Python module for for creating and manipulating an array of crops (or regions of interest) from images obtained using single-molecule microscopy.
Project description
croparray
Authors: Tim Stasevich and Luis Aguilera.
Description
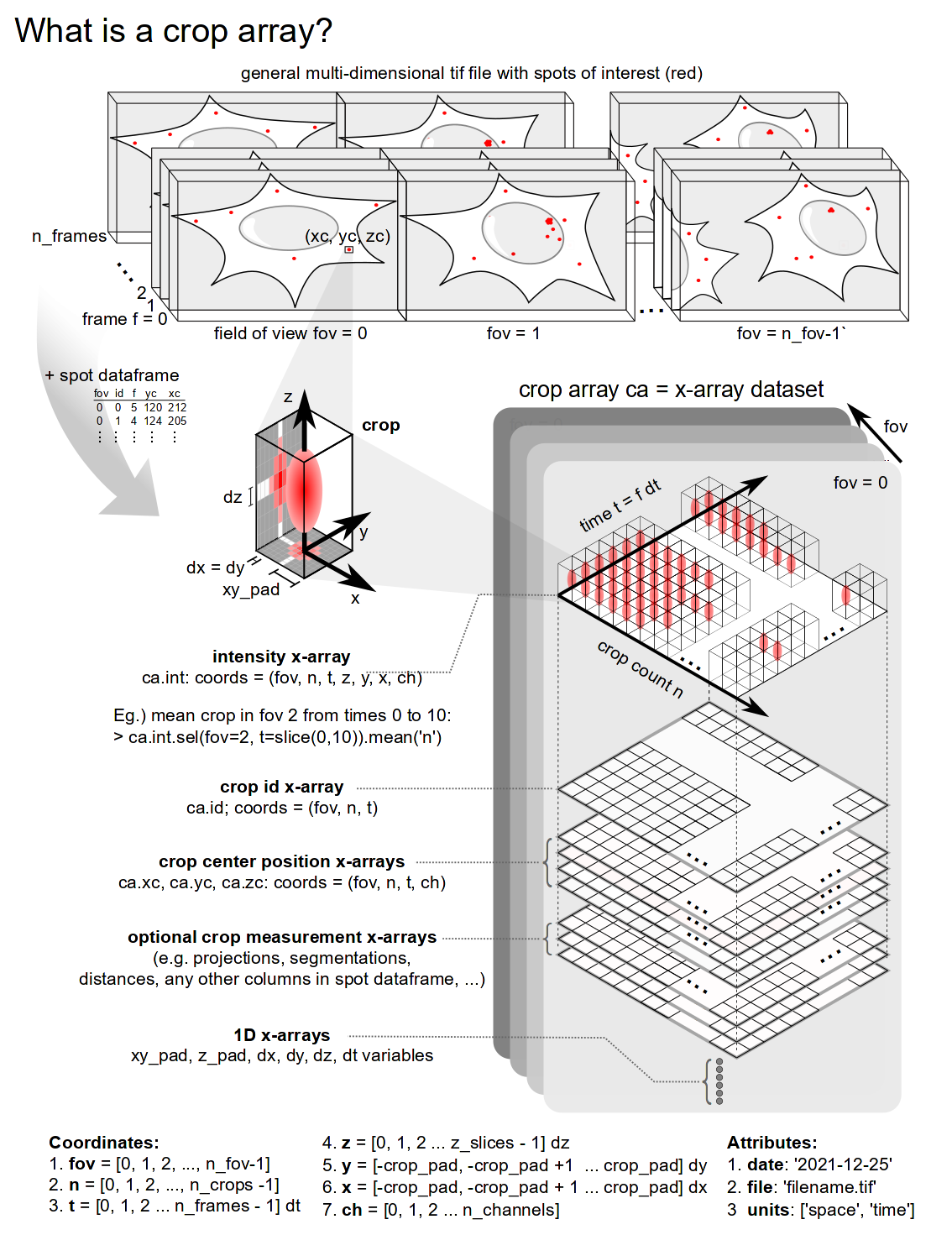
This module is intended for creating and manipulating an array of crops (or regions of interest) that were generated from a multicolor TIF video obtained from single-molecule microscopy.
Documentation
- Documentation is accessible via croparray.readthedocs
Colab implementation

Local installation from the Github repository
-
Install anaconda.
-
Clone the Github repository
git clone https://github.com/Colorado-State-University-Stasevich-Lab/croparray.git
- To create a virtual environment navigate to your local repository and use:
conda create -n croparray_env python=3.8 -y
source activate croparray_env
- To install the rest of requirements use:
pip install -r requirements.txt
- To install napari use:
python -m pip install "napari[all]"
Local installation using PIP
- To create a virtual environment using:
conda create -n croparray_env python=3.8 -y
source activate croparray_env
- Open the terminal and use pip for the installation:
pip install croparray
- To install napari use:
python -m pip install "napari[all]"
Deactivating and removing the environment
- To deactivate or remove the environment from your computer use:
conda deactivate
- To remove the environment use:
conda env remove -n croparray_env
- To unistall croparray use
pip uninstall croparray
additional troubleshooting information
- If you cannot see the package installed on your computer, try using
pip3. For example:
pip3 install croparray
Installing from yml env
- To creating an environment file (yml) use:
source activate croparray_env
conda env export > croparray_env.yml
- ToCreate an environment from this yml file.
conda env create -f croparray_env.yml
Usage
- Organizes crops and measurements of spots of interest from tif images in a convenient x-array format for reduced filesize and more open and reproducible analyses.
- Visualizes crops of detected spots from super-resolution microscope images.
- Calculates the best maximum projection for each crop containing a detected spot.
- Measures intensity of detected spots within crops.
- Calculates the correlation between two equal-length, 1D signals.
- Saves the crop array as a netcdf file at output_direction/output_filename.
- Integrates with Napari for fast and convenient review of crops of detected spots.
Licenses for dependencies
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file croparray-0.1.2.tar.gz.
File metadata
- Download URL: croparray-0.1.2.tar.gz
- Upload date:
- Size: 30.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/54.2.0 requests-toolbelt/0.9.1 tqdm/4.62.3 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dea46108701ba38bfea4d230da48c8f4aa4893d0749fb156f2f440a860cfabe7
|
|
| MD5 |
ab69c3a857d7afffd4e1c0c3fa1d890b
|
|
| BLAKE2b-256 |
8fde6a7a1c5aaa112522f0f7606c79cbe0c73a120e76f49a4622dd1f2ce3beec
|
File details
Details for the file croparray-0.1.2-py3-none-any.whl.
File metadata
- Download URL: croparray-0.1.2-py3-none-any.whl
- Upload date:
- Size: 29.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/54.2.0 requests-toolbelt/0.9.1 tqdm/4.62.3 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
603da1099c66b3b908612845cfa7c76521d70b3fb9669aefcbdbffaff5b690a2
|
|
| MD5 |
6c17a2e3115681fb228c134dc068b138
|
|
| BLAKE2b-256 |
4b9c257287e4f8d2523ef5112a12a59f4d35cfd2ede11e7dbce5ca9f99c594ab
|












