An Evolutionary-scale Model (ESM) for protein function prediction from amino acid sequences using the Gene Ontology (GO).
Project description
ESMC Protein Function Predictor
An Evolutionary-scale Model (ESM) for protein function prediction from amino acid sequences using the Gene Ontology (GO). Based on the ESM Cambrian Transformer architecture, pre-trained on UniRef, MGnify, and the Joint Genome Institute's database and fine-tuned on the AmiGO Boost protein function dataset, this model predicts the GO subgraph for a particular protein sequence - giving you insight into the molecular function, biological process, and location of the activity inside the cell.
What are GO terms?
"The Gene Ontology (GO) is a concept hierarchy that describes the biological function of genes and gene products at different levels of abstraction (Ashburner et al., 2000). It is a good model to describe the multi-faceted nature of protein function."
"GO is a directed acyclic graph. The nodes in this graph are functional descriptors (terms or classes) connected by relational ties between them (is_a, part_of, etc.). For example, terms 'protein binding activity' and 'binding activity' are related by an is_a relationship; however, the edge in the graph is often reversed to point from binding towards protein binding. This graph contains three subgraphs (subontologies): Molecular Function (MF), Biological Process (BP), and Cellular Component (CC), defined by their root nodes. Biologically, each subgraph represent a different aspect of the protein's function: what it does on a molecular level (MF), which biological processes it participates in (BP) and where in the cell it is located (CC)."
From CAFA 5 Protein Function Prediction
Pretrained Models
The following pretrained models are available on HuggingFace Hub.
| Name | Embedding Dim. | Attn. Heads | Encoder Layers | Context Length | Total Parameters |
|---|---|---|---|---|---|
| andrewdalpino/ESMC-300M-Protein-Function | 960 | 15 | 30 | 2048 | 361M |
| andrewdalpino/ESMC-600M-Protein-Function | 1152 | 18 | 36 | 2048 | 644M |
Cloning the Repo
You'll need the code in the repository to run the model. To clone the repo onto your local machine enter the command like in the example below.
git clone https://github.com/andrewdalpino/ESMC-Function-Classifier
Install Project Dependencies
Project dependencies are specified in the requirements.txt file. You can install them with pip using the following command from the project root. We recommend using a virtual environment such as venv to keep package dependencies on your system tidy.
python -m venv ./.venv
source ./.venv/bin/activate
pip install -r requirements.txt
Using a Pretrained Model
You'll need the code in this repository to load the pretrained model weights. Import the EsmGoTermClassifier class and call from_pretrained() with the name of the model you want to load from the HuggingFace Hub.
from esm.tokenization import EsmSequenceTokenizer
from model import EsmcGoTermClassifier
model_name = "andrewdalpino/ESMC-300M-Protein-Function"
tokenizer = EsmSequenceTokenizer()
model = EsmcGoTermClassifier.from_pretrained(model_name)
Fine-tuning
We'll be fine-tuning the pre-trained ESMC model with a multi-label binary classification head on the AmiGO Boost dataset of GO term-annotated protein sequences. To begin training with the default arguments, you can enter the command below.
python fine-tune.py
You can change the base model and dataset subset like in the example below.
python fine-tune.py --base_model="esmc_600m" --dataset_subset="biological_process"
You can also adjust the batch_size, gradient_accumulation_steps, and learning_rate like in the example below.
python fine-tune.py --batch_size=16 --gradient_accumulation_step=8 --learning_rate=5e-4
Training checkpoints will be saved at the checkpoint_path location. You can change the location and the checkpoint_interval like in the example below.
python fine-tune.py --checkpoint_path="./checkpoints/biological-process-large.pt" --checkpoint_interval=3
If you would like to resume training from a previous checkpoint, make sure to add the resume argument. Note that if the checkpoint path already exists, the file will be overwritten.
python fine-tune.py --checkpoint_path="./checkpoints/checkpoint.pt" --resume
Training Arguments
| Argument | Default | Type | Description |
|---|---|---|---|
| --base_model | "esmc_300m" | str | The base model name, choose from esmc_300m, esmc_600m. |
| --dataset_subset | "all" | str | The subset of the dataset to train on, choose from all, mf for molecular function, cc for cellular component, or bp for biological process. |
| --num_dataset_processes | 1 | int | The number of CPU processes to use to process and load samples. |
| --min_sequence_length | 1 | int | The minimum length of the input sequences. |
| --max_sequence_length | 2048 | int | The maximum length of the input sequences. |
| --unfreeze_last_k_layers | 7 | int | Fine-tune the last k layers of the pre-trained encoder network. |
| --add_lora | False | bool | Should we add LoRA adapters to the frozen encoder layers? |
| --lora_rank | 8 | int | The rank of the matrices used to approximate the weights of frozen encoder layers during fine-tuning. |
| --batch_size | 8 | int | The number of samples to pass through the network at a time. |
| --gradient_accumulation_steps | 16 | int | The number of batches to pass through the network before updating the weights. |
| --max_gradient_norm | 1.0 | float | Clip gradients above this threshold norm before stepping. |
| --learning_rate | 5e-4 | float | The learning rate of the Adam optimizer. |
| --num_epochs | 40 | int | The number of epochs to train for. |
| --classifier_hidden_ratio | 1 | {1, 2, 4} | The ratio of hidden nodes to embedding dimensions in the classifier head. |
| --eval_interval | 2 | int | Evaluate the model after this many epochs on the testing set. |
| --checkpoint_interval | 2 | int | Save the model parameters to disk every this many epochs. |
| --checkpoint_path | "./checkpoints/checkpoint.pt" | string | The path to the training checkpoint. |
| --resume | False | bool | Should we resume training from the last checkpoint? |
| --run_dir_path | "./runs" | str | The path to the TensorBoard run directory for this training session. |
| --device | "cuda" | str | The device to run the computation on ("cuda", "cuda:1", "mps", "cpu", etc). |
| --seed | None | int | The seed for the random number generator. |
Training Dashboard
We use TensorBoard to capture and display training events such as loss and gradient norm updates. To launch the dashboard server run the following command from the terminal.
tensorboard --logdir=./runs
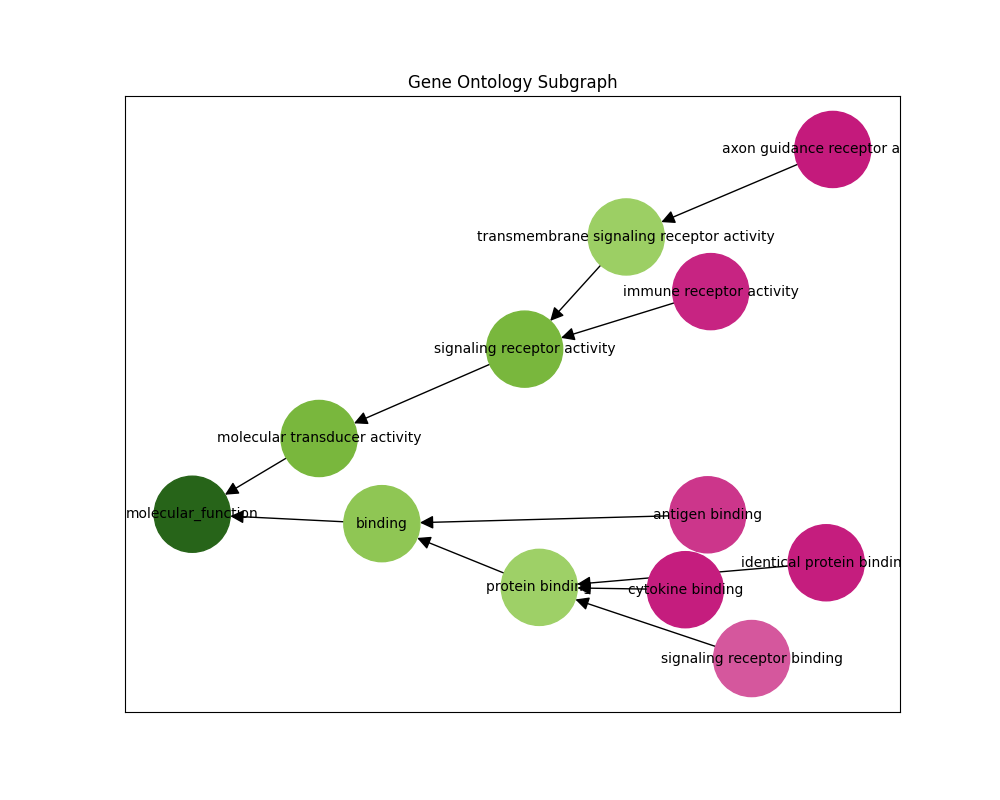
GO Subgraph Prediction
We can also infer the gene ontology subgraph of a particular sequence. The predict-subgraph.py script outputs a graphical representation of the predictions where green nodes have high probability and pink nodes have low probability.
python predict-subgraph.py --checkpoint_path="./checkpoints/checkpoint.pt" --top_p=0.1
Checkpoint loaded successfully
Enter a sequence: MPNERLKWLMLFAAVALIACGSQTLAANPPDADQKGPVFLKEPTNRIDFSNSTG
Prediction Arguments
| Argument | Default | Type | Description |
|---|---|---|---|
| --checkpoint_path | "./checkpoints/checkpoint.pt" | str | The path to the training checkpoint. |
| --go_db_path | "./dataset/go-basic.obo" | str | The path to the Gene Ontology basic obo file. |
| --context_length | 2048 | int | The maximum length of the input sequences. |
| --top_p | 0.5 | float | Only display nodes with the top p probability. |
| --device | "cuda" | str | The device to run the computation on ("cuda", "cuda:1", "mps", "cpu", etc). |
| --seed | None | int | The seed for the random number generator. |
References:
- T. Hayes, et al. Simulating 500 million years of evolution with a language model, 2024.
- M. Ashburner, et al. Gene Ontology: tool for the unification of biology, 2000.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file esmc_function_classifier-0.0.1.tar.gz.
File metadata
- Download URL: esmc_function_classifier-0.0.1.tar.gz
- Upload date:
- Size: 10.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
60806ca596461b64c794fa2c3dc9341c63386008c88500125b4b8cd074a4f75a
|
|
| MD5 |
c84c05bbf4835ccee469abdc256921d4
|
|
| BLAKE2b-256 |
43088ca660053119003aecab1b4e85b14e9abc7f21b0ccdc06415611b0bd399c
|
File details
Details for the file esmc_function_classifier-0.0.1-py3-none-any.whl.
File metadata
- Download URL: esmc_function_classifier-0.0.1-py3-none-any.whl
- Upload date:
- Size: 7.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d1448c2f2272e534251075590cbf2383266293f0798f12224c3179db3bc82266
|
|
| MD5 |
80b9c3262b00b6187b319ee588fdfef1
|
|
| BLAKE2b-256 |
ca56d832eb706723b1d3e7df3c123d66ea96dbe70c14035909a50b14c107c657
|