Statistical Analysis and Simulations for Plant Breeding Experiments
Project description
dgNova (Designs Nova)
A Python library for Statistical Analysis and Simulations of Plant Breeding Experiments, designed specifically for researchers and students who are new to programming but want to understand field design concepts through practical implementation.
Overview
dgNova ("Designs Nova") makes it easy to:
- Simulate field experiments with realistic spatial patterns
- Analyze unreplicated field trials using moving grid methods
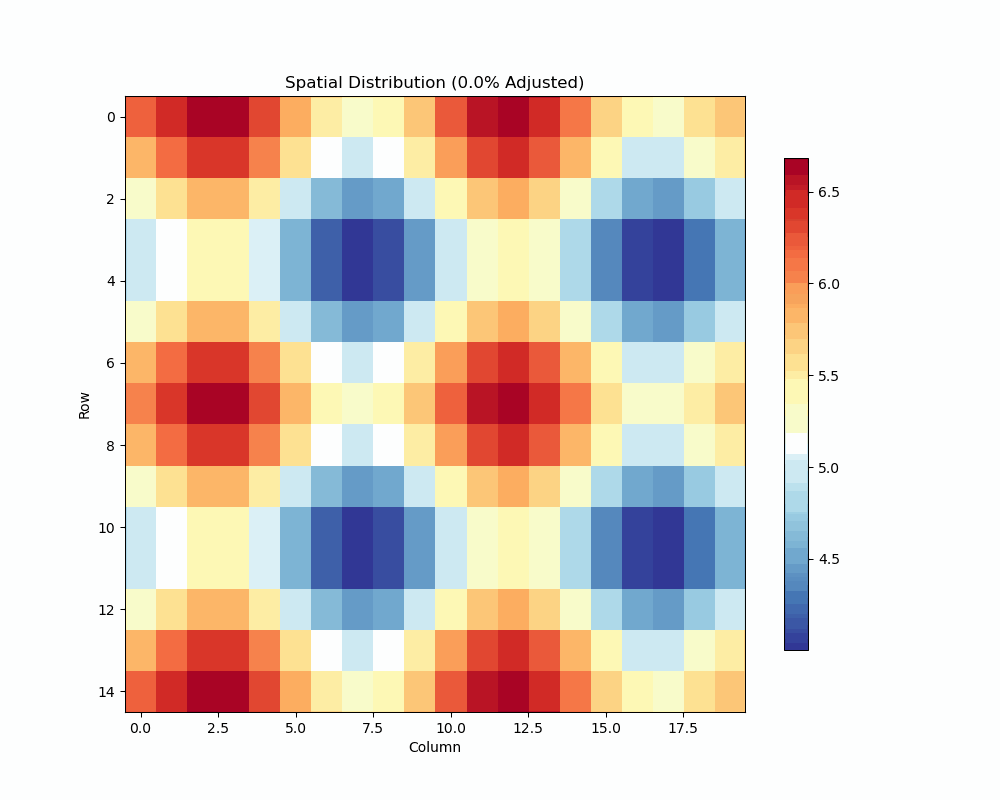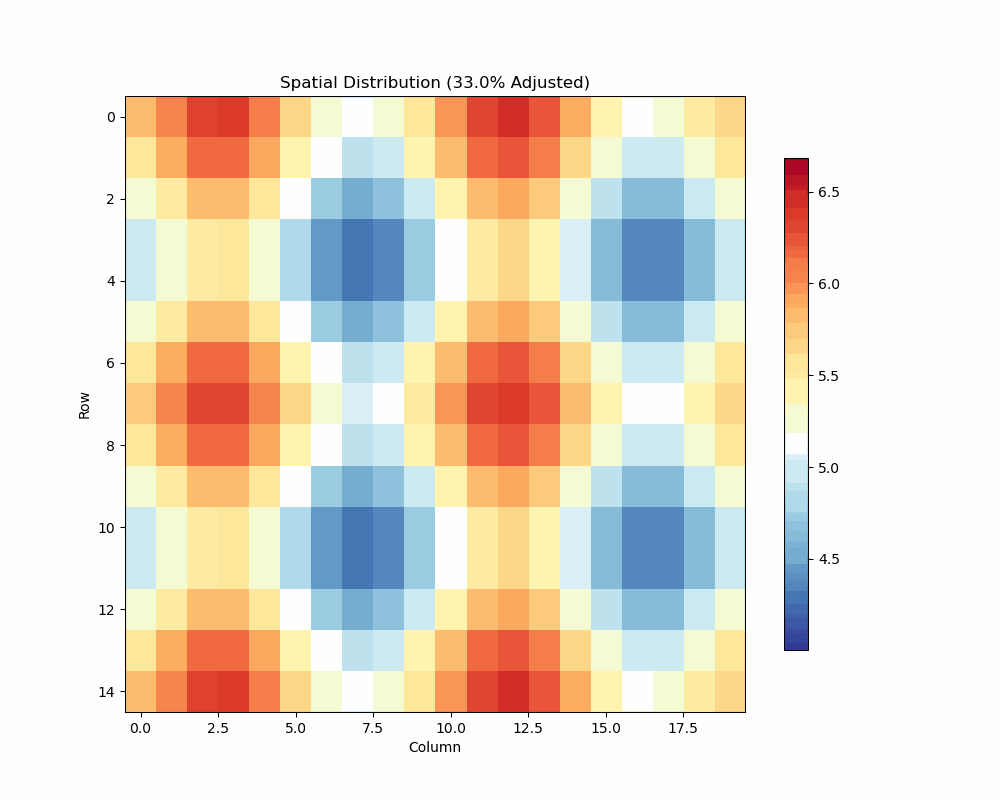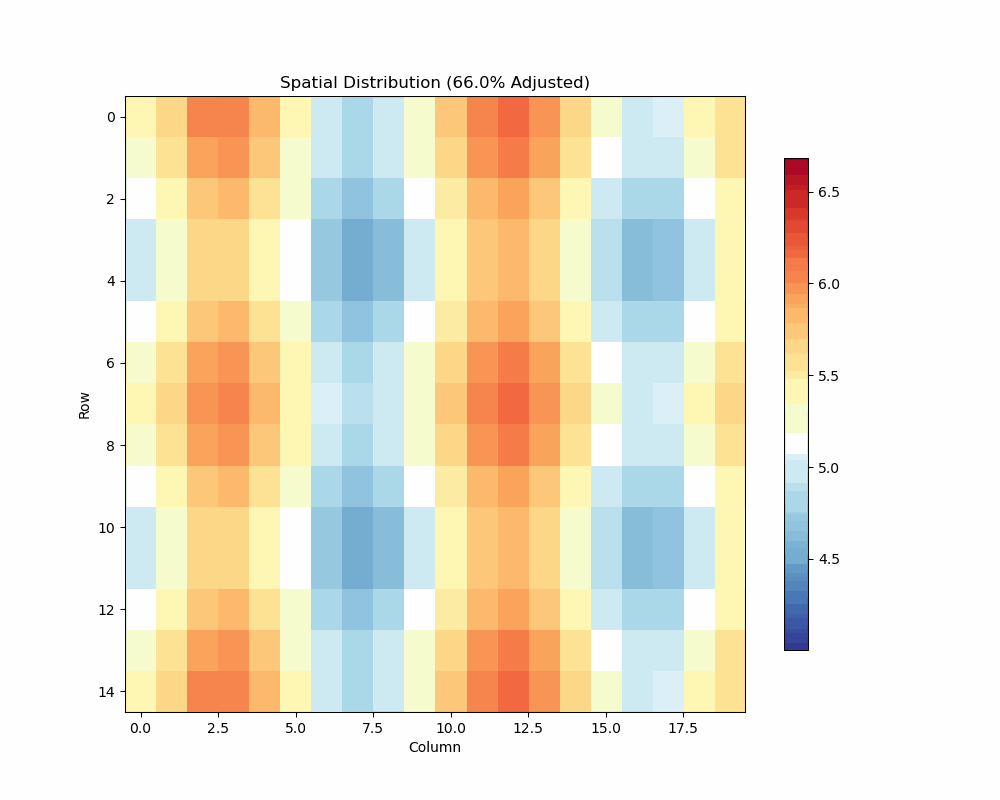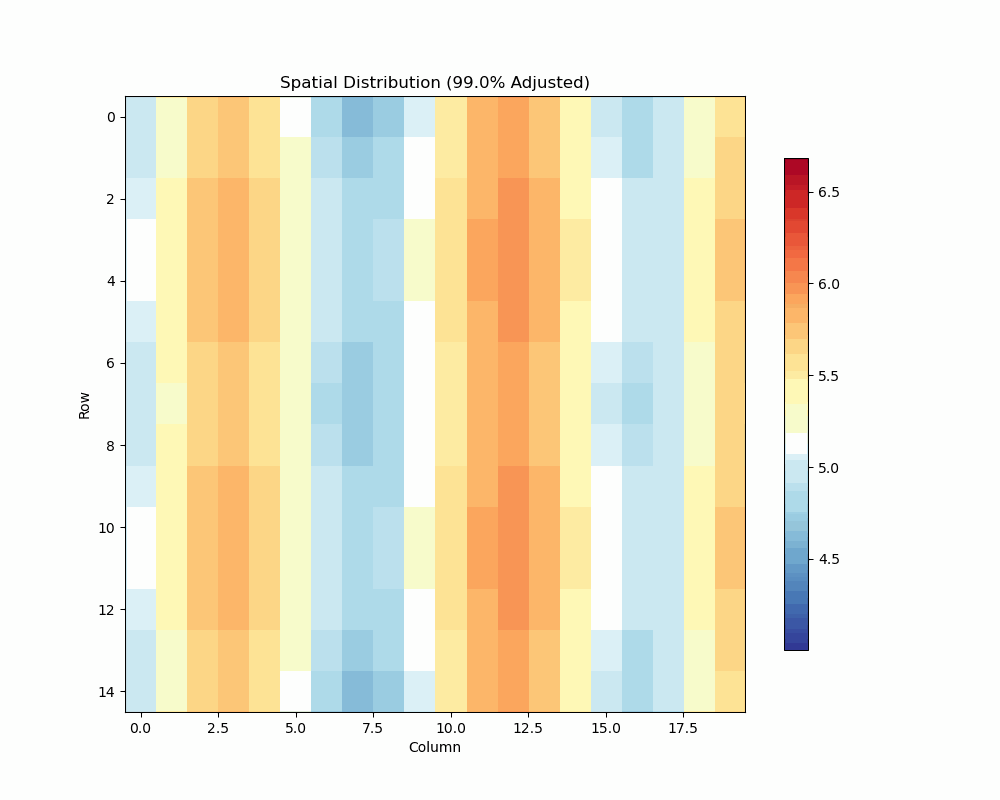
- Visualize spatial patterns and adjustments
- Learn experimental design concepts through interactive simulations
Dependencies
dgNova is built using robust scientific Python libraries:
- NumPy: For efficient numerical computations and array operations
- Pandas: For data manipulation and analysis
- Matplotlib: For creating static visualizations and plots
- Seaborn: For enhanced statistical visualizations
- Pillow: For image processing and animation support
These dependencies are automatically installed when you install dgNova using pip.
Current Status
The library is in active development. Currently implemented:
- Moving Grid Design (fully functional)
- Unreplicated Trial Analysis (UNREP class)
Future implementations will include:
- RCBD (Randomized Complete Block Design)
- Alpha-Lattice Design
- Augmented Design
- Split-Plot Design
- Mating Design
- Fieldbook Generation
- Spatial Analysis
Installation
pip install dgNova
Quick Start: Moving Grid Analysis
Example 1: Using Real Data
from dgNova import UNREP
# Load and analyze real field data
unrep = UNREP(
data='test_data/UNREP_Triticale_Unreplicated_Trial_Moving_Grids.csv',
response='Yield', # yield measurements
row='Row', # row positions
column='Column', # column positions
genotype='Genotype', # genotype labels
plot='Plot', # plot identifiers
design='moving_grid' # analysis method
)
# Run analysis
results = unrep.analyze()
Example 2: Simulation Study
# Simulate a field experiment
unrep_sim = UNREP(
row=15, # field dimensions
column=20,
heterogeneity=0.8, # spatial trend intensity (0-1)
mean=5.3, # base yield level
sd=0, # random variation
ne=1, # neighbor effects (0-1)
design='moving_grid'
)
# Analyze simulated data
unrep_sim.analyze()
Key Features of UNREP Class
Initialization Parameters
data: CSV file, DataFrame, or None (for simulation)response: Name of response variable columnrow,column: Field dimensions or column namesgenotype: Genotype identifier columnplot: Plot identifier columndesign: Analysis method (currently 'moving_grid')
Simulation Parameters
heterogeneity: Spatial trend intensity (0-1)mean: Base response levelsd: Random variationne: Neighbor effects strength (0-1)
Analysis Methods
-
analyze(): Performs moving grid analysis- Adjusts for spatial trends
- Calculates efficiency metrics
- Returns adjusted values and statistics
-
Visualization Methods:
# Plot spatial distribution unrep_sim.plot_spatial_analysis() # Animate adjustment process unrep_sim.animate(frames=100, interval=100) # Show detailed regions unrep_sim.plot_zoomed_regions()
Output Statistics
- Mean response
- Standard deviation
- Regression coefficient
- CV% (Adjusted)
- Relative efficiency
- Error variance
- LSD (5%)
Example Output
Using the Triticale trial data:
unrep = UNREP(data='UNREP_Triticale_Unreplicated_Trial_Moving_Grids.csv')
results = unrep.analyze()
# Sample output:
# Mean: 3.89
# Std: 1.14
# CV% (Adjusted): 29.37
# Relative Efficiency: 1.07
# Error Variance: 1.3040
# LSD (5%): 2.49
Understanding Moving Grid Design
The moving grid method adjusts plot values based on local spatial patterns by:
- Analyzing neighboring plots
- Detecting systematic field trends
- Adjusting values to account for spatial variation
- Providing more accurate genotype estimates
Visualization Tools
-
Spatial Analysis Plot:
unrep.plot_spatial_analysis(use_adjusted=True)
- Shows raw vs adjusted values
- Highlights spatial patterns
- Displays adjustment effects
-
Animation of Adjustment Process:
unrep.animate(frames=100, interval=100)
- Visualizes gradual transformation
- Helps understand adjustment mechanism
Contributing
dgNova is in active development. Contributions are welcome! Areas of focus:
- Additional design implementations
- Enhanced visualization options
- Documentation improvements
- Test cases and examples
License
MIT License
Contact
- Author: Nadim Khan
- Email: nfornadim@gmail.com
- Website: https://nadimkhan.org
- GitHub: https://github.com/nfornadimkhan/dgNova
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dgnova-1.2.12.tar.gz.
File metadata
- Download URL: dgnova-1.2.12.tar.gz
- Upload date:
- Size: 47.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.9.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7e2c705c6e5fe2c669d9a52564348dcce9b5404c82e2128f6c72622732a7805e
|
|
| MD5 |
a397eb1227f4bc834f5584112ec8bf47
|
|
| BLAKE2b-256 |
f6ad7e07e179b2d6f6268cf27a7d6f35a83e7e320a5727b3570b4c446dfd17fe
|
File details
Details for the file dgNova-1.2.12-py3-none-any.whl.
File metadata
- Download URL: dgNova-1.2.12-py3-none-any.whl
- Upload date:
- Size: 55.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.9.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
800bec8847dd7f8f7cd0e8fd565150278ceb3cfcc69b6bb41de0651f148b0ce7
|
|
| MD5 |
d54678ce8d13b903f1cf2d215b06805c
|
|
| BLAKE2b-256 |
e39ee7711eb01b6df88e4592e411df2b81a6d0438d0c7c2e598e51d6fb9768ea
|