A Python tool for standardizing drug names using the latest FDA's UNII Names list.
Project description
Drugname Standardizer
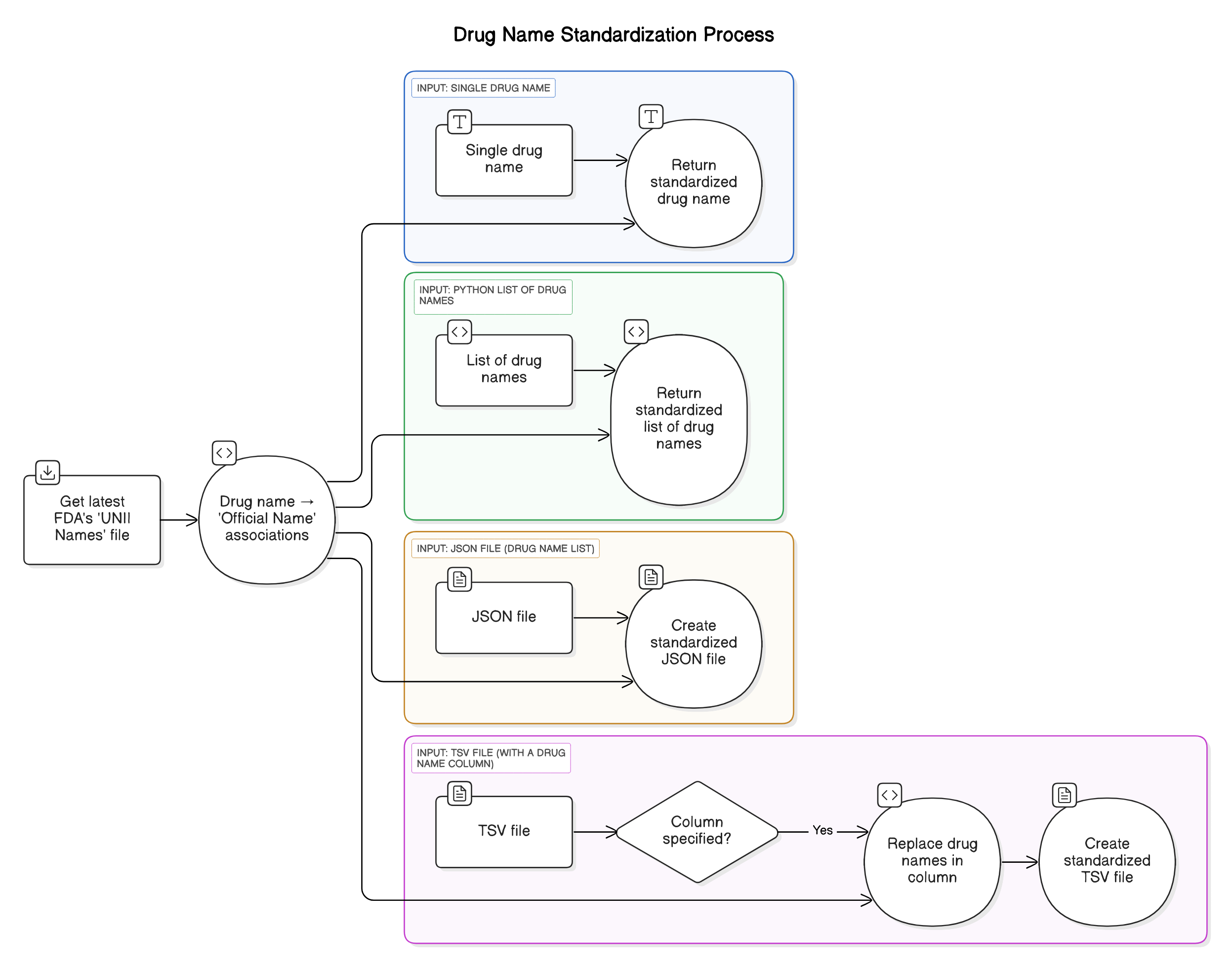
The Drugname Standardizer is a Python tool for standardizing drug names using the official FDA's UNII Names List archive. It notably supports both JSON and TSV/CSV input formats, making it easy to ensure consistent drug naming in datasets.
Features
-
A trusted source for drug synonyms : the package automatically downloads the latest version of the UNII Names file from the official FDA repository (montly check of an update). The
UNII_Names.txtis saved to the package'sdata/folder for use without redownload before next update. The user can also choose to indicate another local UNII Names file if a particular version is preferred. -
Parsing of the FDA's UNII Names List to map drug names (code / official / systematic / common / brand names) to a single preferred name (i.e. the Display Name of the UNII Names file).
-
Input versatility:
- a single drug name,
- a list of drug names,
- a JSON input file (a list of drugs to standardize)
- a TSV input file (a dataframe containing a column of drugs to standardize)
-
Provides both a Python package interface for scripting and a command-line interface (CLI) for direct use.
-
Resolves naming ambiguities of the FDA's UNII Names file by selecting the shortest Display Names. Rare but exists: 55 / 986397 associations in
UNII_Names_20Dec2024.txt. For example, forPRN1008the ambiguity is solved by keepingRILZABRUTINIBwhereas 2 associations exist:PRN1008... ...RILZABRUTINIB, (.ALPHA.E,3S)-PRN1008... ...RILZABRUTINIB
Warning:
There are code / official / systematic / common / brand names for drugs. Some are linked to different level of details about the compound.
The standardization proposed here gathers information at the "upper" level (i.e. the less detailled one). I relied on the "Preferred Substance Name" (= the Display name field) indicated in the correspondence table provided by the FDA.
For instance : both 3'-((1R)-1-((6R)-5,6-DIHYDRO-4-HYDROXY-2-OXO-6-PHENETHYL-6-PROPYL-2H-PYRAN-3-YL)PROPYL)-5-(TRIFLUOROMETHYL)-2-PYRIDINESULFONANILIDE (systematic name) and Aptivus (brand name) become TIPRANAVIR.
Usage
Python API
You can use the package programmatically in your Python scripts:
from drugname_standardizer import standardize
Examples:
- Get the preferred name for a specific drug:
drug_name = "GDC-0199"
preferred_name = standardize(drug_name)
print(preferred_name) # Outputs: VENETOCLAX
- Standardize a list of drugs:
drug_names = ["GDC-0199", "Aptivus", "diodrast"]
preferred_names = standardize(drug_names)
print(preferred_names) # Outputs: ["VENETOCLAX", "TIPRANAVIR", "IODOPYRACET"]
- Standardize a JSON file:
standardize(
input_file = "drugs.json",
file_type = "json",
output_file = "standardized_drugs.json"
)
# Outputs: Standardized JSON file saved as standardized_drugs.json
- Standardize a TSV/CSV file:
standardize(
input_file = "dataset.csv",
file_type = "tsv",
column_drug = 0,
separator = ","
)
# Outputs: Standardized TSV file saved as dataset_drug_standardized.csv
Command-Line Interface
You can also use a CLI for standardizing a drug name, a JSON file or a TSV/CSV file.
- Required arguments:
--input,-i: A drug name or the path to a JSON/TSV file
- Optional arguments:
--file_type,-f: Type of the input file (jsonortsv)--output,-o: The output file name (relative path can be given). Defaults: the input file name with_drug_standardizedadded before the extension.--column_drug,-c: Index of the column containing the drug names to standardize (required for TSV files). Starts at 0: 1st column = column 0.--separator,-s: Field separator for TSV files. Defaults:\t.--unii_file,-u: Path to a UNII Names List file. Defaults: automatic download of the latest version.
Examples:
- Get the preferred name for a specific drug:
drugname_standardizer -i DynaCirc
- Standardize a JSON file:
drugname_standardizer -i drugs.json -f json
- Standardize a TSV file: e.g., using a pipe as separator and a custom file name for the output:
drugname_standardizer -i dataset.tsv -f tsv -c 2 -s "|" -o standardized_dataset.tsv
Installation
Using pip
python3 -m pip install drugname_standardizer
GitHub repository
git clone https://github.com/StephanieChevalier/drugname_standardizer.git
cd drugname_standardizer
pip install -r requirements.txt
Requirements:
- Python 3.7+
- Dependencies:
requests >= 2tqdm >= 4
How it works
-
Parse UNII File:
- Reads the UNII Names List to create a mapping of drug names to the Display Name (i.e. the preferred name).
- Resolves potential naming conflicts by selecting the shortest Display Name (55 / 986397 associations).
-
Standardize Names:
- For a single drug name: return the preferred name.
- For a list of drug names: maps drug names to their preferred names and return the updated list.
- For JSON input: Maps drug names to their preferred names and saves the results to a JSON file.
- For TSV input: Updates the specified column with standardized drug names and saves the modified DataFrame to a TSV file.
Package structure
drugname_standardizer/
├── drugname_standardizer/
│ ├── __init__.py # Package initialization
│ ├── standardizer.py # Core logic for name standardization
│ └── data/
│ ├── UNII_Names.txt # UNII Names List file (ensured to be no older than 1 month when available)
│ └── UNII_dict.pkl # parsed UNII Names List
├── tests/
│ ├── __init__.py
│ └── test_standardizer.py # Unit tests for the package
├── LICENSE # MIT License
├── pyproject.toml # Package configuration
├── README.md # Project documentation
└── requirements.txt # Development dependencies
License
This project is licensed under the MIT License - see the LICENSE file for details.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file drugname_standardizer-1.2.5.tar.gz.
File metadata
- Download URL: drugname_standardizer-1.2.5.tar.gz
- Upload date:
- Size: 13.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.12.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
85a5b146e198ed601e96619d191febdf6ee535a3ab1ebe1655cde94ecef04d42
|
|
| MD5 |
c77a7261a3476fcf6652c78fc71966f5
|
|
| BLAKE2b-256 |
018c7db9c4bd212d1da52158a5b7dfa24f330cf36598bdcb66192ad0f310af20
|
File details
Details for the file drugname_standardizer-1.2.5-py3-none-any.whl.
File metadata
- Download URL: drugname_standardizer-1.2.5-py3-none-any.whl
- Upload date:
- Size: 10.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.12.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4856b302621eadfa856f47eb829f07ec4a8608fd3ad56e92188286b9d2270540
|
|
| MD5 |
c3bf0ff7416189ebadba051a1dd9be1d
|
|
| BLAKE2b-256 |
95c37a23c8c44ba9685a2c8b69af42ba40bbef6b2863c150a67ff9d6973283ce
|