Data Analysis for X-ray Spectroscopy
Project description
Data Analysis for X-ray Spectroscopy
Usage at the ESRF
In scripts
If you want to use the library in scripts you execute on the ESRF computing cluster, follow the steps below.
- Use a terminal to log in on one of the computing cluster front ends:
ssh -Y account@cluster-access. Theaccountcan be your personal SMIS account or the user experiment number. Enter the associated password when prompted. - Ask for resources:
srun --x11 --pty bash -l. This will give you an interactive shell. In addition, you can add--time=hh:mm:ssto specify the maximum time the resources will be available; by default, it will be 1 hour. - Load the spectroscopy environment module:
module load spectroscopy. The command loads an environment that contains the latest stable version ofdaxs. - Print the version of the library to test that everything went smoothly:
python -c "import daxs; print(daxs.__version__)".
If all goes well with the previous command and you don't get an error, you should be able to use the library in your scripts.
In Jupyter notebooks
You can also use the library in Jupyter notebooks.
- Connect to https://jupyter-slurm.esrf.fr.
- Select the
Spectroscopy (latest)in theJupyter environmentdrop-down menu. - Change the
Job durationin case you need to run your notebook for a longer time, than the default 1 hour. - Press
Startat the bottom of the page.
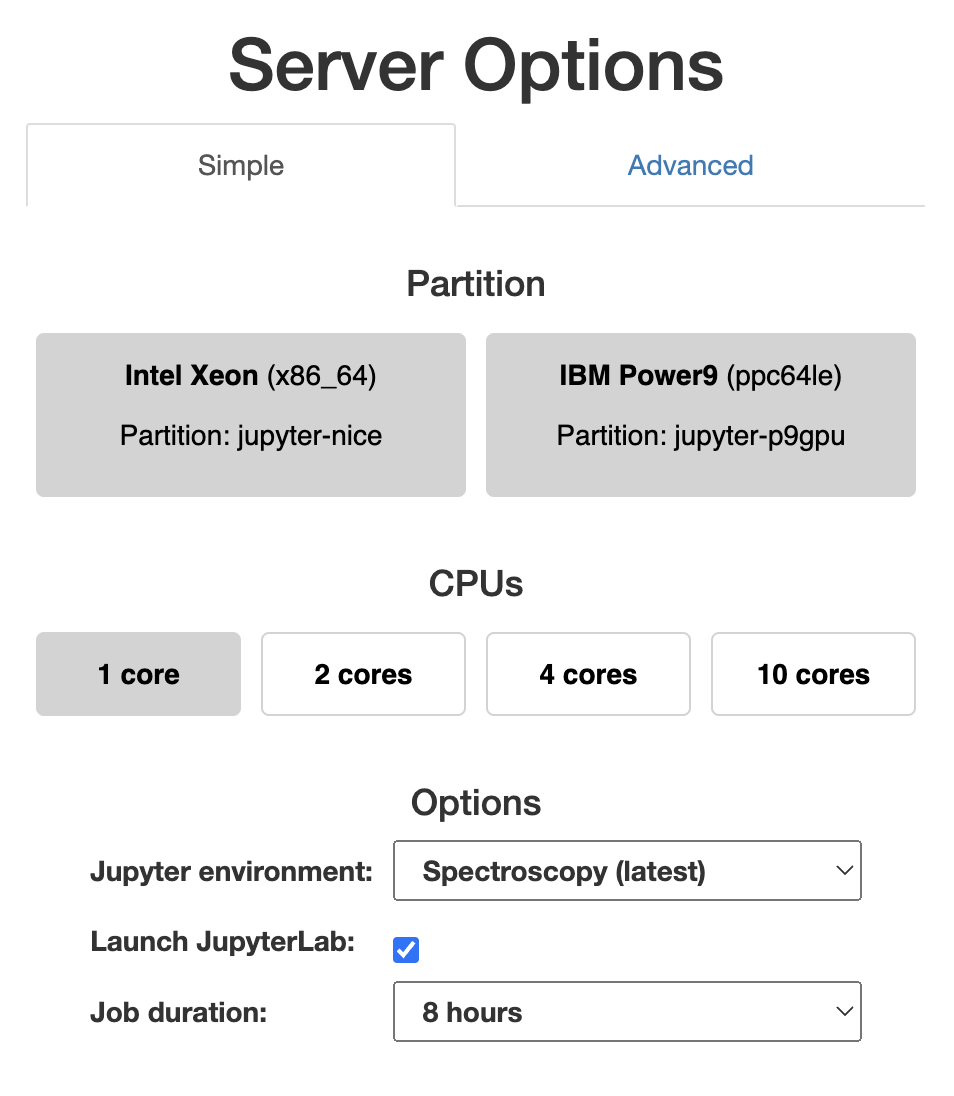
You can find more information about Jupyter at ESRF here.
While this simplifies the usage, you will not be able to add Python packages to the virtual environment. If you want to use additional packages not present in the environment, either open an issue here or install the library in your home directory, in a virtual environment (see below).
Local installation
The latest stable version of the library can be installed using:
pip install daxs
The development version can be installed using:
pip install [--ignore-installed] git+https://gitlab.esrf.fr/spectroscopy/daxs.git
The --ignore-installed argument is required to upgrade an existing installation.
Installing the library in a virtual environment is best to avoid messing up other Python packages. See the official documentation on how to create and use virtual environments.
Documentation
The documentation can be found at https://daxs.readthedocs.io.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file daxs-2026.2.tar.gz.
File metadata
- Download URL: daxs-2026.2.tar.gz
- Upload date:
- Size: 40.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.6.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6d75528a06aa4aefd0887efcc19e208dd0c7cf8327470cd284c508931525ccb0
|
|
| MD5 |
b41e8defdc776b6c1319d2dbe9ce7d47
|
|
| BLAKE2b-256 |
77eb92a72e413e60a818ab5278551b4d1d0bca4f9bcb794ead44755f74700ef2
|
File details
Details for the file daxs-2026.2-py3-none-any.whl.
File metadata
- Download URL: daxs-2026.2-py3-none-any.whl
- Upload date:
- Size: 44.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.6.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
04ea6a3676f34a53fea614192fb0def65344003bf6d7f98b60a454b274cdd5ef
|
|
| MD5 |
17355e3a193de81ac7193cac1673a7a0
|
|
| BLAKE2b-256 |
7e44870abfc80d918025b3635a19f599330eb621e5b46c83397184bc796b6070
|











