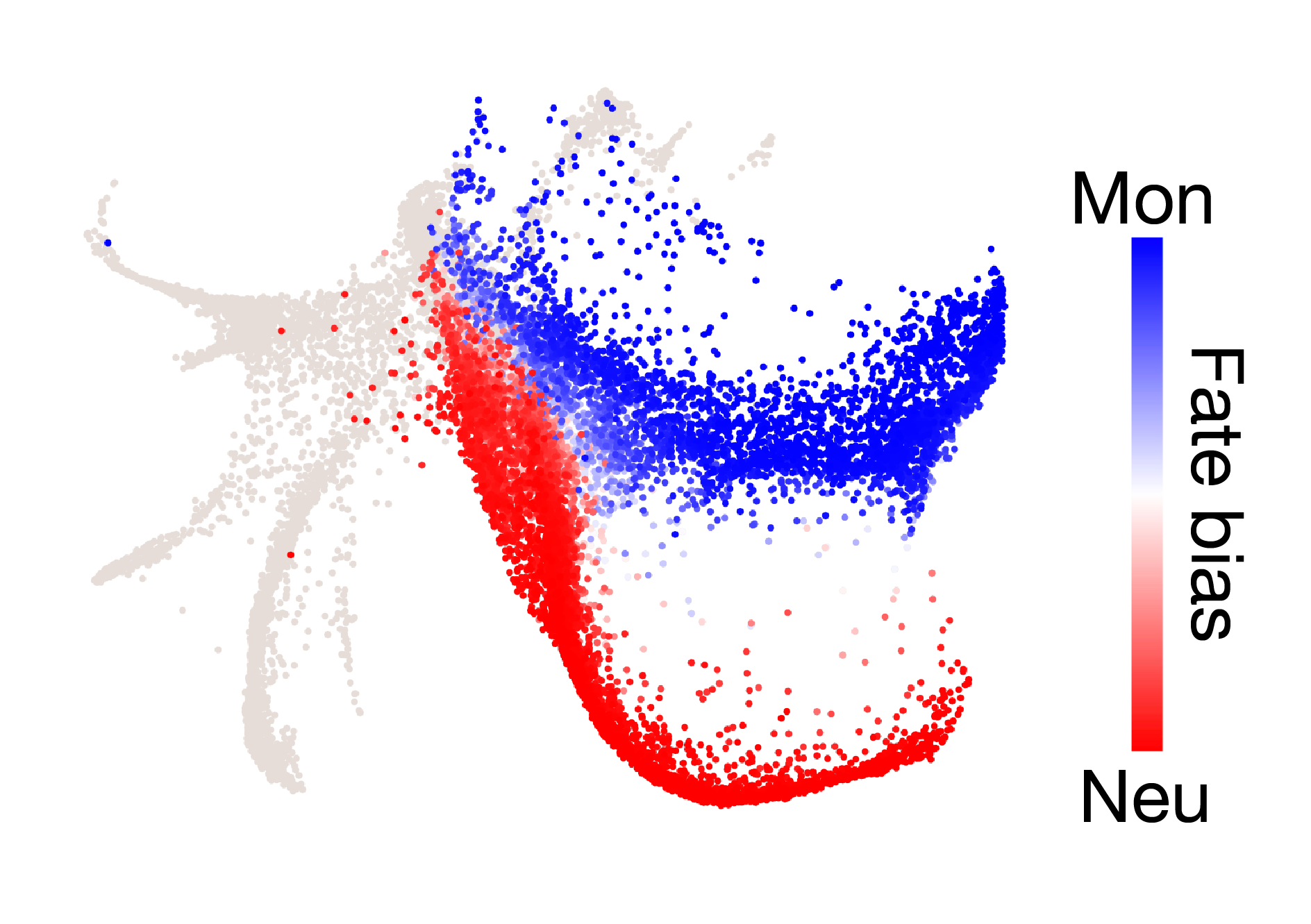
CoSpar is a toolkit for dynamic inference from lineage-traced single cells. The methods are based on Wang et al. Nat. Biotech. (2022).
Dynamic inference based on single-cell state measurement alone requires serious simplifications. On the other hand, direct dynamic measurement via lineage tracing only captures partial information and its interpretation is challenging. CoSpar integrates both state and lineage information to infer a finite-time transition map of a development/differentiation system. It gains superior robustness and accuracy by exploiting both the local coherence and sparsity of differentiation transitions, i.e., neighboring initial states share similar yet sparse fate outcomes. Building around the anndata object, CoSpar provides an integrated analysis framework for datasets with both state and lineage information. When only state information is available, CoSpar also improves upon existing dynamic inference methods by imposing sparsity and coherence. It offers essential toolkits for analyzing lineage data, state information, or their integration.
See https://cospar.readthedocs.io for documentation and tutorials.
Community usage
Our analysis of in vivo cell fate choice of HSCs from DARLIN mice, having only a single time point (with both lineage and transcriptome information in single cells). Ref: L. Li,…,S.-W. Wang, F. Camargo, Cell (2023). Notebooks available here. It is also available in the EXAMPLES session of this website.
Analysis of in vitro lineage tracing data of HSPCs in Jindal et.al., Nat. Biotech. (2023).
Recorded talks
Jun 1: Single-Cell Data Science 2022. This is a 20-min short talk focusing more on the utility of CoSpar: talk video.
Oct 19, 2022: Invited MIA talk at Broad Institute. This is an one-hour talk focusing on the Machine Learning part of CoSpar: talk video.
Reference
S.-W. Wang*, M. Herriges, K. Hurley, D. Kotton, A. M. Klein*, CoSpar identifies early cell fate biases from single cell transcriptomic and lineage information, Nat. Biotech. (2022). [* corresponding authors]
Support
Feel free to submit an issue or send us an email. Your help to improve CoSpar is highly appreciated.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cospar-0.5.0.tar.gz.
File metadata
- Download URL: cospar-0.5.0.tar.gz
- Upload date:
- Size: 106.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.3.2 CPython/3.13.12 Darwin/23.3.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4ad09dd1c841e8e7fef433d671f039b35eb687eb6bb12c63f4478bcf3a339c20
|
|
| MD5 |
d60c258475cb63994173d427d90adf32
|
|
| BLAKE2b-256 |
98d8dedfb05630ae913ad6708890a46a2673026a9fa4561fe6576827d019afc3
|
File details
Details for the file cospar-0.5.0-py3-none-any.whl.
File metadata
- Download URL: cospar-0.5.0-py3-none-any.whl
- Upload date:
- Size: 122.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.3.2 CPython/3.13.12 Darwin/23.3.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b7489850b5ae6b4379c9ad151b3a09604bc68a1e59fb20de5423be94be32dbc2
|
|
| MD5 |
bf3c0f5cad545ba055efb8386d86dbb3
|
|
| BLAKE2b-256 |
fd7669ecbb49c5a463f9cb2d0665a5eb4479cbcb160aad970ca6b5739f797860
|
















