The official SetBERT implementation using the deepbio-toolkit.
Project description
SetBERT
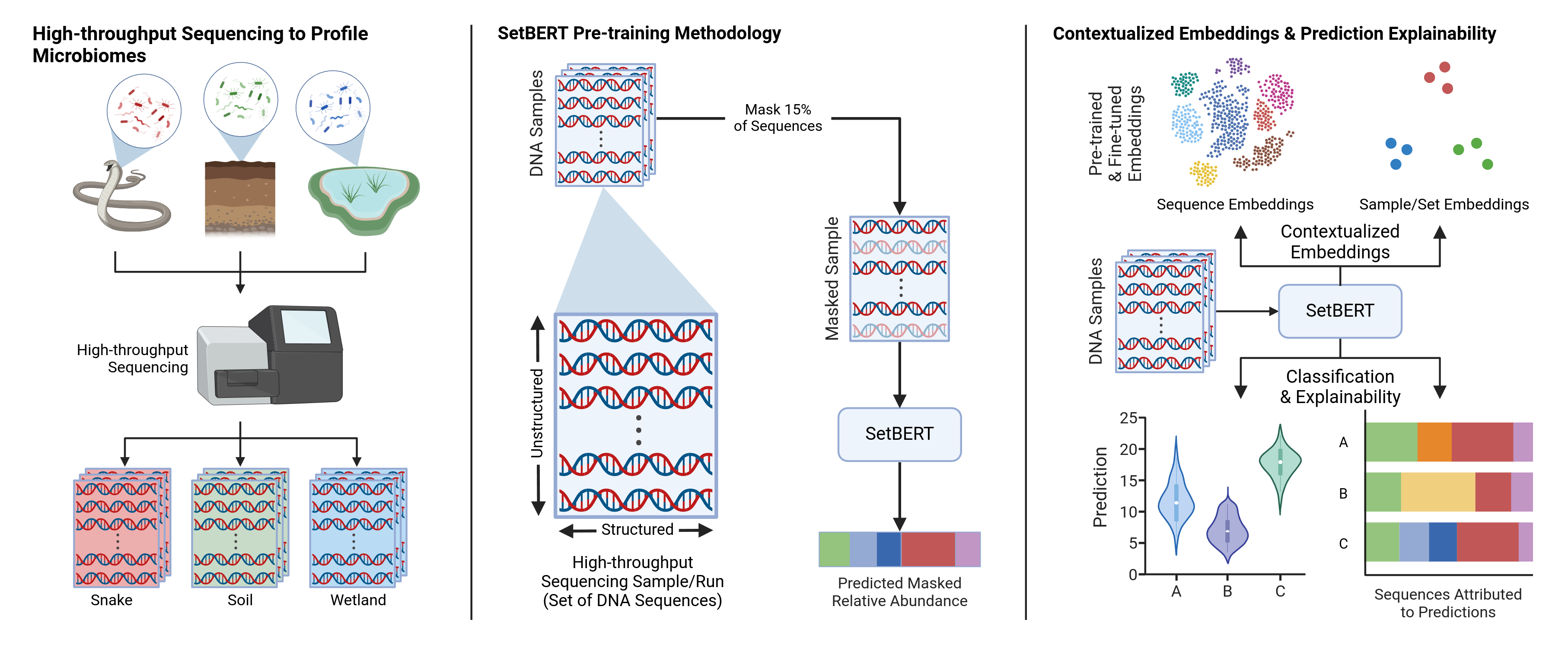
The official code repository for SetBERT: SetBERT: the deep learning platform for contextualized embeddings and explainable predictions from high-throughput sequencing
Quick Start
Installation from PyPI:
pip install dbtk-setbert
Download SetBERT pre-trained on the Qiita 16S platform (see Available Models for other options):
from setbert import SetBert
# Download the model
model = SetBert.from_pretrained("sirdavidludwig/setbert", revision="qiita-16s")
# Get the tokenizer
tokenizer = model.sequence_encoder.tokenizer
Example sample embedding
# Input sample
sequences = [
"ACTGCAG",
"TGACGTA",
"ATGACGA"
]
# Tokenize sequences in the sample
sequence_tokens = torch.stack([tokenizer(s) for s in sequences])
# Compute embeddings
output = model(sequence_tokens)
# Sample level representation
sample_embedding = output["class"]
# Contextualized sequence representations
sequence_embeddings = output["sequences"]
Available Models:
| Model Revision | Platform | Pre-training Dataset Description |
|---|---|---|
qiita-16s |
16S Amplicon | ~280k 16S amplicon samples from the Qiita platform |
Configuration
SetBERT embeds the DNA sequences in chunks using activation checkpointing. This chunk size is specified
by the sequence_encoder_chunk_size parameter in the SetBert.Config class and adjusted freely at any point.
# Set chunk size
model.config.sequence_encoder_chunk_size = 256 # default
# Remove chunking and embed all sequences in parallel
model.config.sequence_encoder_chunk_size = None
Manual Installation
git clone https://github.com/DLii-Research/setbert
pip install -e ./new-setbert
Citation
@article{ludwig_setbert_2025,
title = {{SetBERT}: the deep learning platform for contextualized embeddings and explainable predictions from high-throughput sequencing},
volume = {41},
issn = {1367-4811},
doi = {10.1093/bioinformatics/btaf370},
number = {7},
journal = {Bioinformatics},
author = {Ludwig, II, David W and Guptil, Christopher and Alexander, Nicholas R and Zhalnina, Kateryna and Wipf, Edi M -L and Khasanova, Albina and Barber, Nicholas A and Swingley, Wesley and Walker, Donald M and Phillips, Joshua L},
month = jul,
year = {2025},
}
Original Experiment Source Code
The original source code used to produce the models and experiments for the manuscript are available in the bioinformatics branch of this repository.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dbtk_setbert-1.0.2.tar.gz.
File metadata
- Download URL: dbtk_setbert-1.0.2.tar.gz
- Upload date:
- Size: 9.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1b4dc660b1517477f137ac6790276993867f2c0d4c0a7f17214d3ff63977bc00
|
|
| MD5 |
33d338c8c45243e79d1e7fa1afa605cb
|
|
| BLAKE2b-256 |
f75385288dd16d03b9238cb2acae3d7fa8dd899c28988246446ed2f1b50c1331
|
File details
Details for the file dbtk_setbert-1.0.2-py3-none-any.whl.
File metadata
- Download URL: dbtk_setbert-1.0.2-py3-none-any.whl
- Upload date:
- Size: 11.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
97f761f0313c64ed292281448a545a28e04a866e5cfd423c7515819a26a16b06
|
|
| MD5 |
f9d313a81cebab2499973d1608412df1
|
|
| BLAKE2b-256 |
bbe6b27b91afba4d7c18af879f8ec3fc655ad3d3b3339eb95f52441f04647ed0
|