User Friendly Data-Driven Numerical Optimization and Exploration
Project description
evopt
User Friendly Black-Box Numerical Optimization, Exploration, and Equation Discovery
evopt is a package for efficient parameter optimization using the CMA-ES (Covariance Matrix Adaptation Evolution Strategy) algorithm, exploration using Sobol sequence sampling, and symbolic regression using PySR. It provides a user-friendly way to find the best set of parameters for a given problem, especially when the problem is complex, non-linear, and doesn't have easily calculable derivatives.
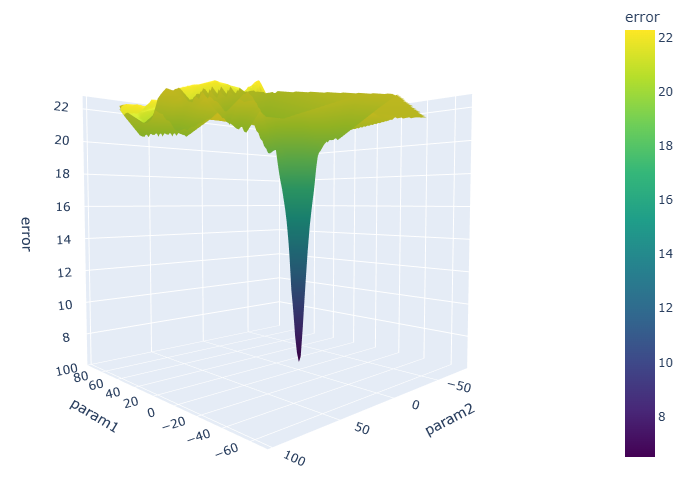
Optimization of the two parameter Ackley function.
Documentation
Complete documentation is available at evopt.readthedocs.io.
Scope
- Focus:
evoptprovides a CMA-ES-based optimization routine that is easy to set up and use. - Black-box Parameter Optimization: The package is designed for problems where you need to find the optimal values for a set of parameters, without being able to parameterise the evaluator function.
- Efficient exploration and equation discovery: You can efficiently sample the parameter space using Sobol sequences, and apply symbolic regression to the results to discover the underlying black-box equations.
- Directory Management: The package includes robust directory management to organise results, checkpoints, and logs.
- Logging: It provides logging capabilities to track the optimization process.
- Checkpointing: It supports saving and loading checkpoints to resume interrupted optimization runs.
- CSV Output: It writes results and epoch data to CSV files for easy analysis.
- Easy results plotting: Simple pain-free methods to plot the results.
- High Performance Computing: It can leverage HPC resources for increased performance.
Installation
You can install the package using pip:
pip install evopt
Usage
Here is an example of how to use the evopt package to optimize the Rosenbrock function:
import evopt
# Define your parameters, their bounds, and evaluator function
params = {
'param1': (-5, 5),
'param2': (-5, 5),
}
def evaluator(param_dict):
# Your evaluation logic here, in this case the Rosenbrock function
p1 = param_dict['param1']
p2 = param_dict['param2']
error = (1 - p1) ** 2 + 100*(p2 - p1 ** 2) ** 2
return error
# Run the optimization using .optimize method
results = evopt.optimize(params, evaluator)
Here is the corresponding output:
Starting new CMAES run in directory path\to\base\dir\evolve_0
Epoch 0 | (1/16) | Params: [1.477, -2.369] | Error: 2069.985
Epoch 0 | (2/16) | Params: [-2.644, -1.651] | Error: 7481.172
Epoch 0 | (3/16) | Params: [0.763, -4.475] | Error: 2557.411
Epoch 0 | (4/16) | Params: [4.269, -0.929] | Error: 36687.174
Epoch 0 | (5/16) | Params: [-1.879, -4.211] | Error: 5999.711
Epoch 0 | (6/16) | Params: [4.665, -2.186] | Error: 57374.982
Epoch 0 | (7/16) | Params: [-1.969, -2.326] | Error: 3856.201
Epoch 0 | (8/16) | Params: [-1.588, -3.167] | Error: 3244.840
Epoch 0 | (9/16) | Params: [-2.191, -2.107] | Error: 4780.562
Epoch 0 | (10/16) | Params: [2.632, -0.398] | Error: 5369.439
Epoch 0 | (11/16) | Params: [-2.525, -1.427] | Error: 6099.094
Epoch 0 | (12/16) | Params: [4.161, -2.418] | Error: 38955.920
Epoch 0 | (13/16) | Params: [-0.435, -1.422] | Error: 261.646
Epoch 0 | (14/16) | Params: [-0.008, -3.759] | Error: 1414.379
Epoch 0 | (15/16) | Params: [-4.243, -0.564] | Error: 34496.083
Epoch 0 | (16/16) | Params: [0.499, -3.170] | Error: 1169.217
Epoch 0 | Mean Error: 13238.614 | Sigma Error: 17251.295
Epoch 0 | Mean Parameters: [0.062, -2.286] | Sigma parameters: [2.663, 1.187]
Epoch 0 | Normalised Sigma parameters: [1.065, 0.475]
...
Epoch 21 | Mean Error: 2.315 | Sigma Error: 0.454
Epoch 21 | Mean Parameters: [-0.391, 0.192] | Sigma parameters: [0.140, 0.154]
Epoch 21 | Normalised Sigma parameters: [0.056, 0.062]
Terminating after meeting termination criteria at epoch 22.
print(results.best_parameters)
{param1: -0.391, param2: 0.192}
Multi-objective target optimization
Sometimes when using black-box functions like simulations, your result may be a specific variable such as mean pressure, temperature, or velocity. With evopt it is possible to specify a target value for the optimizer to reach, and in cases where targets are in conflict, you can specify hard or soft target preference such that the optimizer can weigh target priority.
For example:
import evopt
# example black-box function
def example_eval(param_dict):
x1 = param_dict['x1']
x2 = param_dict['x2']
target1 = (1 - 2 * (x1 - 3))
target2 = x1 ** 2 + 1 + x2
return {'target1': target1, 'target2': target2}
# define objectives
target_dict={
"target1": {"value": (2.8), "hard": True},
"target2": {"value": (2.9), "hard": False},
}
# define free parameters (evaluated by black-box function)
params = {
"x1": (-5, 5),
"x2": (-5, 5),
}
results = evopt.optimize(params, example_eval, target_dict=target_dict)
and corresponding output:
Starting new CMAES run in directory path\to\base\dir\evolve_0
target1: 100% of values outside [2.66e+00, 2.94e+00]
target1: 16.10 | loss: 4.47e-01 | Hard: True | Constraint met: False
target2: 100% of values outside [2.75e+00, 3.04e+00]
target2: 23.90 | loss: 5.71e-01 | Hard: False | Constraint met: False
Epoch 0 | (1/64) | Params: [-4.551, 2.191] | Error: 0.472
target1: 100% of values outside [2.66e+00, 2.94e+00]
target1: 15.94 | loss: 4.43e-01 | Hard: True | Constraint met: False
target2: 100% of values outside [2.75e+00, 3.04e+00]
target2: 23.39 | loss: 5.64e-01 | Hard: False | Constraint met: False
Epoch 0 | (2/64) | Params: [-4.468, 2.431] | Error: 0.467
target1: 100% of values outside [2.66e+00, 2.94e+00]
target1: 15.39 | loss: 4.30e-01 | Hard: True | Constraint met: False
target2: 100% of values outside [2.75e+00, 3.04e+00]
target2: 21.51 | loss: 5.36e-01 | Hard: False | Constraint met: False
Epoch 0 | (3/64) | Params: [-4.196, 2.901] | Error: 0.452
...
Epoch 11 | Mean Error: 0.000 | Sigma Error: 0.000
Epoch 11 | Mean Parameters: [2.105, -2.501] | Sigma parameters: [0.039, 0.202]
Epoch 11 | Normalised Sigma parameters: [0.015, 0.081]
Terminating after meeting termination criteria at epoch 12.
Note that verbosity can be controlled with verbose: bool option in evopt.optimize().
Parameter Space Sampling
In addition to optimization, evopt provides functionality for efficient parameter space exploration using Sobol sequences. This is useful for generating diverse sets of parameter values and understanding the response surface of your system before optimization.
import evopt
# Define your parameters and evaluator function
params = {
'x1': (-5, 5),
'x2': (-5, 5)
}
def evaluator(param_dict):
x1 = param_dict['x1']
x2 = param_dict['x2']
result = (1 - x1) ** 2 + 100*(x2 - x1 ** 2) ** 2
return result
# Sample 32 points using Sobol sampling
results = evopt.sample(
params=params,
evaluator=evaluator,
n_samples=32,
verbose=True
)
Sample output:
Running evaluations in serial mode.
Sample 0 | (1/32) | Params: [-2.500, -2.500] | Error: None
Sample 1 | (2/32) | Params: [2.500, 2.500] | Error: None
Sample 2 | (3/32) | Params: [-1.250, -1.250] | Error: None
Sample 3 | (4/32) | Params: [3.750, 3.750] | Error: None
...
The sampling function organizes results in a directory structure similar to the optimization function, with CSV files containing all sample data for further analysis.
Keywords for sample() Function
The evopt.sample() function takes several keyword arguments to control the sampling process:
params (dict): A dictionary defining the parameters to sample. Keys are parameter names, and values are tuples of(min, max)bounds.evaluator (Callable): A callable that evaluates the parameters and returns an error value or result dictionary.n_samples (int, optional): The number of Sobol samples to generate. Defaults to32.rand_seed (int, optional): Random seed for reproducible results. Defaults toNone.target_dict (dict, optional): Dictionary of target values for comparison. Defaults toNone.max_workers (int, optional): Maximum number of worker processes for parallel evaluation. Defaults to1.cores_per_worker (int, optional): Number of CPU cores per worker process. Defaults to1.base_dir (str, optional): Base directory for storing results. Defaults to the current working directory.dir_id (str, optional): Directory ID for organizing results. Defaults toNone.verbose (bool, optional): Whether to print detailed information during sampling. Defaults toTrue.
Keywords for optimize() Function
The evopt.optimize() function takes several keyword arguments to control the optimization process:
params (dict): A dictionary defining the parameters to optimize. Keys are parameter names, and values are tuples of(min, max)bounds.evaluator (Callable): A callable (usually a function) that evaluates the parameters and returns an error value. This function is the core of your optimization problem.optimizer (str, optional): The optimization algorithm to use. Currently, only 'cmaes' (Covariance Matrix Adaptation Evolution Strategy) is supported. Defaults to'cmaes'.base_dir (str, optional): The base directory where the optimization results (checkpoints, logs, CSV files) will be stored. If not specified, it defaults to the current working directory.dir_id (int, optional): A specific directory ID for the optimization run. If provided, the results will be stored in base_dir/evolve_{dir_id}. If not provided, a new unique ID will be generated automatically.sigma_threshold (float, optional): The threshold for the sigma values (step size) of the CMA-ES algorithm. The optimization will terminate when all sigma values are below this threshold, indicating convergence. Defaults to0.1.batch_size (int, optional): The number of solutions to evaluate in each epoch (generation) of the CMA-ES algorithm. A larger batch size can speed up the optimization but may require more computational resources. Defaults to16.start_epoch (int, optional): The epoch number to start from. This is useful for resuming an interrupted optimization run from a checkpoint. Defaults toNone.verbose (bool, optional): Whether to print detailed information about the optimization process to the console. IfTrue, the optimization will print information about each epoch and solution. Defaults toTrue.num_epochs (int, optional): The maximum number of epochs to run the optimization for. If specified, the optimization will terminate after this number of epochs, even if the convergence criteria (sigma_threshold) has not been met. If None, the optimization will run until the convergence criteria is met. Defaults toNone.max_workers (int, optional): The number of multi-processing workers to operate concurrently. Defaults to 1. Each worker operates on a different processor.rand_seed (int, optional): Specify the deterministic seed.hpc_cores_per_worker (int, optional): Number of CPU cores to allocate per HPC worker.hpc_memory_gb_per_worker (int): Memory in GB to allocate per worker on the HPC.hpc_wall_time (str): Wall time limit for each HPC worker, must be in the format "DD:HH:MM:SS" or "HH:MM:SS".hpc_qos (str): Quality of Service for HPC jobs.
Plotting convergence
Evopt provides an overview of the convergence for each parameter over the epochs, through the evopt.Plotting.plot_epochs() method.
# path to your evolve folder that contains epochs.csv and results.csv
evolve_dir = r"path\to\base\dir\evolve_0"
evopt.Plotting.plot_epochs(evolve_dir_path=evolve_dir)
Output:
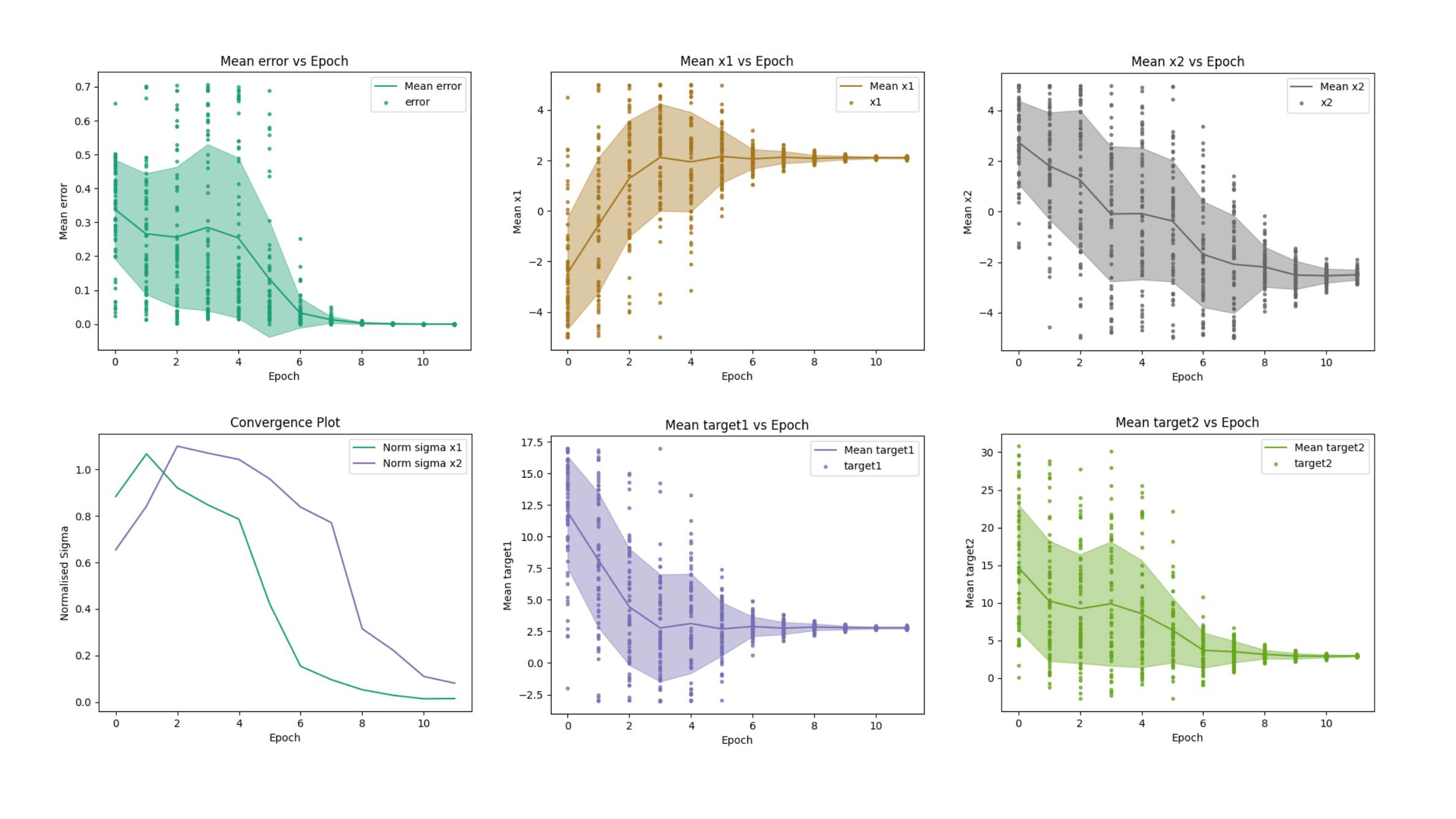
Convergence plots displaying error, parameters, targets, and normalised standard-deviation of the solution (normalised sigma) as a function of the number of epochs.
Plotting variables
Evopt also supports hassle free plotting of 1-D, 2-D, 3-D, and even 4-D results data using the same method: evopt.Plotting.plot_vars(). Simply specify the Evolve_{dir_id} file directory and the columns of the results.csv file you want to plot. By default the figures will save to Evolve_{dir_id}\figures.
2-D example (simple xy plot):
evopt.Plotting.plot_vars(evolve_dir_path=evolve_dir, x="x1", y="error")
Output:
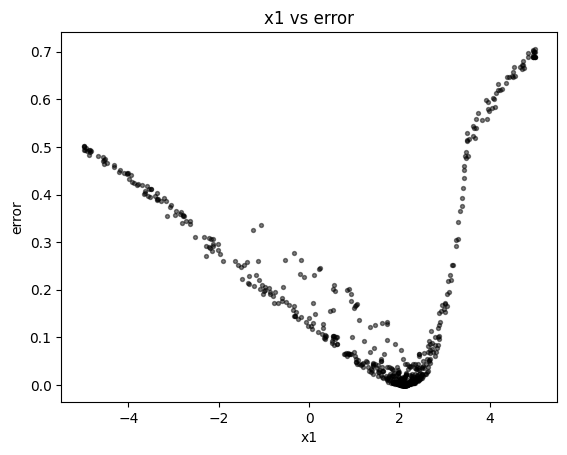
Scatter plot showing parameter versus error. The axis handle is returned to the user for any modifications.
2-D example (Voronoi plot):
evopt.Plotting.plot_vars(evolve_dir_path=evolve_dir, x="x1", y="x2", cval="error")
Output:
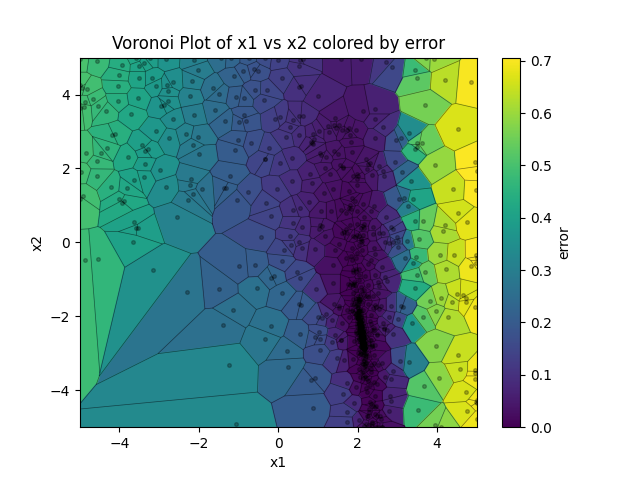
2-D Voronoi plot illustrating parameters versus error. Each cell contains a single solution, with cell line is equidistant between points on either size. In this sense the plot conveys the exploration/explotation nature of the evolutionary algorithm as it hones in on the global optimum. The axis handle is returned to the user for any modifications.
4-D example (interactive html surface plot with color)
evopt.Plotting.plot_vars(evolve_dir_path=evolve_dir, x="x1", y="x2", z="error", cval="epoch")
Output:
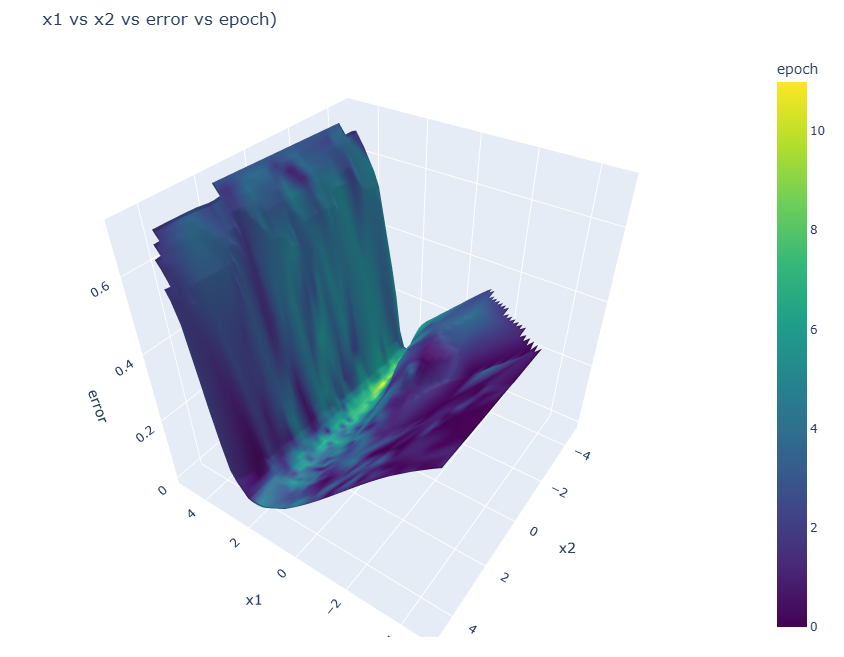
3-D surface plot of the parameters versus the error values, coloured by epoch. As is the nature of convergent optimization, the latest epochs show the lowest error values.
Directory Structure
When you run an optimization with evopt, it creates the following directory structure to organise the results:
Each evaluation function call operates in its respective solution directory. This means that files can be created locally without needing absolute paths.
For example:
def evaluator(dict_params:dict) -> float:
...
with open("your_file.txt", 'a') as f:
f.write(error)
...
return error
Would result in the creation of a file "your_file.txt" in each solution folder:
base_directory/
└── evolve_{dir_id}/
├── epochs/
│ └── epoch0000/
│ └── solution0000/
| └── your_file.txt
│ └── solution0001/
| └── your_file.txt
│ └── ...
│ └── epoch0001/
│ └── ...
│ └── ...
├── checkpoints/
│ └── checkpoint_epoch0000.pkl
│ └── checkpoint_epoch0001.pkl
│ └── ...
├── logs/
│ └── logfile.log
├── epochs.csv
└── results.csv
base_directory: This is the base directory where the optimization runs are stored. If not specified, it defaults to the current working directory.evolve_{dir_id}: Each optimization run gets its own directory namedevolve_{dir_id}, wheredir_idis a unique integer.epochs: This directory contains subdirectories for each epoch of the optimization.epoch####: Each epoch directory contains subdirectories for each solution evaluated in that epoch. Epoch folders are only produced if solution files contain files.solution####: Each solution directory can contain files generated by the evaluator function for that specific solution. Solution folders are only produced if files are created during an evaluation.checkpoints: This directory stores checkpoint files, allowing you to resume interrupted optimization runs.logs: This directory contains the log file (logfile.log) which captures the output of the optimization process.epochs.csv: This file contains summary statistics for each epoch, such as mean error, parameter values, and sigma values.results.csv: This file contains the results for each solution evaluated during the optimization, including parameter values and the corresponding error.
Citing
If you publish research making use of this library, we encourage you to cite this repository:
Hart-Villamil, R. (2024). Evopt, simple but powerful gradient-free numerical optimization.
This library makes fundamental use of the pycma implementation of the state-of-the-art CMA-ES algorithm.
Hence we kindly ask that research using this library cites:
Nikolaus Hansen, Youhei Akimoto, and Petr Baudis. CMA-ES/pycma on Github. Zenodo, DOI:10.5281/zenodo.2559634, February 2019.
This work was also inspired by 'ACCES', a package for derivative-free numerical optimization designed for simulations.
Nicusan, A., Werner, D., Sykes, J. A., Seville, J., & Windows-Yule, K. (2022). ACCES: Autonomous Characterisation and Calibration via Evolutionary Simulation (Version 0.2.0) [Computer software]
The symbolic regression functionality of this package is built upon PySR. We ask that research cites:
Cranmer, M. (2023). Interpretable machine learning for science with PySR and SymbolicRegression. jl. arXiv preprint arXiv:2305.01582.
License
This project is licensed under the GNU General Public License v3.0 License.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file evopt-0.15.0.tar.gz.
File metadata
- Download URL: evopt-0.15.0.tar.gz
- Upload date:
- Size: 1.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
32a7eea79307d652b0f76b1d556c9d87babdc7d026fd3952c120be35dc3e99e5
|
|
| MD5 |
03aa86ad2cbd0f983f3a01869609293e
|
|
| BLAKE2b-256 |
a829adc5ae39b630f60b34abe6e0b5c2130979be7807fca584e0e3a8cfb14b58
|
File details
Details for the file evopt-0.15.0-py3-none-any.whl.
File metadata
- Download URL: evopt-0.15.0-py3-none-any.whl
- Upload date:
- Size: 75.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7696e4697d9dba527320d1a5ee273b598fdf806dc13a22c92ea80a636295e088
|
|
| MD5 |
8f4d39d4c1729bdbe4b98cc9269458ef
|
|
| BLAKE2b-256 |
95788166c560e77d3232eeefb0732fc0cbf0d05e477cd031c77ec5b373739c8b
|











