Data Science Utils extends the Scikit-Learn API and Matplotlib API to provide simple methods that simplify tasks and visualizations for data science projects.
Project description
Data Science Utils: Frequently Used Methods for Data Science
Data Science Utils extends the Scikit-Learn API and Matplotlib API to provide simple methods that simplify tasks and visualizations for data science projects.
Code Examples and Documentation
This README provides several examples of the package's capabilities. For complete documentation, including more methods and additional examples, please visit: https://datascienceutils.readthedocs.io/en/stable/
The API of the package is built to work with the Scikit-Learn API and Matplotlib API.
Metrics
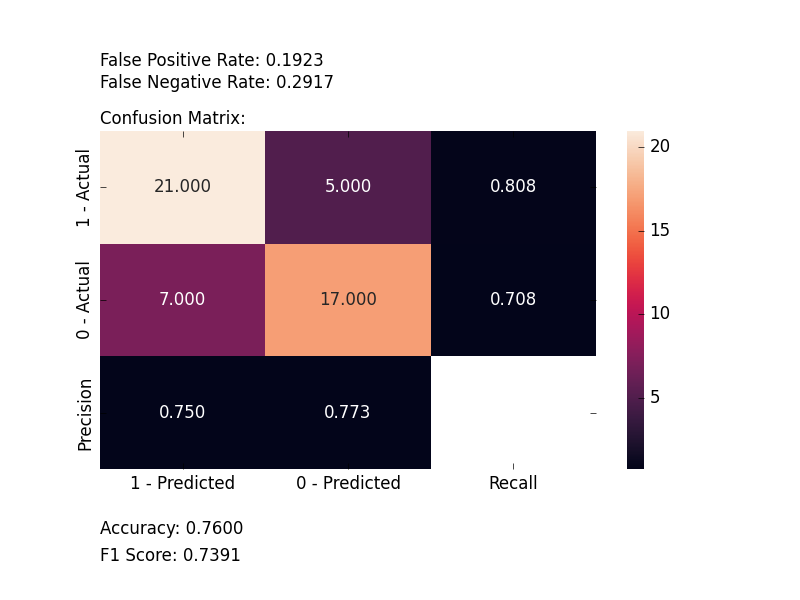
Plot Confusion Matrix
Computes and plots a confusion matrix, False Positive Rate, False Negative Rate, Accuracy, and F1 score of a classification.
from ds_utils.metrics import plot_confusion_matrix
plot_confusion_matrix(y_test, y_pred, [0, 1, 2])
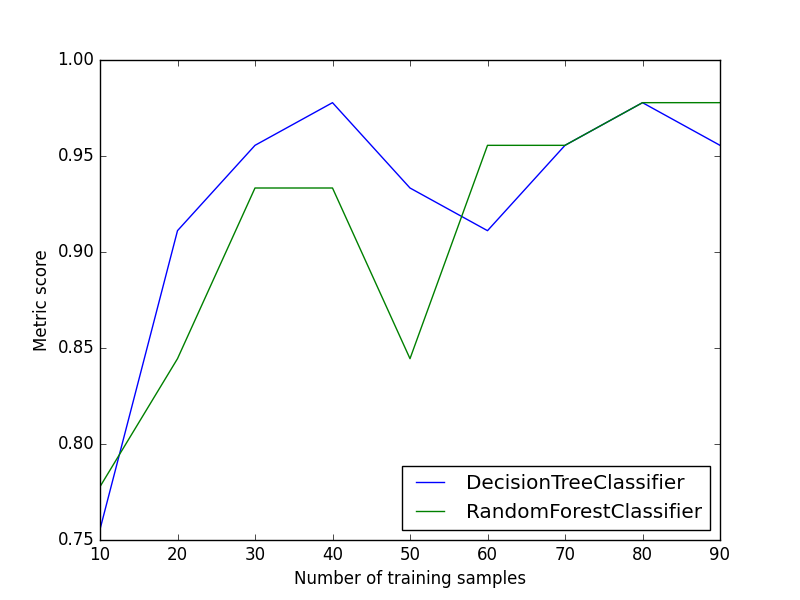
Plot Metric Growth per Labeled Instances
Receives train and test sets, and plots the given metric change with an increasing number of trained instances.
from ds_utils.metrics import plot_metric_growth_per_labeled_instances
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
plot_metric_growth_per_labeled_instances(
x_train, y_train, x_test, y_test,
{
"DecisionTreeClassifier": DecisionTreeClassifier(random_state=0),
"RandomForestClassifier": RandomForestClassifier(random_state=0, n_estimators=5)
}
)
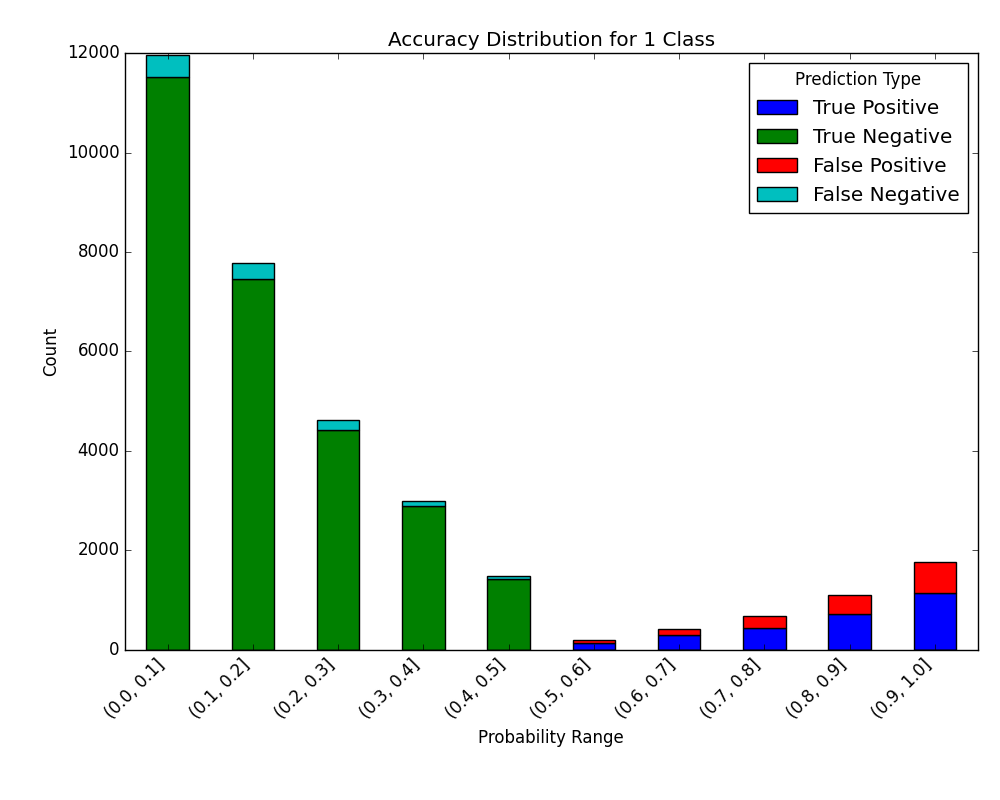
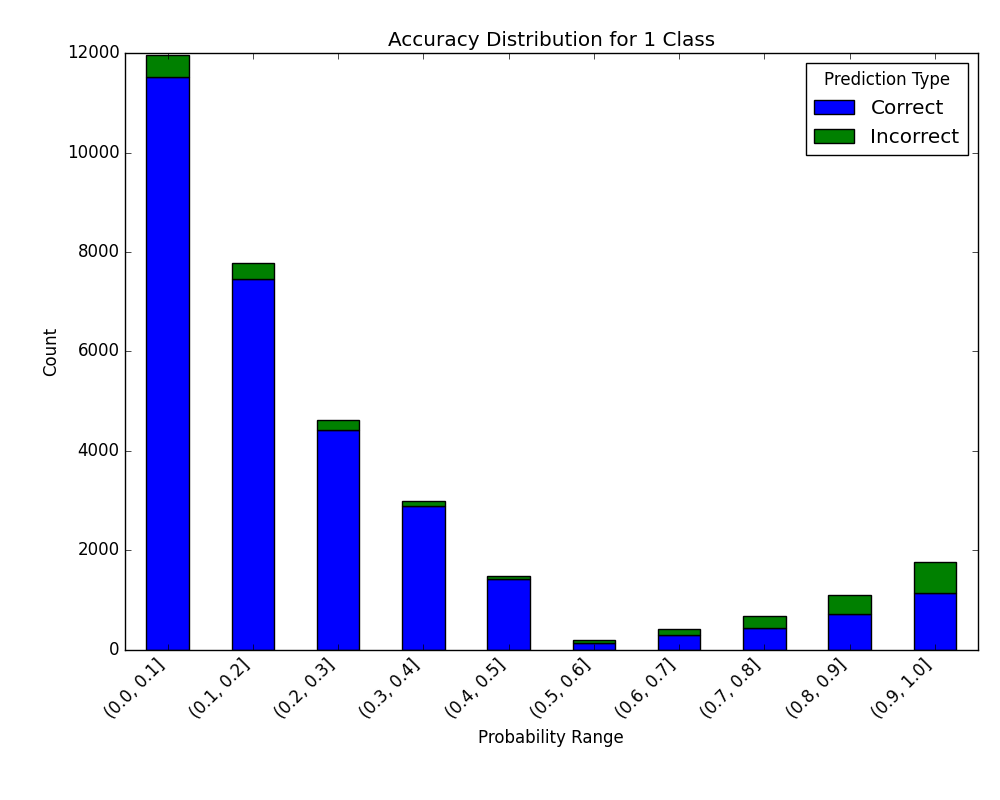
Visualize Accuracy Grouped by Probability
Receives test true labels and classifier probability predictions, divides and classifies the results, and finally plots a stacked bar chart with the results. Original code
from ds_utils.metrics import visualize_accuracy_grouped_by_probability
visualize_accuracy_grouped_by_probability(
test["target"],
1,
classifier.predict_proba(test[selected_features]),
display_breakdown=False
)
Without breakdown:
With breakdown:
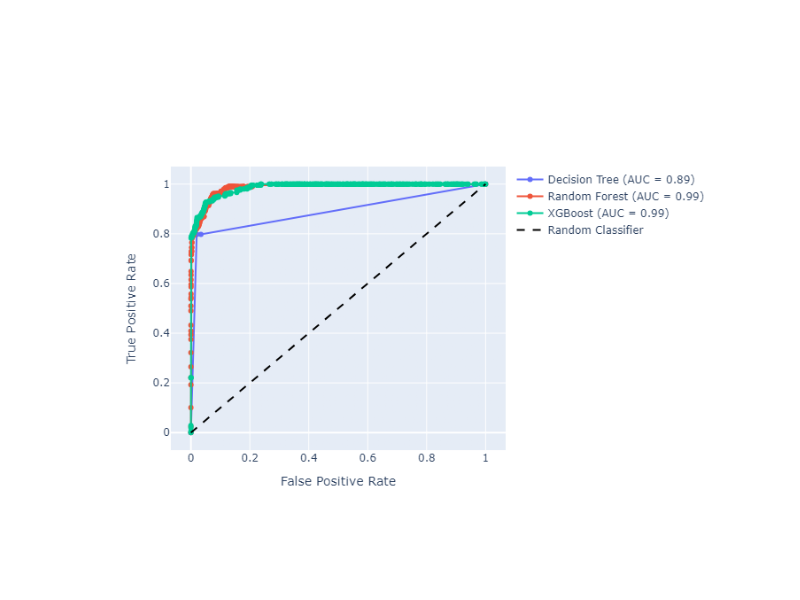
Receiver Operating Characteristic (ROC) Curve with Probabilities (Thresholds) Annotations
Plot ROC curves with threshold annotations for multiple classifiers, using plotly as a backend.
from ds_utils.metrics import plot_roc_curve_with_thresholds_annotations
classifiers_names_and_scores_dict = {
"Decision Tree": tree_clf.predict_proba(X_test)[:, 1],
"Random Forest": rf_clf.predict_proba(X_test)[:, 1],
"XGBoost": xgb_clf.predict_proba(X_test)[:, 1]
}
fig = plot_roc_curve_with_thresholds_annotations(
y_true,
classifiers_names_and_scores_dict,
positive_label=1
)
fig.show()
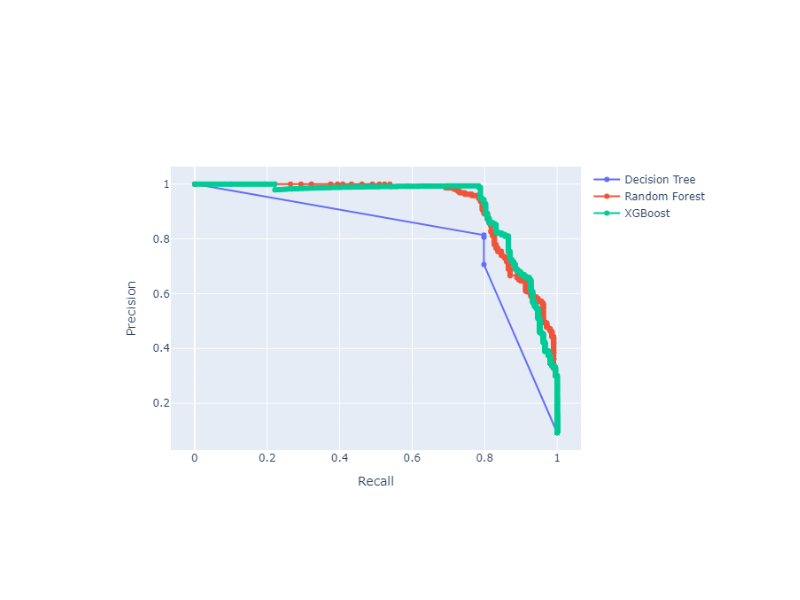
Precision-Recall Curve with Probabilities (Thresholds) Annotations
Plot Precision-Recall curves with threshold annotations for multiple classifiers, using plotly as a backend.
from ds_utils.metrics import plot_precision_recall_curve_with_thresholds_annotations
classifiers_names_and_scores_dict = {
"Decision Tree": tree_clf.predict_proba(X_test)[:, 1],
"Random Forest": rf_clf.predict_proba(X_test)[:, 1],
"XGBoost": xgb_clf.predict_proba(X_test)[:, 1]
}
fig = plot_precision_recall_curve_with_thresholds_annotations(
y_true,
classifiers_names_and_scores_dict,
positive_label=1
)
fig.show()
Preprocess
Visualize Feature
Receives a feature and visualizes its values on a graph:
- If the feature is float, the method plots the distribution plot.
- If the feature is datetime, the method plots a line plot of progression over time.
- If the feature is object, categorical, boolean, or integer, the method plots a count plot (histogram).
from ds_utils.preprocess import visualize_feature
visualize_feature(X_train["feature"])
| Feature Type | Plot |
|---|---|
| Float | 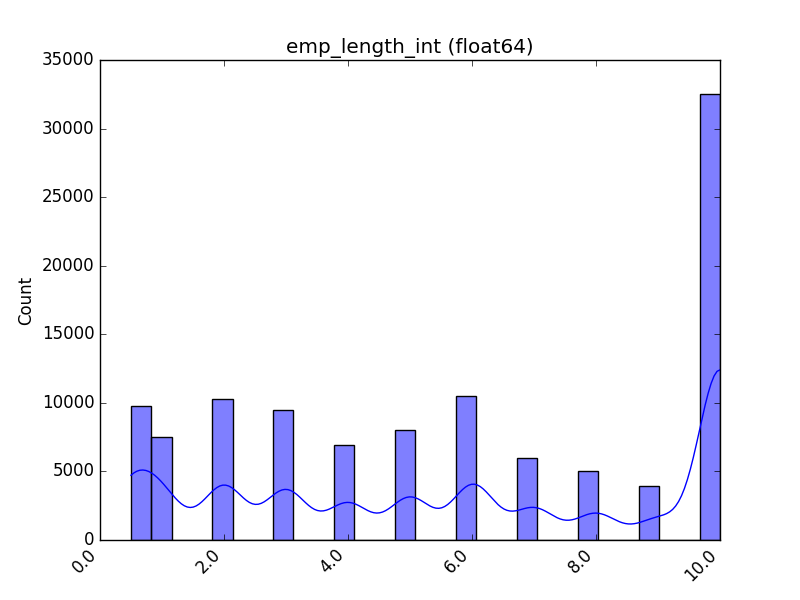 |
| Integer | 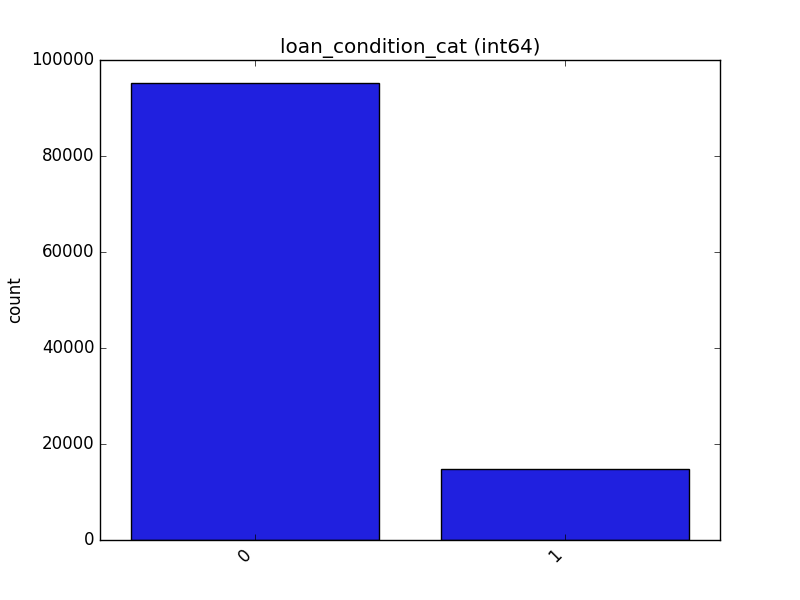 |
| Datetime | 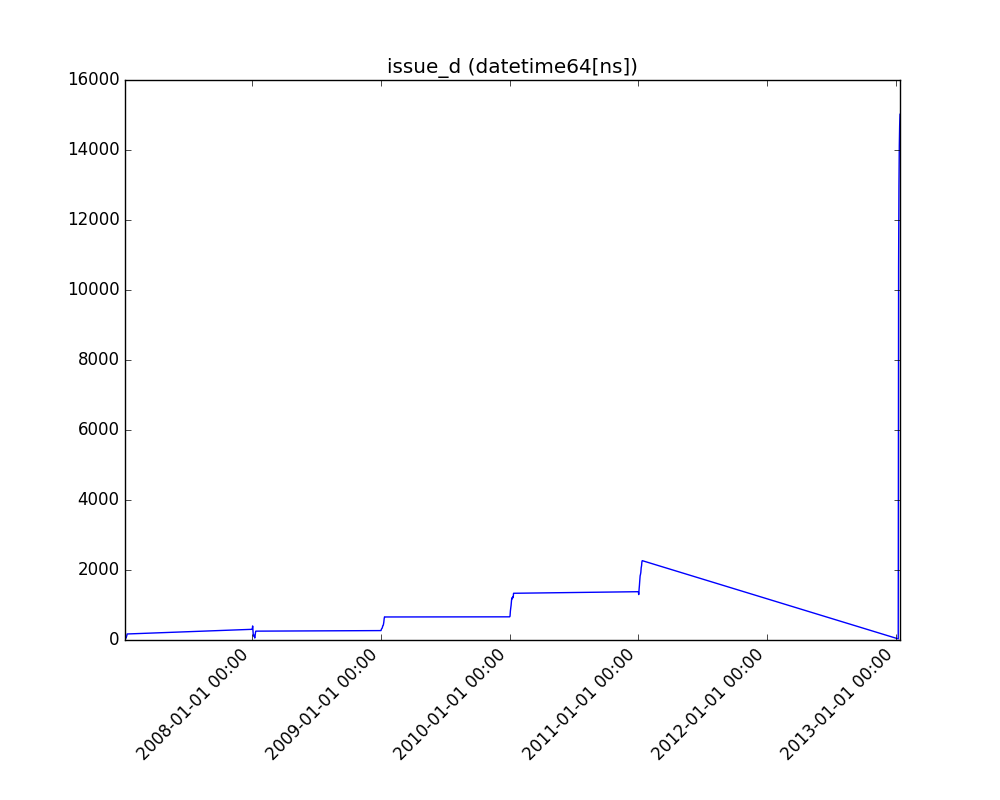 |
| Category / Object | 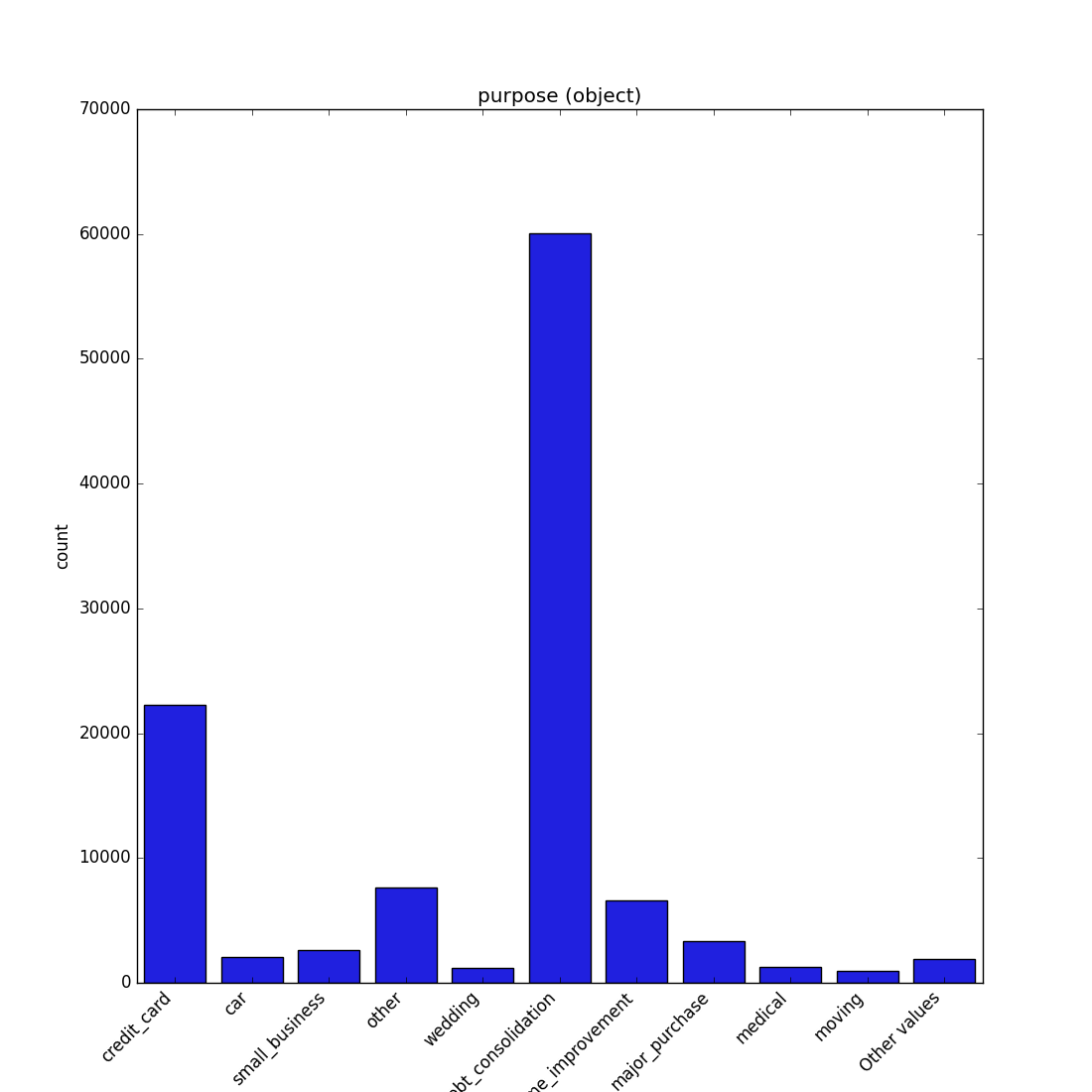 |
| Boolean | 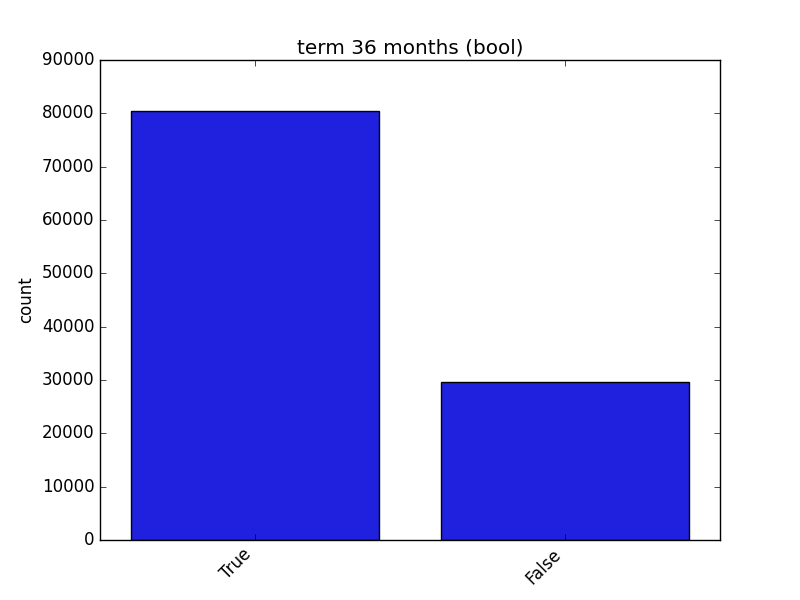 |
Get Correlated Features
Calculate which features are correlated above a threshold and extract a data frame with the correlations and correlation to the target feature.
from ds_utils.preprocess import get_correlated_features
correlations = get_correlated_features(correlation_matrix, features, target)
| level_0 | level_1 | level_0_level_1_corr | level_0_target_corr | level_1_target_corr |
|---|---|---|---|---|
| income_category_Low | income_category_Medium | 1.0 | 0.1182165609358650 | 0.11821656093586504 |
| term_ 36 months | term_ 60 months | 1.0 | 0.1182165609358650 | 0.11821656093586504 |
| interest_payments_High | interest_payments_Low | 1.0 | 0.1182165609358650 | 0.11821656093586504 |
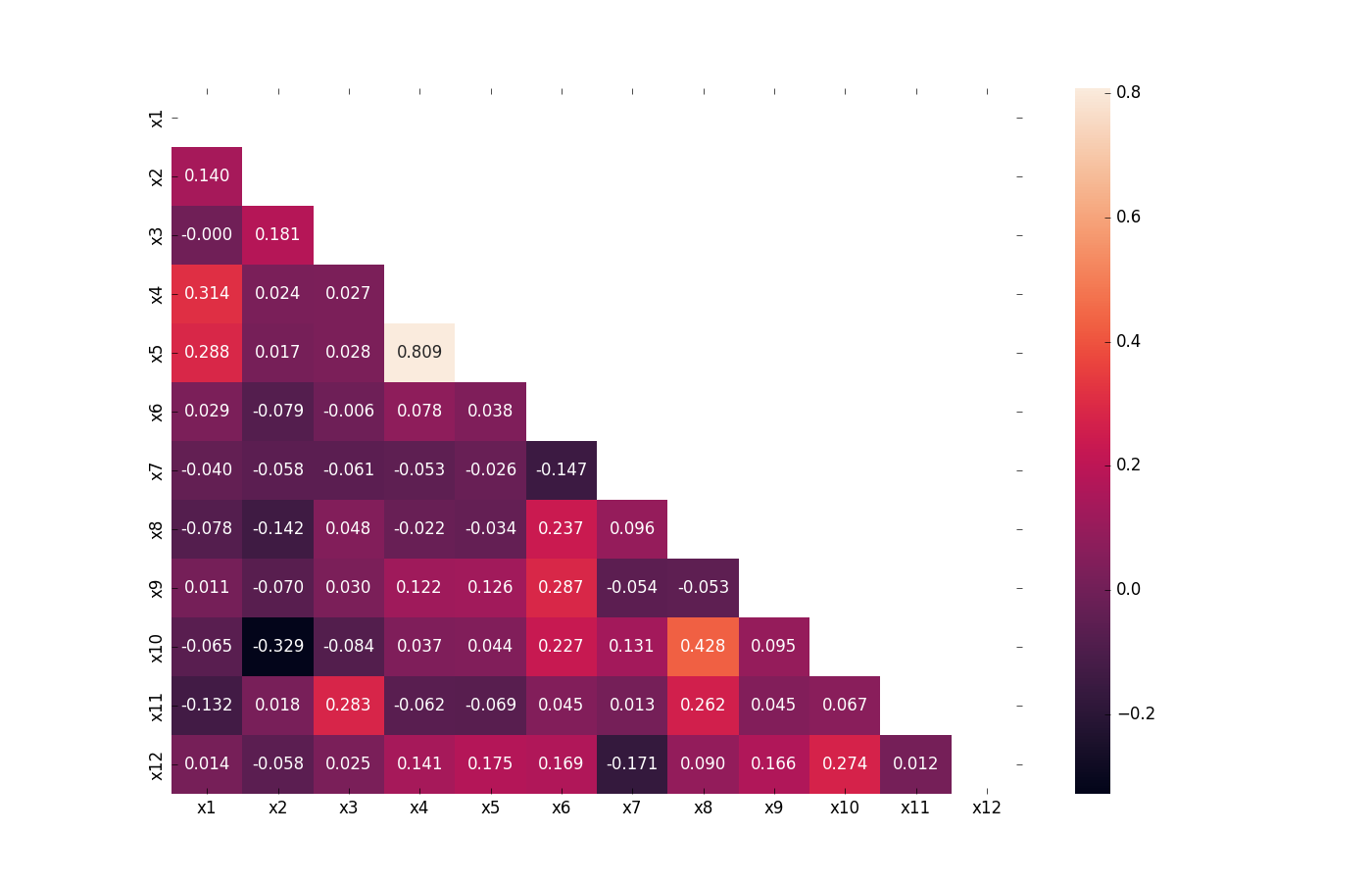
Visualize Correlations
Compute pairwise correlation of columns, excluding NA/null values, and visualize it with a heat map. Original code
from ds_utils.preprocess import visualize_correlations
visualize_correlations(correlation_matrix)
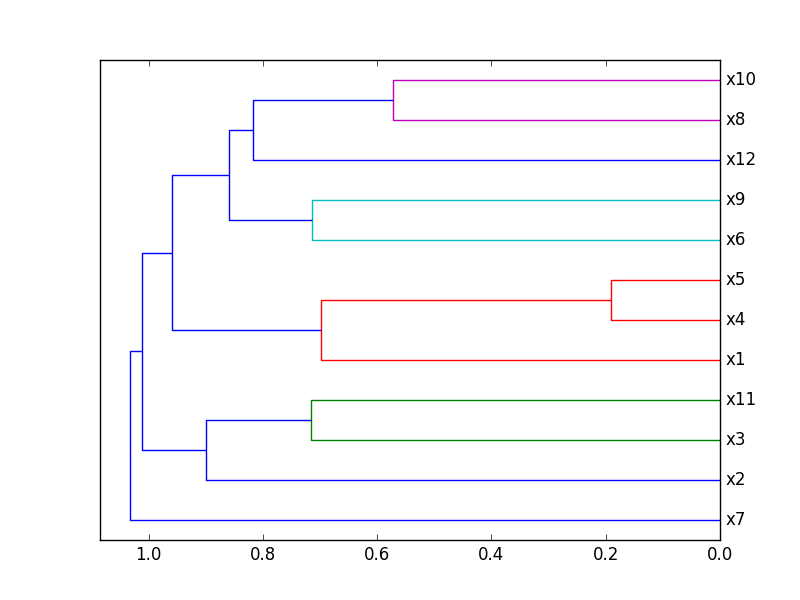
Plot Correlation Dendrogram
Plot a dendrogram of a correlation matrix. This chart hierarchically displays the most correlated variables by connecting them in a tree-like structure. The closer to the right that the connection is, the more correlated the features are. Original code
from ds_utils.preprocess import plot_correlation_dendrogram
plot_correlation_dendrogram(correlation_matrix)
Plot Features' Interaction
Plots the joint distribution between two features:
- If both features are either categorical, boolean, or object, the method plots the shared histogram.
- If one feature is either categorical, boolean, or object and the other is numeric, the method plots a boxplot chart.
- If one feature is datetime and the other is numeric or datetime, the method plots a line plot graph.
- If one feature is datetime and the other is either categorical, boolean, or object, the method plots a violin plot (combination of boxplot and kernel density estimate).
- If both features are numeric, the method plots a scatter graph.
from ds_utils.preprocess import plot_features_interaction
plot_features_interaction(data, "feature_1", "feature_2")
| Numeric | Categorical | Boolean | Datetime | |
|---|---|---|---|---|
| Numeric | 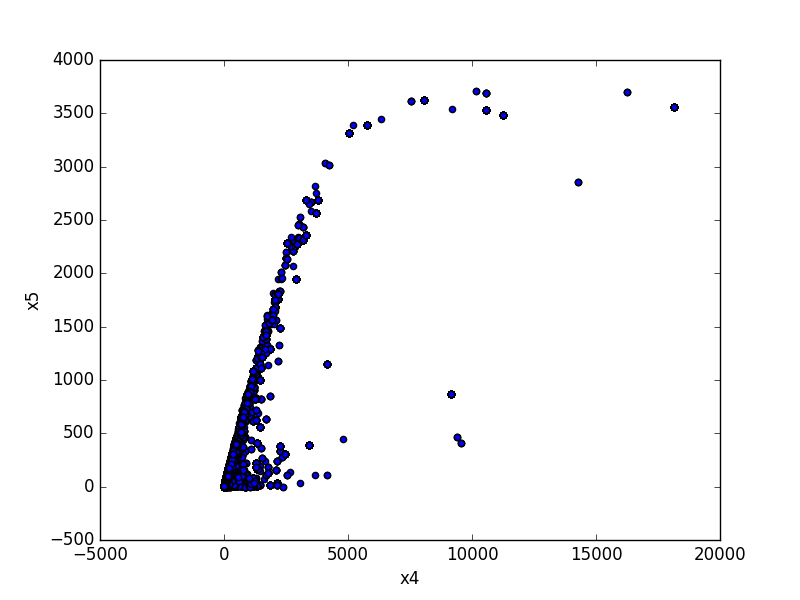 |
|||
| Categorical | 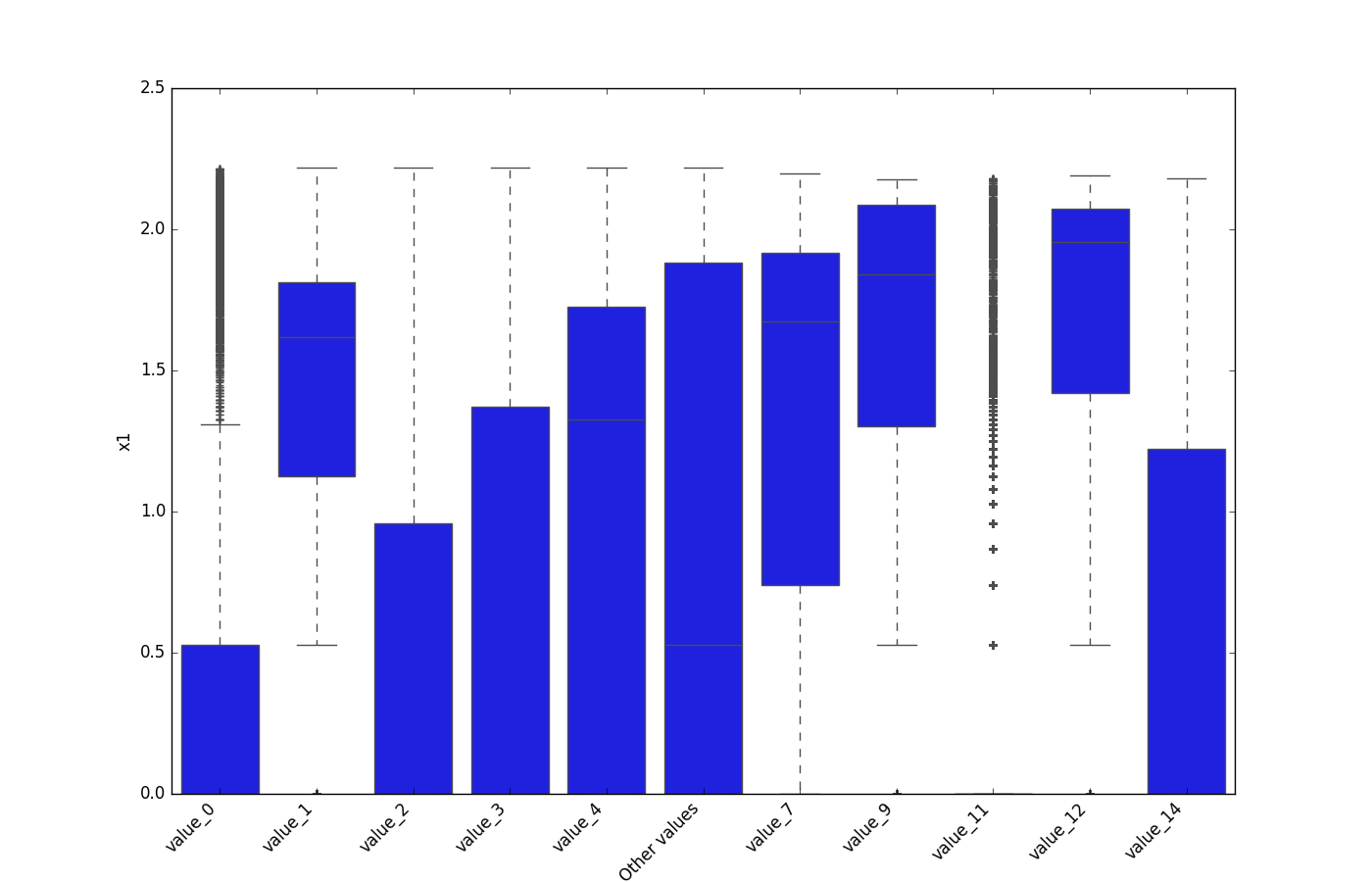 |
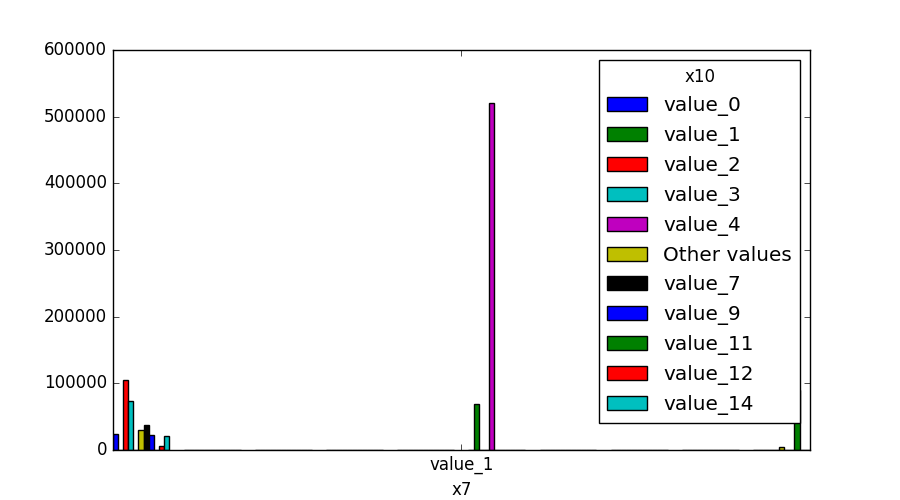 |
||
| Boolean | 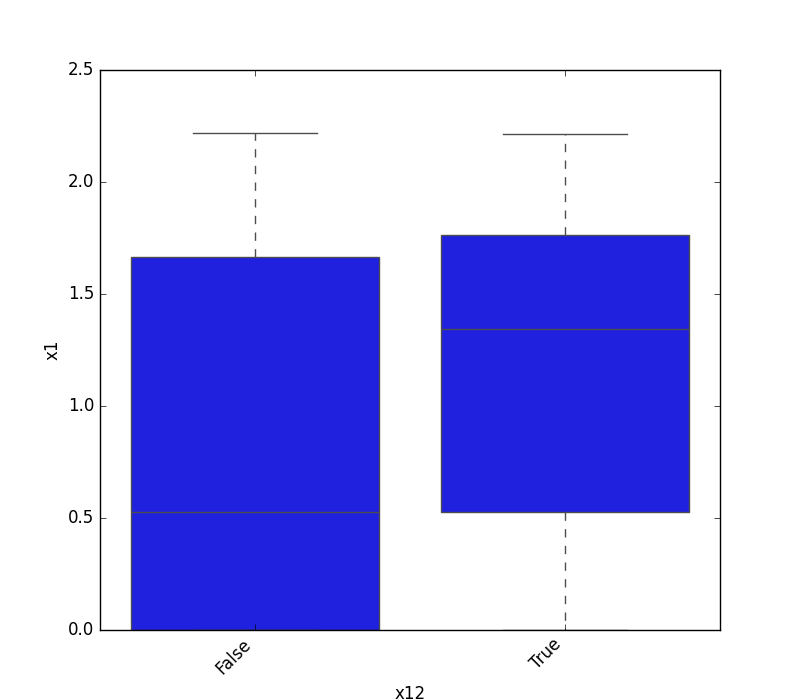 |
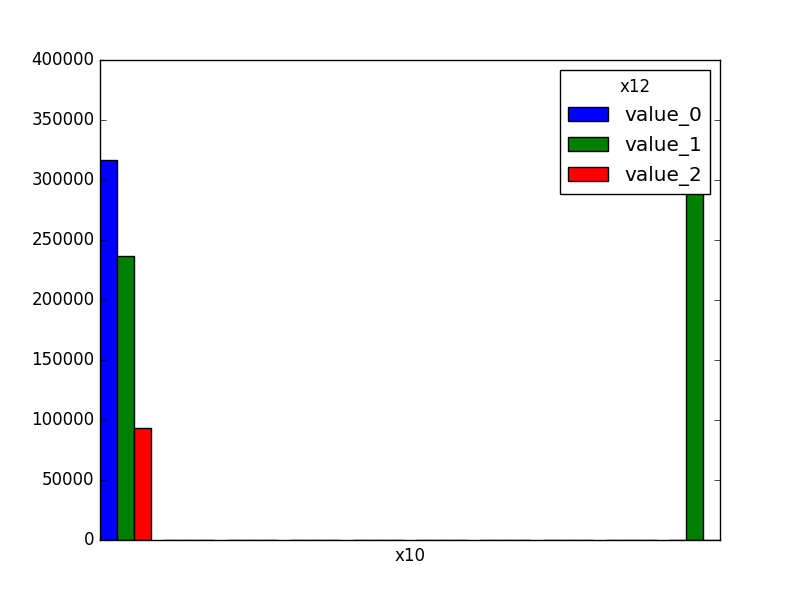 |
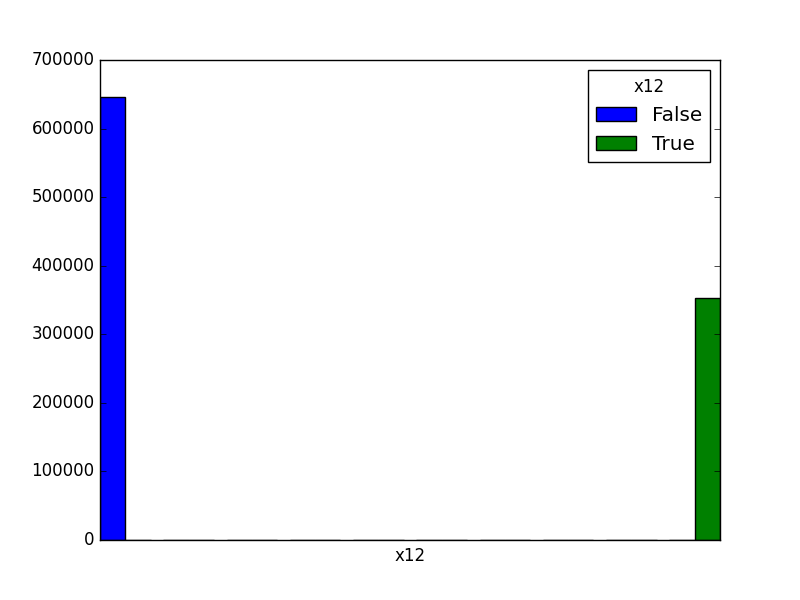 |
|
| Datetime | 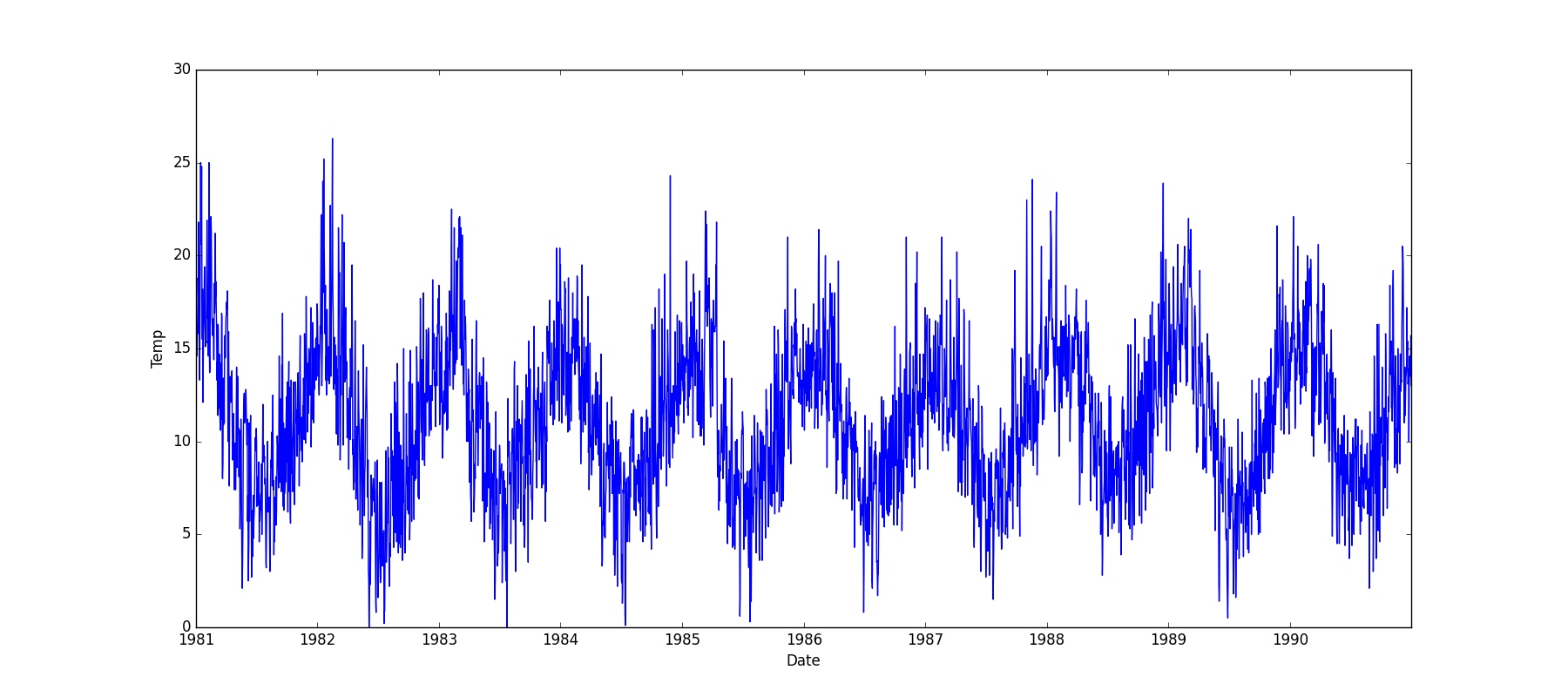 |
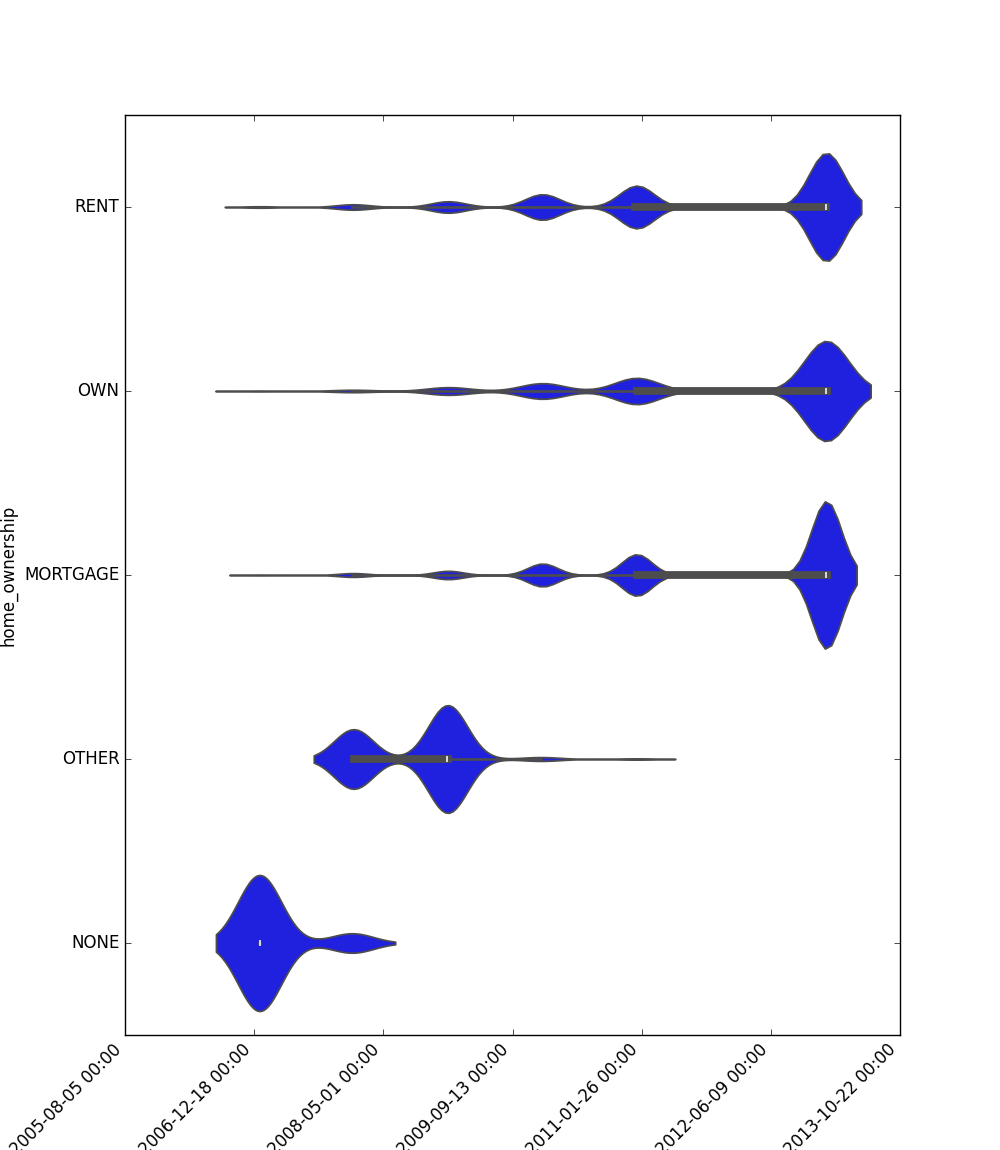 |
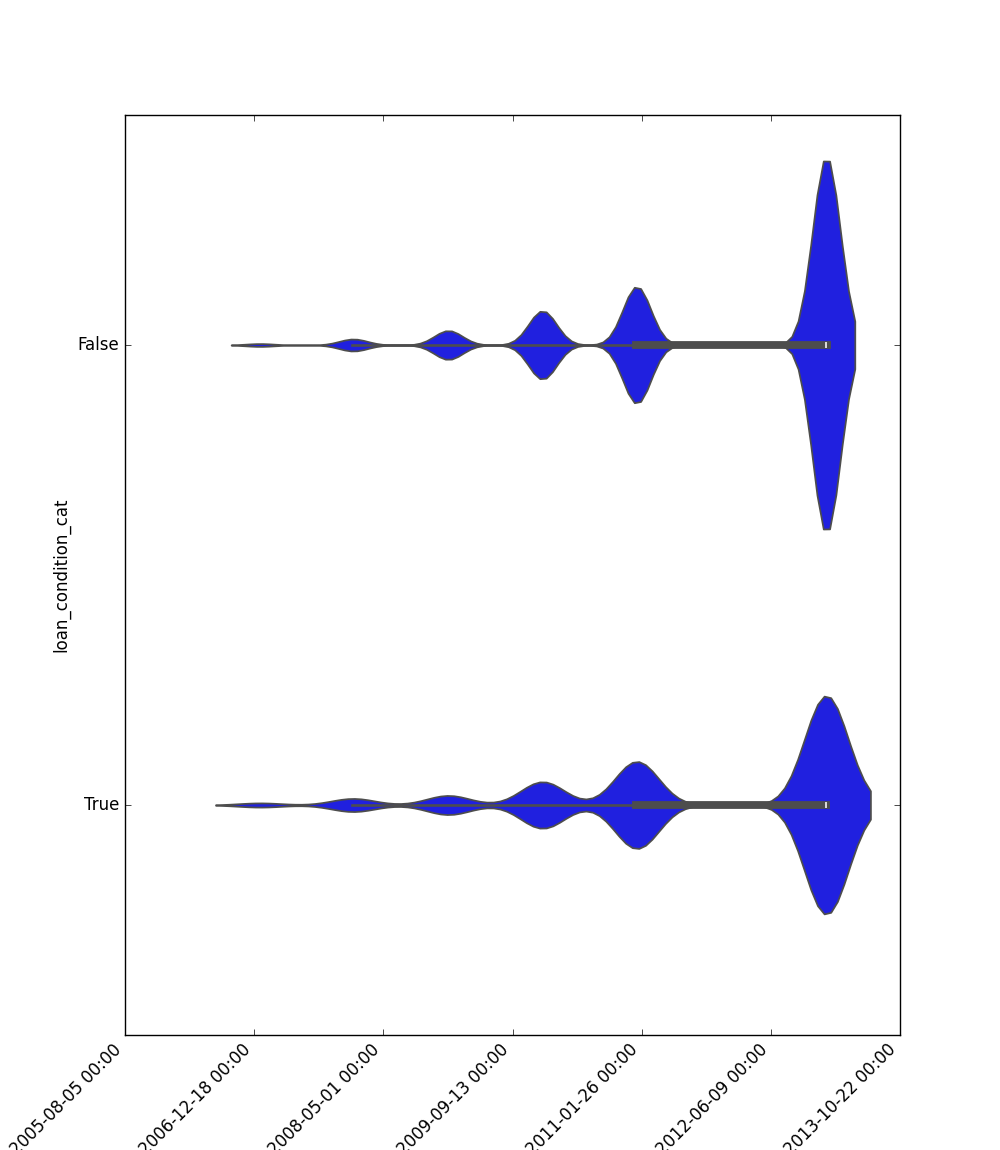 |
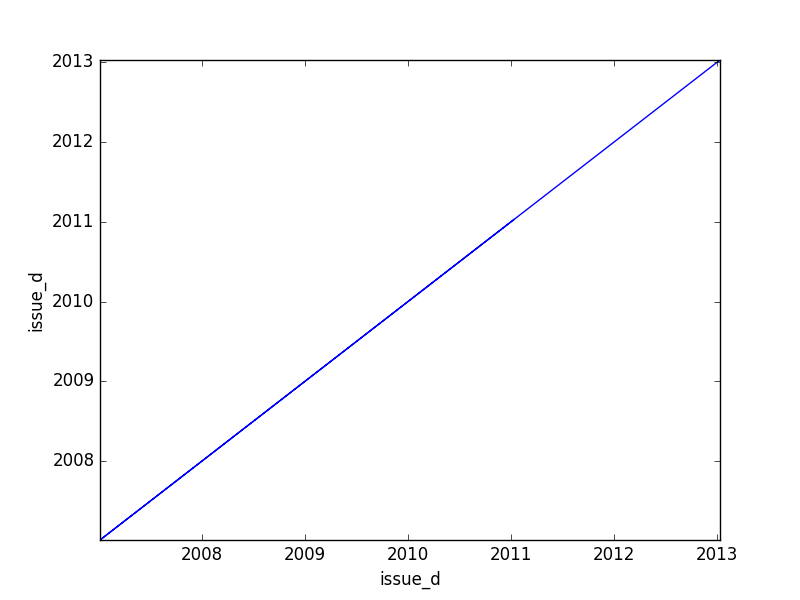 |
Extract Statistics DataFrame per Label
This method calculates comprehensive statistical metrics for numerical features grouped by label values. Use this when you want to:
- Analyze how a numerical feature's distribution varies across different categories
- Detect potential patterns or anomalies in feature behavior per group
- Generate detailed statistical summaries for reporting or analysis
- Understand the relationship between features and target variables
from ds_utils.preprocess import extract_statistics_dataframe_per_label
extract_statistics_dataframe_per_label(
df=df,
feature_name='amount',
label_name='category'
)
| category | count | null_count | mean | min | 1_percentile | 5_percentile | 25_percentile | median | 75_percentile | 95_percentile | 99_percentile | max |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A | 2 | 0 | 150.0 | 100 | 100.0 | 100.0 | 100.0 | 150.0 | 200.0 | 200.0 | 200.0 | 200.0 |
| B | 2 | 0 | 225.0 | 150 | 150.0 | 150.0 | 150.0 | 225.0 | 300.0 | 300.0 | 300.0 | 300.0 |
| C | 2 | 0 | 212.5 | 175 | 175.0 | 175.0 | 175.0 | 212.5 | 250.0 | 250.0 | 250.0 | 250.0 |
Compute Mutual Information
This method computes mutual information scores between features and a target label. Mutual information measures the mutual dependence between two variables - higher scores indicate stronger relationships between features and the target label. Features are automatically categorized as numerical or discrete and preprocessed accordingly.
Use this method when you want to:
- Identify which features have the strongest relationship with your target variable
- Perform feature selection based on statistical dependence
- Understand feature importance from an information theory perspective
- Compare the predictive value of different types of features
from ds_utils.preprocess import compute_mutual_information
# Compute mutual information scores for all features
mi_scores = compute_mutual_information(
df=df,
features=['feature1', 'feature2', 'feature3'],
label_col='target',
random_state=42
)
The result will be a DataFrame sorted by MI score (descending):
| feature_name | mi_score |
|---|---|
| feature1 | 0.245 |
| feature3 | 0.182 |
| feature2 | 0.091 |
Strings
Append Tags to Frame
This method extracts tags from a given field and appends them as new columns to the dataframe.
Consider a dataset that looks like this:
x_train:
| article_name | article_tags |
|---|---|
| 1 | ds,ml,dl |
| 2 | ds,ml |
x_test:
| article_name | article_tags |
|---|---|
| 3 | ds,ml,py |
Using this code:
import pandas as pd
from ds_utils.strings import append_tags_to_frame
x_train = pd.DataFrame([{"article_name": "1", "article_tags": "ds,ml,dl"},
{"article_name": "2", "article_tags": "ds,ml"}])
x_test = pd.DataFrame([{"article_name": "3", "article_tags": "ds,ml,py"}])
x_train_with_tags, x_test_with_tags = append_tags_to_frame(x_train, x_test, "article_tags", "tag_")
The result will be:
x_train_with_tags:
| article_name | tag_ds | tag_ml | tag_dl |
|---|---|---|---|
| 1 | 1 | 1 | 1 |
| 2 | 1 | 1 | 0 |
x_test_with_tags:
| article_name | tag_ds | tag_ml | tag_dl |
|---|---|---|---|
| 3 | 1 | 1 | 0 |
Extract Significant Terms from Subset
This method returns interesting or unusual occurrences of terms in a subset. It is based on the elasticsearch significant_text aggregation.
import pandas as pd
from ds_utils.strings import extract_significant_terms_from_subset
corpus = ['This is the first document.', 'This document is the second document.',
'And this is the third one.', 'Is this the first document?']
data_frame = pd.DataFrame(corpus, columns=["content"])
# Let's differentiate between the last two documents from the full corpus
subset_data_frame = data_frame[data_frame.index > 1]
terms = extract_significant_terms_from_subset(data_frame, subset_data_frame,
"content")
The output for terms will be the following table:
| third | one | and | this | the | is | first | document | second |
|---|---|---|---|---|---|---|---|---|
| 1.0 | 1.0 | 1.0 | 0.67 | 0.67 | 0.67 | 0.5 | 0.25 | 0.0 |
Unsupervised
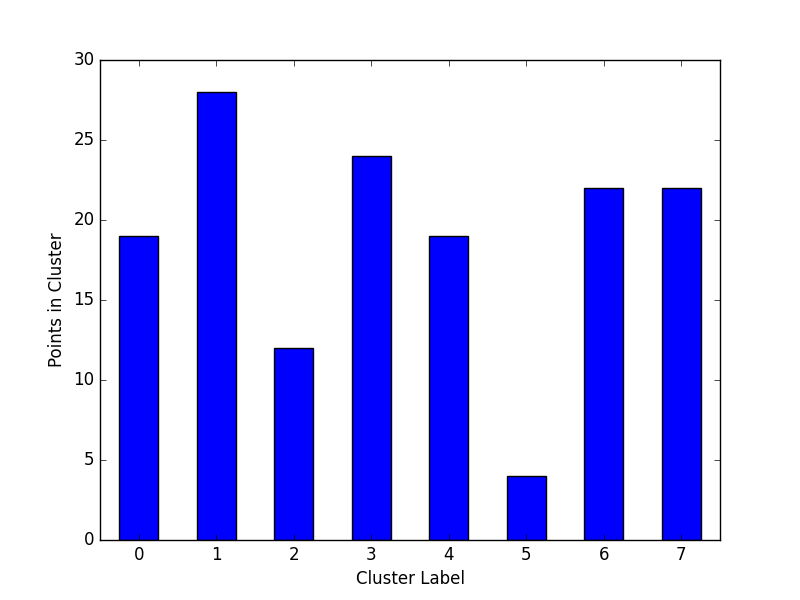
Cluster Cardinality
Cluster cardinality is the number of examples per cluster. This method plots the number of points per cluster as a bar chart.
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.cluster import KMeans
from ds_utils.unsupervised import plot_cluster_cardinality
data = pd.read_csv(path / to / dataset)
estimator = KMeans(n_clusters=8, random_state=42)
estimator.fit(data)
plot_cluster_cardinality(estimator.labels_)
plt.show()
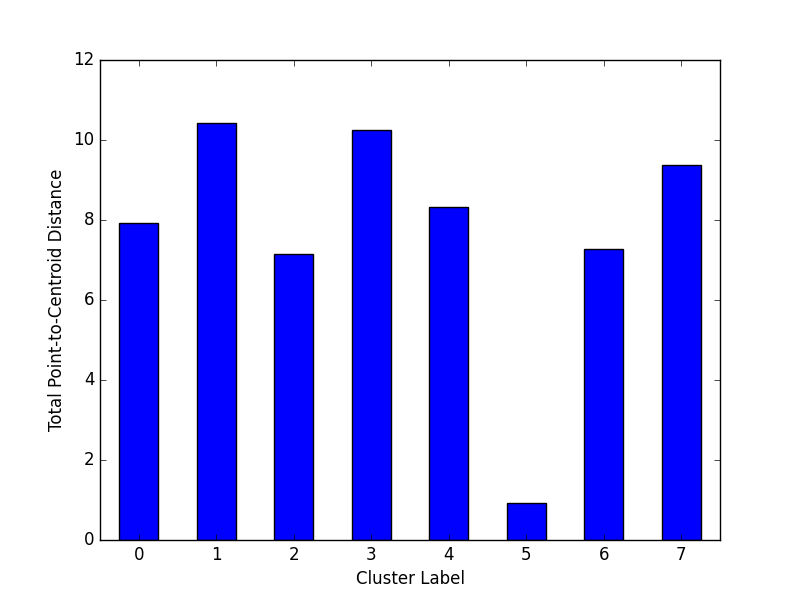
Plot Cluster Magnitude
Cluster magnitude is the sum of distances from all examples to the centroid of the cluster. This method plots the Total Point-to-Centroid Distance per cluster as a bar chart.
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.cluster import KMeans
from scipy.spatial.distance import euclidean
from ds_utils.unsupervised import plot_cluster_magnitude
data = pd.read_csv(path / to / dataset)
estimator = KMeans(n_clusters=8, random_state=42)
estimator.fit(data)
plot_cluster_magnitude(data, estimator.labels_, estimator.cluster_centers_, euclidean)
plt.show()
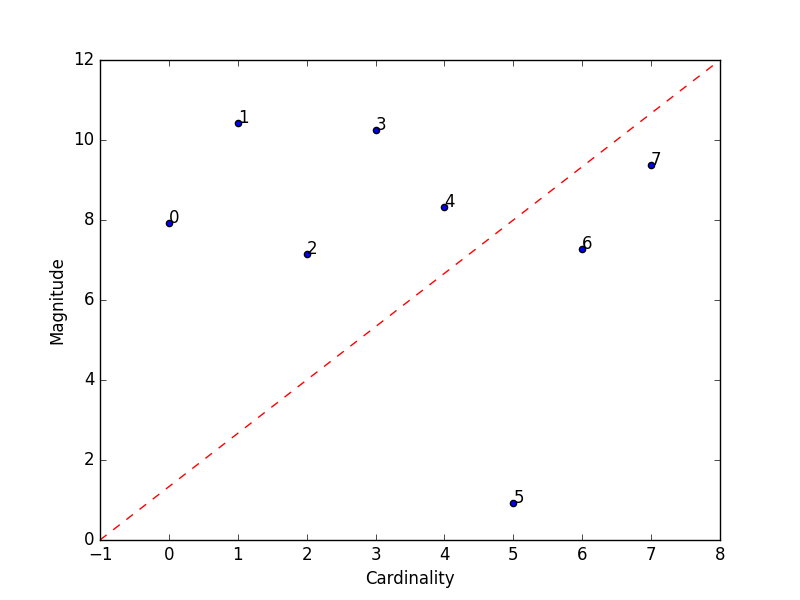
Magnitude vs. Cardinality
Higher cluster cardinality tends to result in a higher cluster magnitude, which intuitively makes sense. Clusters are considered anomalous when cardinality doesn't correlate with magnitude relative to the other clusters. This method helps find anomalous clusters by plotting magnitude against cardinality as a scatter plot.
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.cluster import KMeans
from scipy.spatial.distance import euclidean
from ds_utils.unsupervised import plot_magnitude_vs_cardinality
data = pd.read_csv(path / to / dataset)
estimator = KMeans(n_clusters=8, random_state=42)
estimator.fit(data)
plot_magnitude_vs_cardinality(data, estimator.labels_, estimator.cluster_centers_, euclidean)
plt.show()
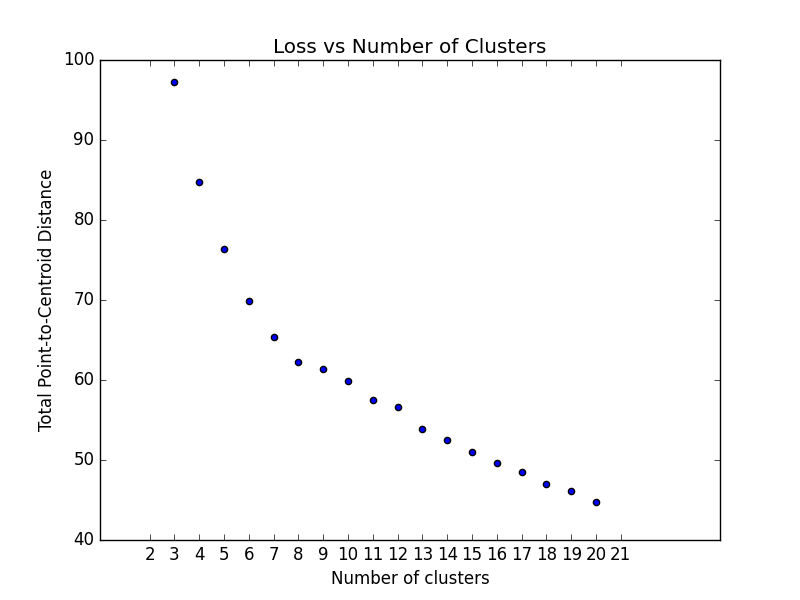
Optimum Number of Clusters
K-means clustering requires you to decide the number of clusters k beforehand. This method runs the KMeans algorithm
and
increases the cluster number at each iteration. The total magnitude or sum of distances is used as the loss metric.
Note: Currently, this method only works with sklearn.cluster.KMeans.
import pandas as pd
from matplotlib import pyplot as plt
from scipy.spatial.distance import euclidean
from ds_utils.unsupervised import plot_loss_vs_cluster_number
data = pd.read_csv(path / to / dataset)
plot_loss_vs_cluster_number(data, 3, 20, euclidean)
plt.show()
XAI (Explainable AI)
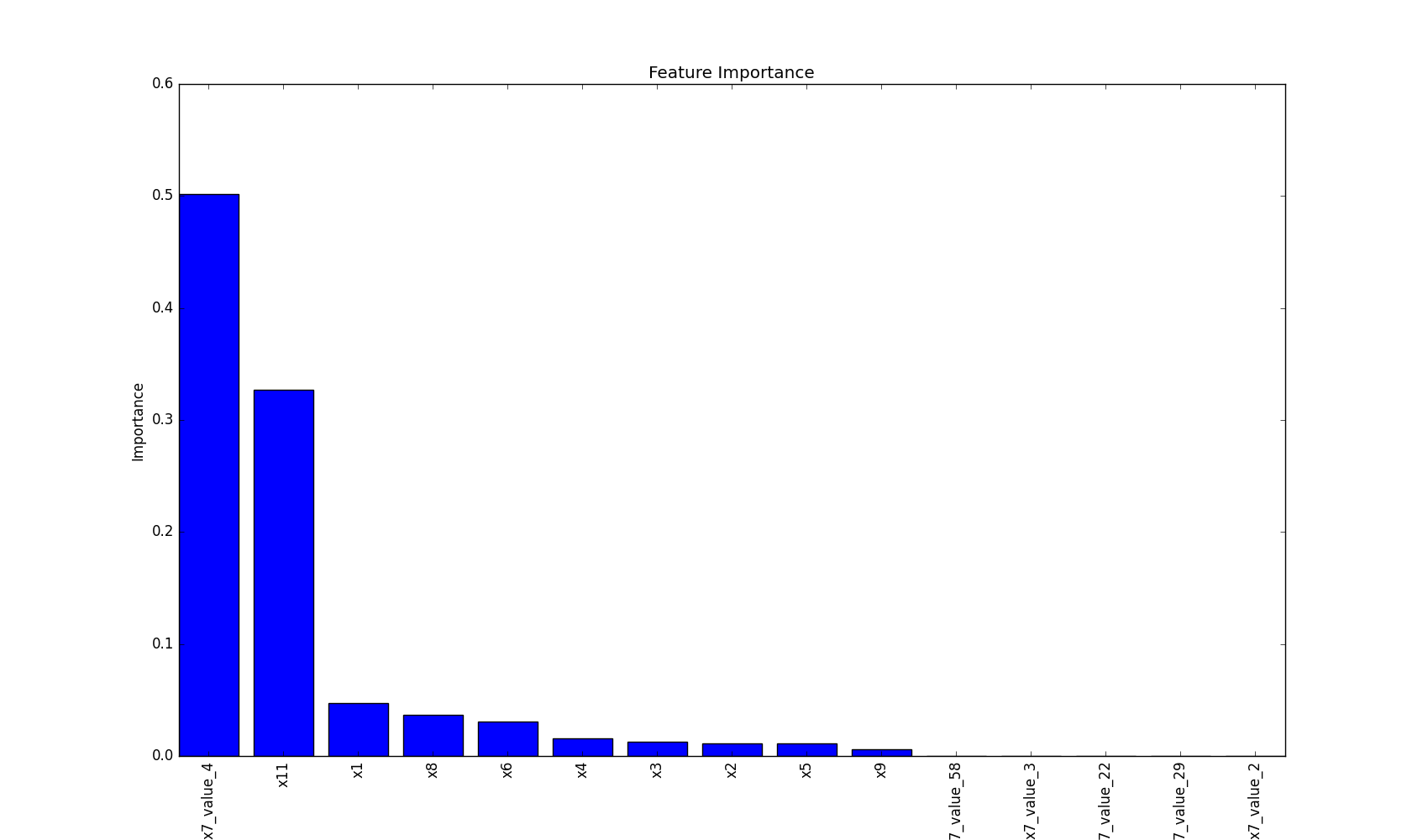
Plot Feature Importance
This method plots feature importance as a bar chart, helping to visualize which features have the most significant impact on the model's decisions.
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.tree import DecisionTreeClassifier
from ds_utils.xai import plot_features_importance
# Load the dataset
data = pd.read_csv(path / to / dataset)
target = data["target"]
features = data.columns.tolist()
features.remove("target")
# Train a decision tree classifier
clf = DecisionTreeClassifier(random_state=42)
clf.fit(data[features], target)
# Plot feature importance
plot_features_importance(features, clf.feature_importances_)
plt.show()
This visualization helps in understanding which features are most influential in the model's decision-making process, providing valuable insights for feature selection and model interpretation.
Explore More
Excited about what you've seen so far? There's even more to discover! Dive deeper into each module to unlock the full potential of DataScienceUtils:
-
Metrics - Powerful methods for calculating and visualizing algorithm performance evaluation. Gain insights into how your models are performing.
-
Preprocess - Essential data preprocessing techniques to prepare your data for training. Improve your model's input for better results.
-
Strings - Efficient methods for manipulating and processing strings in dataframes. Handle text data with ease.
-
Unsupervised - Tools for calculating and visualizing the performance of unsupervised models. Understand your clustering and dimensionality reduction results better.
-
XAI - Methods to help explain model decisions, making your AI more interpretable and trustworthy.
Each module is designed to streamline your data science workflow, providing you with the tools you need to preprocess data, train models, evaluate performance, and interpret results. Check out the detailed documentation for each module to see how DataScienceUtils can enhance your projects!
Contributing
We're thrilled that you're interested in contributing to Data Science Utils! Your contributions help make this project better for everyone. Whether you're a seasoned developer or just getting started, there's a place for you here.
How to Contribute
-
Find an area to contribute to: Check out our issues page for open tasks, or think of a feature you'd like to add.
-
Fork the repository: Make your own copy of the project to work on.
-
Create a branch: Make your changes in a new git branch.
-
Make your changes: Add your improvements or fixes. We appreciate:
- Bug reports and fixes
- Feature requests and implementations
- Documentation improvements
- Performance optimizations
- User experience enhancements
-
Test your changes: Ensure your code works as expected and doesn't introduce new issues.
-
Submit a pull request: Open a PR with a clear title and description of your changes.
Coding Guidelines
We follow the Python Software Foundation Code of Conduct and the Matplotlib Coding Guidelines. Please adhere to these guidelines in your contributions.
Getting Help
If you're new to open source or need any help, don't hesitate to ask questions in the issues section or reach out to the maintainers. We're here to help!
Code Quality
This project uses Ruff for linting and code formatting. Contributors are encouraged to use it to ensure code quality and consistency. You can check your code by running:
ruff check .
ruff format .
These checks are also part of our CI pipeline.
Why Contribute?
- Improve your skills: Gain experience working on a real-world project.
- Be part of a community: Connect with other developers and data scientists.
- Make a difference: Your contributions will help others in their data science journey.
- Get recognition: All contributors are acknowledged in our project.
Remember, no contribution is too small. Whether it's fixing a typo in documentation or adding a major feature, all contributions are valued and appreciated.
Thank you for helping make Data Science Utils better for everyone!
Installation Guide
Here are several ways to install the package:
1. Install from PyPI (Recommended)
The simplest way to install Data Science Utils and its dependencies is from PyPI using pip, Python's preferred package installer:
pip install data-science-utils
To upgrade Data Science Utils to the latest version, use:
pip install -U data-science-utils
2. Install from Source
If you prefer to install from source, you can clone the repository and install:
git clone https://github.com/idanmoradarthas/DataScienceUtils.git
cd DataScienceUtils
pip install .
Alternatively, you can install directly from GitHub using pip:
pip install git+https://github.com/idanmoradarthas/DataScienceUtils.git
3. Install using Anaconda
If you're using Anaconda, you can install using conda:
conda install idanmorad::data-science-utils
Note on Dependencies
Data Science Utils has several dependencies, including numpy, pandas, matplotlib, plotly and scikit-learn. These will be automatically installed when you install the package using the methods above.
Staying Updated
Data Science Utils is an active project that routinely publishes new releases with additional methods and improvements. We recommend periodically checking for updates to access the latest features and bug fixes.
If you encounter any issues during installation, please check our GitHub issues page or open a new issue for assistance.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file data_science_utils-1.9.0.tar.gz.
File metadata
- Download URL: data_science_utils-1.9.0.tar.gz
- Upload date:
- Size: 9.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b67119f5bc8060328cfb9c42be8552c0896efbdef662d5f1a68753471523f5fd
|
|
| MD5 |
11b8379cd1b0d208f58423ca1e62e079
|
|
| BLAKE2b-256 |
584709f78ed024788585aa196fc62d0ac8a9cc5dce44124e08c9db3bd7adde5a
|
File details
Details for the file data_science_utils-1.9.0-py3-none-any.whl.
File metadata
- Download URL: data_science_utils-1.9.0-py3-none-any.whl
- Upload date:
- Size: 29.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ed04d997a9823795601fb1a7e0054cceb414eb21c1e1f0689135796b9affb0ae
|
|
| MD5 |
70dbcc9dee2d307b27a8bdc071c86a66
|
|
| BLAKE2b-256 |
685effe42a75fdd496024e699f1c708314e5301bf7341b5600d0f0927ba281b5
|