Mapping Vector Field of Single Cells
Project description
Dynamo: Mapping Vector Field of Single Cells
Inclusive model of expression dynamics with metabolic labeling based scRNA-seq / multiomics, vector field reconstruction, potential landscape mapping and differential geometry analyses.
Installation - Ten minutes to dynamo - Tutorials - API - Citation - Theory
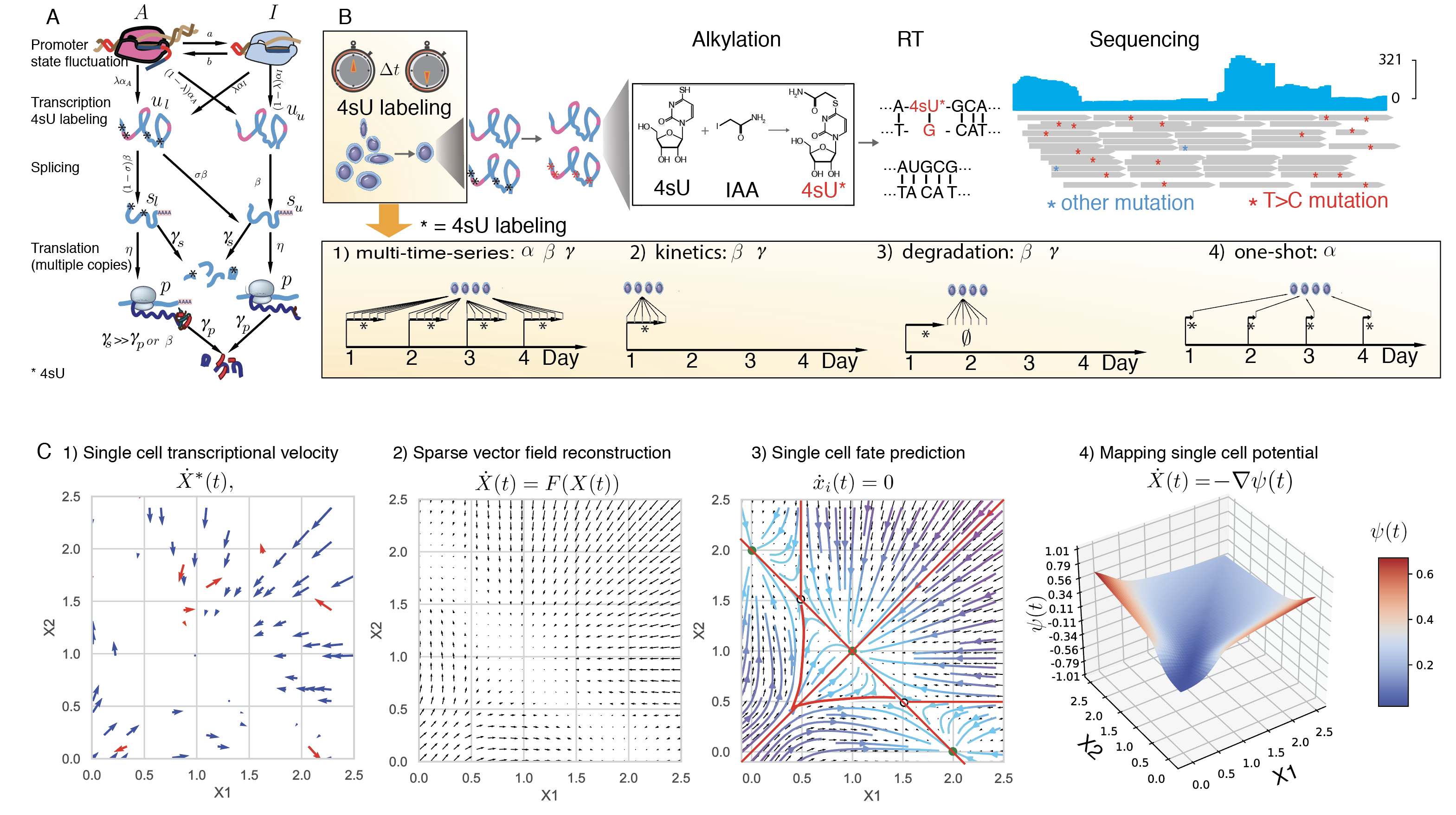
Single-cell RNA-seq, together with RNA velocity and metabolic labeling, reveals cellular states and transitions at unprecedented resolution. Fully exploiting these data, however, requires dynamical models capable of predicting cell fate and unveiling the governing regulatory mechanisms. Here, we introduce dynamo, an analytical framework that reconciles intrinsic splicing and labeling kinetics to estimate absolute RNA velocities, reconstructs velocity vector fields that predict future cell fates, and finally employs differential geometry analyses to elucidate the underlying regulatory networks. We applied dynamo to a wide range of disparate biological processes including prediction of future states of differentiating hematopoietic stem cell lineages, deconvolution of glucocorticoid responses from orthogonal cell-cycle progression, characterization of regulatory networks driving zebrafish pigmentation, and identification of possible routes of resistance to SARS-CoV-2 infection. Our work thus represents an important step in going from qualitative, metaphorical conceptualizations of differentiation, as exemplified by Waddington’s epigenetic landscape, to quantitative and predictive theories.
Highlights of dynamo
- Robust and accurate estimation of RNA velocities for regular scRNA-seq datasets:
- Three methods for the velocity estimations (including the new negative binomial distribution based approach)
- Improved kernels for transition matrix calculation and velocity projection
- Strategies to correct RNA velocity vectors (when your RNA velocity direction is problematic)
- Inclusive modeling of time-resolved metabolic labeling based scRNA-seq:
- Explicitly model RNA metabolic labeling, in conjunction with RNA bursting, transcription, splicing and degradation
- Comprehensive RNA kinetic rate estimation for one-shot, pulse, chase and mixture metabolic labeling experiments
- Move beyond RNA velocity to continuous vector field function for functional and predictive analyses of cell fate transitions:
- Dynamical systems approaches to identify stable cell types (fixed points), boundaries of cell states (separatrices), etc
- Calculate RNA acceleration (reveals early drivers), curvature (reveals master regulators of fate decision points), divergence (stability of cell states) and RNA Jacobian (cell-state dependent regulatory networks)
- Various downstream differential geometry analyses to rank critical regulators/effectors, and visualize regulatory networks at key fate decision points
Discussion
Please use github issue tracker to report coding related issues of dynamo. For community discussion of novel usage cases, analysis tips and biological interpretations of dynamo, please join our public slack workspace: dynamo-discussion (Only a working email address is required from the slack side).
Contribution
If you want to contribute to the development of dynamo, please check out CONTRIBUTION instruction: Contribution
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dynamo-release-1.0.0.tar.gz.
File metadata
- Download URL: dynamo-release-1.0.0.tar.gz
- Upload date:
- Size: 427.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.5.0 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.61.0 CPython/3.6.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7ca57db299ba009f66fccaeb73f7aeb3a0204e676b752e6ccd0550a96f57cdb3
|
|
| MD5 |
d9f24e84622ad2d0156ca39a8deb34ae
|
|
| BLAKE2b-256 |
6105ac84f9076ca2cc06ecebd26347bb2917a41404a08c7e97f4b13b928af26c
|
File details
Details for the file dynamo_release-1.0.0-py3-none-any.whl.
File metadata
- Download URL: dynamo_release-1.0.0-py3-none-any.whl
- Upload date:
- Size: 492.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.5.0 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.61.0 CPython/3.6.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4b18c990b8144b2d4e30d11a6d533440d485176d0366f54a766a00d5cc532fb0
|
|
| MD5 |
1077cac10d530f0612b7748aa71c1224
|
|
| BLAKE2b-256 |
8cbb4d12c7f9c91a7ccc7a59d9fd10cff04fca9f4e47c8500f5b1e5978d94c4f
|