An adapter for transfer DigitalTWIN Clinic Description to FHIR
Project description
Digitaltwins on FHIR
Usage
- Setup and connect to FHIR server
from digitaltwins_on_fhir.core import Adapter
adapter = Adapter("http://localhost:8080/fhir/")
Load data to FHIR server
Primary measurements
- Load FHIR bundle
await adapter.loader().load_fhir_bundle('./dataset/dataset-fhir-bundles')
- Load DigitalTWIN Clinical Description (primary measurements)
measurements = adapter.loader().load_sparc_dataset_primary_measurements()
with open('./dataset/measurements.json', 'r') as file:
data = json.load(file)
await measurements.add_measurements_description(data).generate_resources()
- Add Practitioner (researcher) to FHIR server
from digitaltwins_on_fhir.core.resource import Identifier, Code, HumanName, Practitioner
await measurements.add_practitioner(researcher=Practitioner(
active=True,
identifier=[
Identifier(use=Code("official"), system="sparc.org",
value='sparc-d557ac68-f365-0718-c945-8722ec')],
name=[HumanName(use="usual", text="Xiaoming Li", family="Li", given=["Xiaoming"])],
gender="male"
))
Workflow
Search
References in Task (workflow tool process) resource
- owner:
Patientreference - for:
ResearchStudy(Assay) reference - focus:
ActivityDefinition(workflow tool) reference - basedOn:
ResearchSubject(patient research subject) reference - requester (Optional):
Practitioner(researcher) reference - references in input
- ImagingStudy
- Observation
- DocumentReference
- references in output
- ImagingStudy
- Observation
- DocumentReference
Example
- Find a specific workflow process
- If known: patient, assay, and workflow tool uuids
client = adapter.async_client
# Step 1: find the patient
patient = await client.resources("Patient").search(
identifier="patient-xxxx").first()
# Step 2: find the assay
assay = await client.resources("ResearchStudy").search(
identifier="dataset-xxxx").first()
# Step 3: find the workflow tool
workflow_tool = await client.resources("ActivityDefinition").search(
identifier="workflow-tool-xxxx").first()
# Step 4: find the research subject (cohort in assay)
research_subject = await client.resources("ResearchSubject").search(
patient=patient.to_reference().reference,
study=assay.to_reference().reference).first()
workflow_tool_process = await client.resources("Task").search(
subject=assay.to_reference(),
focus=workflow_tool.to_reference(),
based_on=research_subject.to_reference(),
owner=patient.to_reference()).first()
- Find all input resources of the workflow tool process
inputs = workflow_tool_process.get("input")
for i in inputs:
input_reference = i.get("valueReference")
input_resource = await input_reference.to_resource()
- Find the input data comes from with dataset
- Assume we don't know the dataset and patient uuids at this stage
composition = await client.resources("Composition").search(
title="primary measurements",
entry=input_reference).first()
dataset_uuid = composition.get_by_path([
'identifier',
{'system':'https://www.auckland.ac.nz/en/abi.html'},
'value'
], '')
dataset = await client.resources("Composition").search(identifier=dataset_uuid).fetch_all()
- Find all output resources of the workflow tool process
outputs = workflow_tool_process.get("output")
for output in outputs:
output_reference = output.get("valueReference")
output_resource = await output_reference.to_resource()
References in PlanDefinition (workflow) resource
- action
- definition_canonical: ActivityDefinition (workflow tool) reference
Example
- If known workflow uuid
- Find all related workflow tools
workflow = await client.resources("PlanDefinition").search( identifier="sparc-workflow-uuid-001").first() actions = workflow.get("action") for a in actions: if a.get("definitionCanonical") is None: continue resource_type, _id = a.get("definitionCanonical").split("/") workflow_tool = await client.reference(resource_type, _id).to_resource()
- Find all related workflow processes
assay = await client.resources("ResearchStudy").search( identifier="dataset-xxxx").first() workflow_tool_processes = await client.resources("Task").search( subject=assay.to_reference()).fetch_all()
- Find all related workflow tools
Search in DigitalTWINS on FHIR methods
search = adapter.search()
- Finding all primary measurements for a patient
measurements = await self.search.get_patient_measurements("xxx-xxxx")
- Find which workflow, tool, and primary data was used to generate a specific derived measurement observation
res = await self.search.get_workflow_details_by_derived_data("Observation", "xxxx-xxxx")
- Find all inputs and their dataset uuid for generating the Observation
res = await self.search.get_all_inputs_by_derived_data("Observation","xxx-xxxx")
- Find all tools and models used by a workflow and their workflow tool processes
res = await self.search.get_all_workflow_tools_by_workflow(name="Automated torso model generation - script")
- Find inputs and outputs of a given tool in a workflow
res = await self.search.get_all_inputs_outputs_of_workflow_tool(name="Tumour Position Correction (Manual) Tool")
Reference in resource
ResearchStudy- Study- principalInvestigator: Practitioner reference
ResearchStudy- Assay- protocol: [ PlanDefinition(Workflow) reference ]
- partOf: [ ResearchStudy(Study) reference ]
ResearchSubject- Assay cohort- individual(patient): Patient reference
- study: ResearchStudy(Assay) reference
- consent: Consent reference
ResearchSubject- dataset cohort- individual(patient): Patient reference
- consent: Consent reference
Composition- primary measurements- author: [ Patient reference, Practitioner reference ]
- subject: ResearchSubject reference
- entry: [ Observation reference, ImagingStudy reference, DocumentReference reference ]
ImagingStudy- subject: Patient reference
- endpoint: [ Endpoint Reference ]
- referrer: Practitioner reference
Observation- primary measurements- subject: Patient reference
DocumentRefernce- subject: Patient reference
PlanDefinition:- action.definitionCanonical: ActivityDefinition reference string
ActivityDefinition:- participant: [ software uuid, model uuid ]
Task:- owner: patient reference
- for(subject): ResearchSubject(Assay) reference
- focus: ActivityDefinition(workflow) tool reference
- basedOn: research subject reference
- requester (Optional): practitioner reference
- input: [ Observation reference, ImagingStudy reference ]
- output: [ Observation reference, ImagingStudy reference ]
Work steps
- Upload measurements dataset (primary measurements)
- Upload workflow / workflow tools
- Create Assay (get practitioner, study, and workflow process information)
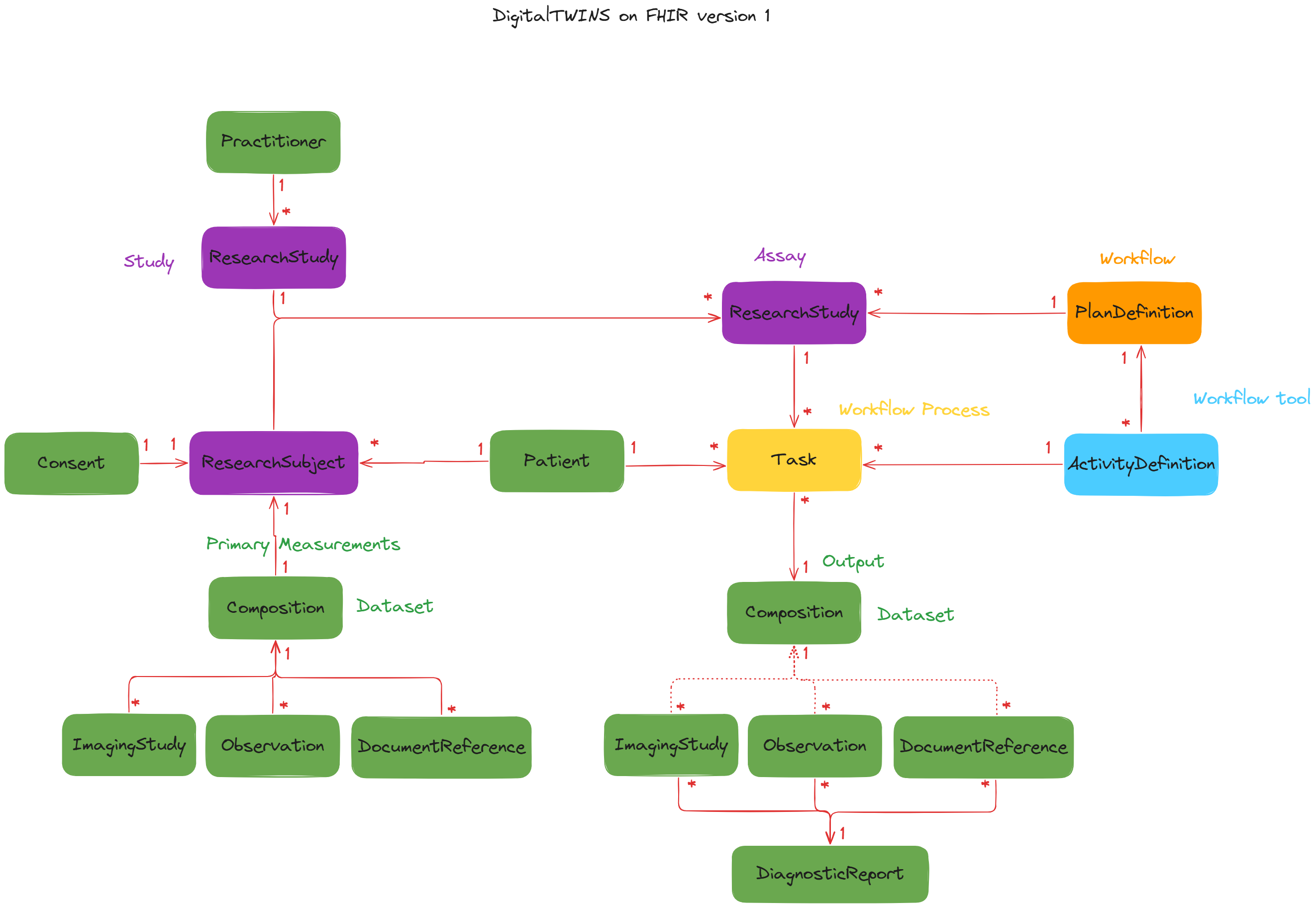
DigitalTWIN on FHIR Diagram
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file digitaltwins_on_fhir-1.3.2.tar.gz.
File metadata
- Download URL: digitaltwins_on_fhir-1.3.2.tar.gz
- Upload date:
- Size: 71.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1422c30d04b282ec1c6fa4d268e38a22f6850fc5c2334ef73cbce68f49d020b7
|
|
| MD5 |
34525421729cdf6bb96b373fde281ea8
|
|
| BLAKE2b-256 |
38838862306f75d4bae48ae046951c21161f5499201644896611f77f001868fd
|
File details
Details for the file digitaltwins_on_fhir-1.3.2-py3-none-any.whl.
File metadata
- Download URL: digitaltwins_on_fhir-1.3.2-py3-none-any.whl
- Upload date:
- Size: 90.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
26f39c80a0f6fb9668c201e2151a6cd11314e083483ff48804e4b0259cfaab1e
|
|
| MD5 |
2866d7b0aa6876a64b08683816a03ec6
|
|
| BLAKE2b-256 |
ce1b48da809d5e25f8106a29fcb851088e7ef16af0cb48f2c4ba60585b939e2b
|