Dissecting Atlas-Registered Tissue (DART): align tissue images to an atlas and generate region ROIs for laser microdissection.
Project description
Overview
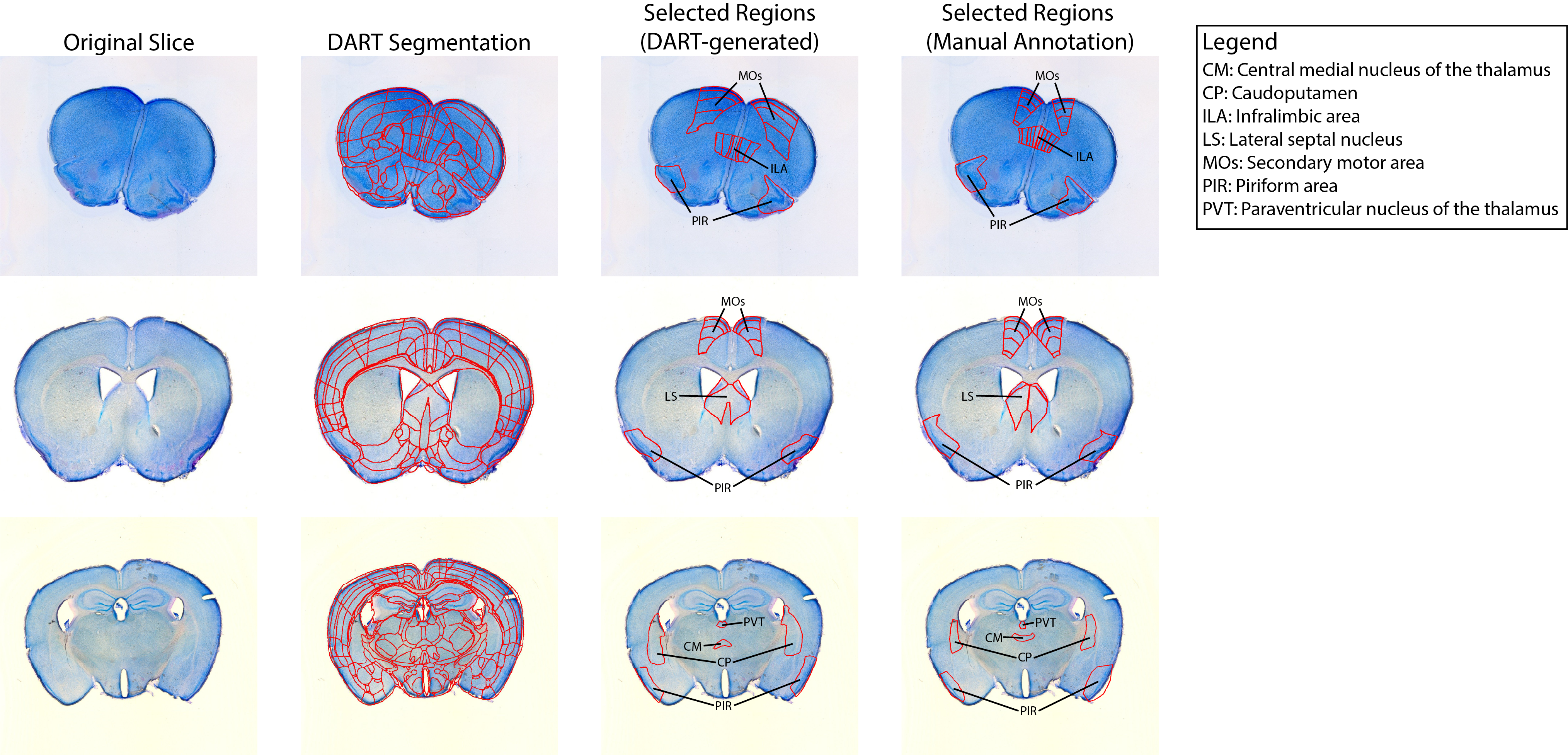
Dissecting Atlas-Registered Tissue (DART) aligns cell-stained images to the Allen Brain Atlas to allow for automatic brain region detection and excision with LEICA laser microdissection software.
DART enables:
- Automated or semi-automated alignment of 2D brain slices to a 3D atlas (e.g., the Allen Brain Atlas CCFv3 2017) and atlas-based segmentation.
- Automatic generation of ROI boundaries.
- Seamless sample handling and integration with Leica LMD software for laser dissection.
Installation
DART can currently be installed via two methods:
Regardless of installation method, DART is available only on Windows 11. It was tested on a computer with 192 GB RAM and 24 GB graphics card. Running DART on less powerful devices may cause issues with loading high-resolution atlases and significantly slow processing time.
Note: the installation processes will install the CPU platform version of PyTorch. To enable GPU-acceleration via CUDA, reinstall PyTorch with the desired version and compute platform by following the directions here.
PIP Install
DART is available as a PIP-installable package. It is recommended to make a virtual environment or custom conda environment, and install DART within that environment. Once the custom environment has been set up, follow these steps:
- PIP install DART with
pip install dart-lmd - Download the atlases with
dart-download-atlases- Note: if this fails, refer to this section
- Run DART with
dart-gui
Cloning this repository
DART can also be installed by cloning the repository and downloading auxiliary files. This is ideal for editing or contributing to the software and is necessary for running unit tests via pytest. To succesfully install DART with this method follow these steps:
- Clone this repository using
git clone https://https://github.com/kebschulllab/Dissecting-Atlas-Registered-Tissue.git - Set up a conda environment by navigating into the repository folder and using
conda env create -f environment.yml - From the repository folder, download the atlases using
python -m dart.download_atlases- Note: if this fails, refer to this section
- To run the software as a user, call
python -m dartfrom the root directory of the repository.
Additional Features of Cloned Repository
Installing DART via repository cloning enables several additional tools for developers:
- The automated tests can be run with pytest using either
pytest --disable-warningsorpython -m pytest --disable-warnings. - Individual pages can be "demoed" by calling
python -m dart.demofrom the root directory of the repository. This opens up a GUI where users can select a specific page and view it without needing to proceed through the previous pages.
If Downloading the Atlases Fails
The atlases can be manually downloaded and extracted from the Zenodo archive. These atlases should be extracted to the folder dart/dart/atlases.
Note: The atlases folder should have 5 subfolders, and each of those should have 3 subfiles titled label.nrrd, reference.nrrd, and names_dict.csv.
Input Data
To use this tool, you will need to provide the following information:
- 3D Reference Atlas with cell stain and region annotations (included in installation if using Allen Brain Atlas)
- A folder containing cell stained images from LEICA LDM on slides
Usage
To use DART, please refer to our tutorial.
License
- DART's source code is available under the GNU General Public License version 3
- VisuAlign is developed by the Neural Systems Laboratory at the Institute of Basic Medical Sciences, University of Oslo, Norway and is licensed under the Creative Commons Attribution-NonCommercial-ShareAlike 4.0 License
Contributing
For support or issues, please open an “Issue” on our GitHub repository. To contribute to the software, please refer to our Contribution guidelines
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dart_lmd-1.2.0.tar.gz.
File metadata
- Download URL: dart_lmd-1.2.0.tar.gz
- Upload date:
- Size: 40.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2c5760f642613542a37e98e771fff6e0c3a256c44bdf5605fb7c3161d5044763
|
|
| MD5 |
08cee2a0833be8d6d2d63ca71d6e2c2a
|
|
| BLAKE2b-256 |
d0d7d5a8830aad349ebca70fa4c858945d1a8ae9dc29889747d7981512452a83
|
File details
Details for the file dart_lmd-1.2.0-py3-none-any.whl.
File metadata
- Download URL: dart_lmd-1.2.0-py3-none-any.whl
- Upload date:
- Size: 40.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
78318f7d9561c6ce282512809d69e2f26e9bd37a0c31e6dd10bb1930bb8b0611
|
|
| MD5 |
46933637a55d5ee63f5d1ec927c1280a
|
|
| BLAKE2b-256 |
c27fb86ee77c68d7c349d33fa67df669d0b942cfaddba65347df605dcd1a0da2
|










