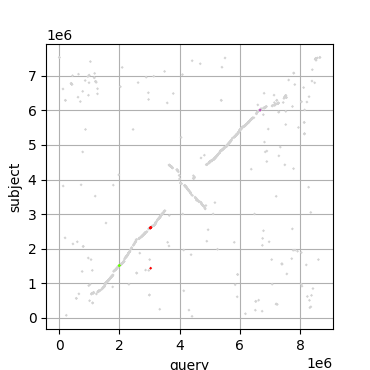
A python for plotting dot plots from blast results.
Project description
dotplotter
build dotplots from blastn results
Description
dotplotter can take blast results, either as standard input or from a .tsv file and plots a standard dot plot. It can also highlight regions based on provided arguments (single region) or from a file (single or multiple regions) see below. Dot plots are a classic way to visualise DNA similarity and can be used for whole chromosomes or small regions.
Installation
The easiest way to install dotplotter is though the python package index.
pip install dotplotter
This will fetch and install the latest version from: LINK
You can also install by cloning this repository.
dotplotter only requires matplotlib and this should be installed automatically.
Usage
Basic Usage
You can use dotplotter in two ways: using stdin, or reading a .tsv file.
Example data can be found in this repository ./example_data.
Important: Regardless of method, make sure your results are in blast outfmt 6. This is specified in the search with -outfmt 6
stdin method
You can pipe your blastn results straight into dotplotter:
blastn -query streptomyces_coelicolor.fna -subject streptomyces_albus.fna -outfmt 6 | dotplotter
.tsv method
You can also read your results in from a previously generated .tsv file (e.g. blastn -query streptomyces_coelicolor.fna -subject streptomyces_albus.fna -outfmt 6 > blastn.tsv)
dotplotter -i blastn.tsv
blastn tips
Setting the parameters of you blast search can be very important. For larger sequences the default values should be fine. However, if you are looking for smaller repeats, you may need to adjust the word size of you blastn search -word_size.
Highlighting
You can also highlight regions of interest.
Important: Highlighted regions are based on the query sequence ONLY.
Single Region Highlighting
The easiest way to highlight a single region is to use the in-built parameters: -hs/highlight-start and -he/--highlight-end. For example:
dotplotter -i blastn.tsv -hs 3024902 -he 3054689
Multiple Region Highlighting
To highlight multiple regions, you can provide a .csv file containing the required information. Each line should contain the start position, end position and the hex value (or colour name).
See .example_data/highlight.csv for an example. You can specify this file with -hf/--highligh-file.
dotplotter -i blastn.tsv -hf highlight.csv
Further Usage
For more usage information, use the help command:
dotplotter -h
Example Output
Comparison of Streptomyces coelicolor and Streptomyces albus with gene clusters for ectoine (green), desferrioxamine (red) and spore pigment (purple) highlighted.
Citation
Mohite, O.S., Jørgensen, T.S., Booth, T.J. et al. Pangenome mining of the Streptomyces genus redefines species’ biosynthetic potential. Genome Biol 26, 9 (2025)
Patch Notes
Version 1
- 1.0.0
- initial release
- 1.0.1
- removed unnecessary print statements
- fixed README.md
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dotplotter-1.0.1.tar.gz.
File metadata
- Download URL: dotplotter-1.0.1.tar.gz
- Upload date:
- Size: 18.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.8.0 colorama/0.4.4 importlib-metadata/4.6.4 keyring/23.5.0 pkginfo/1.8.2 readme-renderer/34.0 requests-toolbelt/0.9.1 requests/2.31.0 rfc3986/1.5.0 tqdm/4.57.0 urllib3/1.26.5 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9a5d3443cbabe0693e34d83158dd3461a90776a2eb71fad64a2d8db195cbd4e5
|
|
| MD5 |
89d00ce86511458b976d4a67786afbbe
|
|
| BLAKE2b-256 |
231754ebb4d578602340306125848f4ee9bb1e1a469fc02565de4fc6ba5a67f5
|
File details
Details for the file dotplotter-1.0.1-py3-none-any.whl.
File metadata
- Download URL: dotplotter-1.0.1-py3-none-any.whl
- Upload date:
- Size: 19.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.8.0 colorama/0.4.4 importlib-metadata/4.6.4 keyring/23.5.0 pkginfo/1.8.2 readme-renderer/34.0 requests-toolbelt/0.9.1 requests/2.31.0 rfc3986/1.5.0 tqdm/4.57.0 urllib3/1.26.5 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d6549df88feb991ebb93b2c202175dda16c43c808ddbc0aee7468ba27dfbe0dd
|
|
| MD5 |
1dd22090cd027e5cb9cc9b0b94d1fc55
|
|
| BLAKE2b-256 |
d5d1f92d663ced0805f08a2e25af11083a658c45374297afa793602e970429cc
|