Feature selection for preserving biological trajectories from single-cell data
Project description
DELVE
Dynamic selection of locally covarying features
Introduction
DELVE is an unsupervised feature selection method for identifying a representative subset of dynamically-expressed molecular features that recapitulate cellular trajectories from single-cell data (e.g. single-cell RNA sequencing, protein iterative immunofluorescence imaging). In contrast to previous work, DELVE uses a bottom-up approach to mitigate the effect of unwanted sources of feature variation confounding inference, and instead models cell states from dynamic feature modules that constitute core regulatory complexes. For more details on the method, please read the associated paper: Ranek JS, Stallaert W, Milner JJ, Redick M, Wolff SC, Beltran AS, Stanley N, and Purvis JE. DELVE: feature selection for preserving biological trajectories in single-cell data. Nature Communications. 2024.
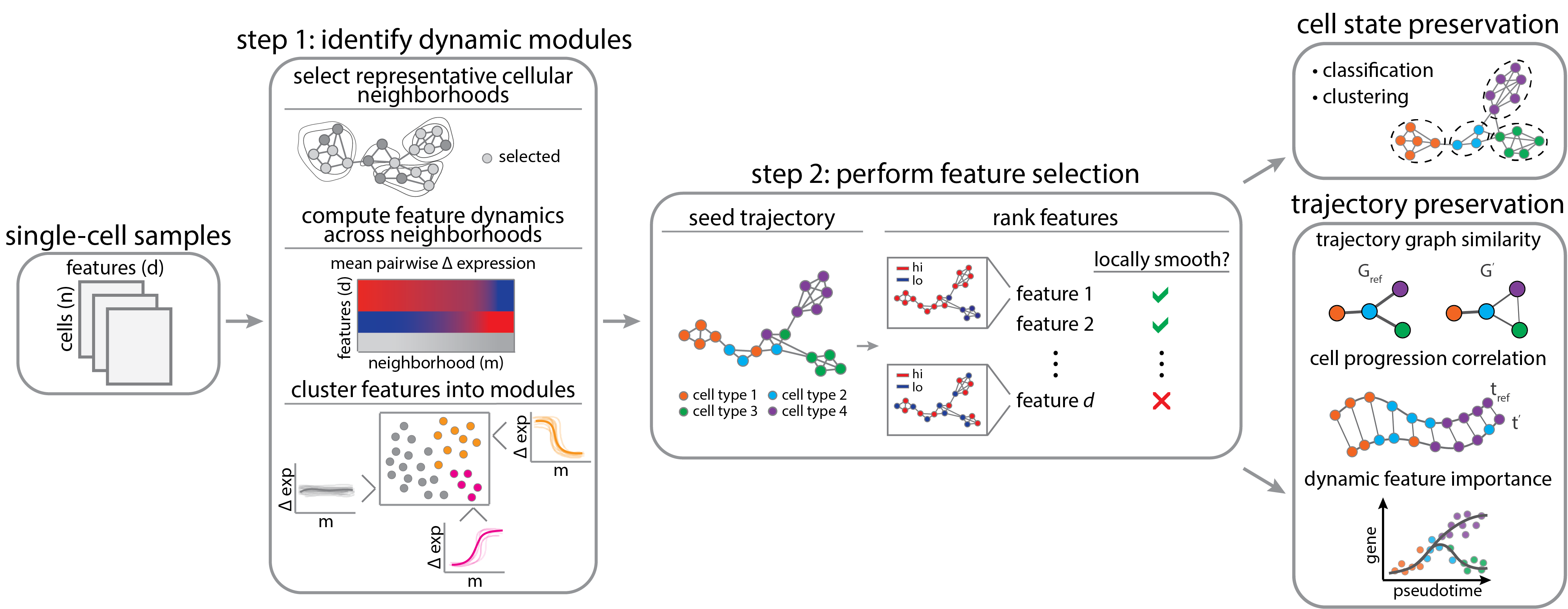
For a comparison of alternative feature selection methods and the overall benchmarking pipeline, please see delve_benchmark.
Installation
Dependencies
- Python == 3.9, sketchKH == 0.1.2, anndata >= 0.7.6, numpy == 1.26.4, scipy >= 1.7.1, pandas >= 1.5.2, umap-learn == 0.5.1, scikit-learn >= 0.23.2, scanpy == 1.8.1, tqdm
You can install the package and necessary dependencies with pip by,
pip install delve-fs
Alternatively, you can clone the git repository and install the necessary dependencies using the provided yml file. First clone the repository by,
git clone https://github.com/jranek/delve.git
Then change the working directory as,
cd delve
You can then create the conda environment using the provided yml file.
conda env create -f venv_delve.yml
Once the environment is created, you can activate it by,
conda activate venv_delve
Data access
You can download all of the preprocessed single-cell datasets (.h5ad files) from the Zenodo repository.
Example usage
To perform trajectory-preserving feature selection with DELVE, first read in a preprocessed .h5ad object. This .h5ad object contains a sample profiled with a single-cell technology (i.e. protein iterative indirect immunofluorescence imaging data).
import anndata
import os
adata = anndata.read_h5ad(os.path.join('data', 'adata_RPE.h5ad'))
Then simply perform DELVE feature selection by,
# Inputs:
# adata: annotated data object (dimensions = cells x features)
# k: number of nearest neighbors in the between-cell kNN affinity graph
# n_pcs: number of principal components. If None (default): will construct a between-cell affinity graph by computing pairwise Euclidean distances according to adata.X. Else: according to PCA of adata.X
# num_subsamples: number of representative cellular neighborhoods. Neighborhoods are subsampled using kernel herding sketching (see https://dl.acm.org/doi/abs/10.1145/3535508.3545539, https://github.com/CompCy-lab/SketchKH)
# n_clusters: number of feature modules
# n_random_state: number of random KMeans clustering initializations when identifying dynamic feature modules
# random_state: random state parameter
# n_jobs: number of tasks
# -----------------------
# Returns:
# delta_mean: average pairwise change in expression across prototypical cellular neighborhoods (dimensions = num_subsamples x features)
# modules: dataframe containing feature-cluster assignments and permutation p-values (dimensions = features x 2)
# ranked_features: ranked set of features that best preserve the local trajectory structure (dimensions = features x 1)
# -----------------------
from delve import *
delta_mean, modules, ranked_features = delve_fs(adata = adata, k = 10, num_subsamples = 1000, n_clusters = 5, random_state = 0, n_jobs = -1)
License
This software is licensed under the MIT license (https://opensource.org/licenses/MIT).
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file delve_fs-0.1.6.tar.gz.
File metadata
- Download URL: delve_fs-0.1.6.tar.gz
- Upload date:
- Size: 10.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.21
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1877dcfae2a7a67b6b835d5e68abaf9b32da56fd6e680dfaa9dd857d9cc519f1
|
|
| MD5 |
3cfa855ddaeb713a46b951cc4d58d27e
|
|
| BLAKE2b-256 |
62b1a2ba73d2c3d7558c86560f385f88811adc9da2816891d0e04f75c8d26cdf
|
File details
Details for the file delve_fs-0.1.6-py3-none-any.whl.
File metadata
- Download URL: delve_fs-0.1.6-py3-none-any.whl
- Upload date:
- Size: 9.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.21
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9206f2f3534024c2ae6dcbbecf846945443cf34650d1107ddc9f25befe88b904
|
|
| MD5 |
1dad296a241385cc4a806b87051a21a4
|
|
| BLAKE2b-256 |
32b4b13a93d326e5563c03cc144d9722c3dcadd581b4bba2eaac351277bf361b
|











