A flexible artifact correction and evaluation toolbox to correct eeg data
Project description
FACETpy — fMRI Artifact Correction and Evaluation Toolbox
A Python toolbox for correcting fMRI-induced EEG artifacts using Averaged Artifact Subtraction (AAS) and other advanced methods. Built on MNE-Python, it provides a modular pipeline architecture that lets researchers process, evaluate, and compare correction results with minimal code.
Key features
- Load EEG from EDF, GDF, and BIDS formats
- Artifact correction: AAS, PCA, Adaptive Noise Cancellation (ANC)
- Full evaluation suite: SNR, RMS, Median Artifact, FFT-based metrics
- One-call results:
result.print_metrics(),result.print_summary() - Batch processing across subjects/sessions with
Pipeline.map() - Generate synthetic EEG for algorithm testing
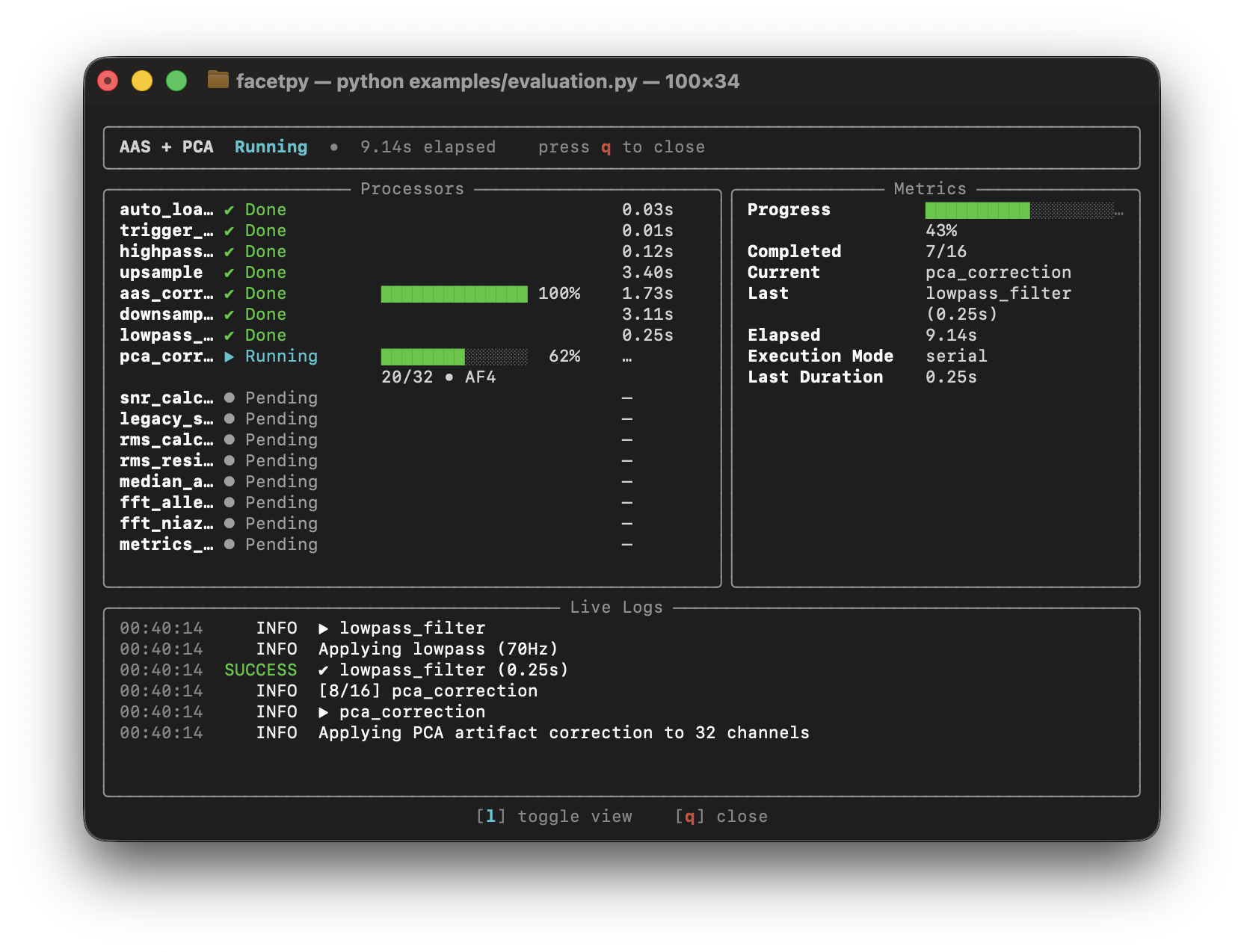
- Rich progress display in the terminal
Quick start
Quick installation (requires Python 3.11/3.12/3.13):
Unix (macOS/Linux):
curl -fsSL https://raw.githubusercontent.com/H0mire/facetpy/main/scripts/bootstrap.sh | sh
cd facetpy
Windows (PowerShell):
git clone https://github.com/H0mire/facetpy.git
Set-Location facetpy
poetry install --no-interaction
from facet import (
Pipeline, Loader, EDFExporter,
TriggerDetector, UpSample, DownSample, AASCorrection,
)
pipeline = Pipeline([
Loader(path="data.edf", preload=True),
TriggerDetector(regex=r"\b1\b"),
UpSample(factor=10),
AASCorrection(window_size=30),
DownSample(factor=10),
EDFExporter(path="corrected.edf", overwrite=True),
], name="Quickstart")
result = pipeline.run()
result.print_summary() # Done in 4.2s snr=18.3 rms_ratio=0.14
Installation
Requires Python 3.11, 3.12, or 3.13 and Poetry >= 1.8. Conda is optional, not required.
Option A (PyPI): install from package index
pip install facetpy
The package name on PyPI is facetpy; import it in Python as facet.
Option B (Unix recommended): bootstrap script
curl -fsSL https://raw.githubusercontent.com/H0mire/facetpy/main/scripts/bootstrap.sh | sh
cd facetpy
This bootstrap script:
- clones the FACETpy repository into
./facetpy - runs
./scripts/install.shinside that clone
Option C (Unix): clone first, then run local installer
git clone https://github.com/H0mire/facetpy.git
cd facetpy
./scripts/install.sh
The script:
- checks for Python 3.11/3.12/3.13
- checks whether Poetry is installed
- asks whether Poetry should be installed if missing
- runs
poetry install --no-interaction
Option D (Windows PowerShell): manual install
git clone https://github.com/H0mire/facetpy.git
Set-Location facetpy
poetry install --no-interaction
Use Option B/C inside WSL or Git Bash if you want to run the .sh installer scripts.
Option E: system Python + Poetry (manual)
Unix (macOS/Linux):
# 1 — verify Python
python3 --version
# 2 — install Poetry (pick one)
pipx install poetry
# or: python3 -m pip install --user poetry
# 3 — install FACETpy and dependencies
poetry install --no-interaction
Windows (PowerShell):
# 1 — verify Python
py --version
# 2 — install Poetry (pick one)
pipx install poetry
# or: py -m pip install --user poetry
# 3 — install FACETpy and dependencies
poetry install --no-interaction
Option F (optional): Conda workflow
Unix (macOS/Linux):
conda create -n facetpy python=3.13 -y
conda activate facetpy
conda install -c conda-forge poetry -y
poetry install --no-interaction
Windows (PowerShell):
conda create -n facetpy python=3.13 -y
conda activate facetpy
conda install -c conda-forge poetry -y
poetry install --no-interaction
Optional extras:
poetry install -E deeplearning # TensorFlow-based models
poetry install -E notebooks # Jupyter notebook support
poetry install -E gui # PyQt6 GUI components
poetry install -E docs # Sphinx documentation toolchain
poetry install -E all # everything above
Run commands with poetry run ... (for example, poetry run python examples/quickstart.py).
Build the C extension (optional)
The fast Adaptive Noise Cancellation (ANC) step uses a compiled C extension. Build it once after installing:
poetry run build-fastranc
If the extension is absent, ANC is skipped automatically and the rest of the toolbox works normally.
Running the examples
All examples are in the examples/ folder and use the bundled
NiazyFMRI.edf dataset. Run them from the project root:
# Recommended order for new users:
poetry run python examples/quickstart.py # minimal pipeline
poetry run python examples/evaluation.py # metrics & comparison
poetry run python examples/advanced_workflows.py # conditional, parallel, factory
poetry run python examples/batch_processing.py # multiple files at once
poetry run python examples/inline_steps.py # custom steps & pipe operator
poetry run python examples/complete_pipeline_example.py # full clinical pipeline
poetry run python examples/eeg_generation_visualization_example.py # synthetic EEG
Testing
# Run the full test suite
poetry run pytest
# Only fast unit tests (skip slow integration tests)
poetry run pytest -m "not slow"
# A single test file
poetry run pytest tests/test_core_pipeline.py -v
# With coverage report
poetry run pytest --cov=facet --cov-report=html
Open the coverage report:
Unix (macOS/Linux):
open htmlcov/index.html
Windows (PowerShell):
start htmlcov/index.html
Documentation
# Install docs dependencies
poetry install -E docs
# Build HTML docs
poetry run sphinx-build -b html docs/source docs/build
Open docs locally:
Unix (macOS/Linux):
open docs/build/index.html
Windows (PowerShell):
start docs/build/index.html
Full online documentation: https://facetpy.readthedocs.io/
For comprehensive build instructions, theme configuration, and contribution guidelines see docs/README.md.
For PyPI release steps, see RELEASING.md.
Project structure
src/facet/
├── core/ Pipeline, Processor, ProcessingContext, BatchResult
├── io/ Loader, BIDSLoader, EDFExporter, BIDSExporter
├── preprocessing/ Filters, Resample, TriggerDetector, Alignment, Transforms
├── correction/ AASCorrection, PCACorrection, ANCCorrection
├── evaluation/ SNRCalculator, RMSCalculator, MetricsReport, RawPlotter
├── misc/ EEGGenerator (synthetic data)
└── pipelines.py create_standard_pipeline() factory
examples/
├── quickstart.py minimal pipeline
├── evaluation.py metrics & comparison
├── advanced_workflows.py conditional, parallel, factory
├── batch_processing.py multiple files
├── inline_steps.py custom steps & pipe operator
├── complete_pipeline_example.py full clinical pipeline
└── eeg_generation_visualization_example.py synthetic EEG
VS Code Tasks
Tasks are defined in .vscode/tasks.json and can be run via Ctrl+Shift+P → Tasks: Run Task.
| Task | Shortcut | Description |
|---|---|---|
| Test: Run All | default test task | Full test suite with coverage report |
| Test: Run Current File | Run pytest on the file open in the editor | |
| Test: Unit Only | Only tests marked @pytest.mark.unit |
|
| Test: Integration Only | Only tests marked @pytest.mark.integration |
|
| Test: Skip Slow | All tests except those marked @pytest.mark.slow |
|
| Test: Show Coverage Report | Open htmlcov/index.html in the browser |
|
| Lint: Check (Ruff) | Check src/ and tests/ for lint errors |
|
| Lint: Fix (Ruff) | Auto-fix lint errors in place | |
| Format: Check (Ruff) | Verify formatting without changing files | |
| Format: Apply (Ruff) | Apply ruff formatting to src/ and tests/ |
|
| Build: FastRANC C Extension | Compile the FastRANC C extension | |
| Build: Install Dependencies | poetry install |
|
| Build: Install All Extras | poetry install -E all |
|
| Build: Update Dependencies | poetry update |
|
| Docs: Build HTML | Build Sphinx documentation | |
| Docs: Open in Browser | Open the built docs in the browser | |
| Docs: Build & Open | Build docs and open immediately | |
| Run: Current Python File | Execute the file open in the editor | |
| Review: Uncommitted Changes (Codex) | Codex AI review of all local changes | |
| Review: Against Branch (Codex) | Codex AI review against a selected base branch (prompts for branch) | |
| QA: Full Check (Lint + Format + Test) | Lint + format check + full test suite in sequence |
License
GPLv3 — see LICENSE for details.
Author: Janik Michael Mueller
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file facetpy-2.0.1.tar.gz.
File metadata
- Download URL: facetpy-2.0.1.tar.gz
- Upload date:
- Size: 163.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ef8288aadae63276824bf422f0d2aab4e2a953240a3eed48048f7a935efd7765
|
|
| MD5 |
8cb002525561f8106113ca142e44e6ac
|
|
| BLAKE2b-256 |
d8cd81c296cfe09c093ae94d41b2efd8cf2ed8408165d672a8241e726487a993
|
Provenance
The following attestation bundles were made for facetpy-2.0.1.tar.gz:
Publisher:
publish.yml on H0mire/FACETpy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
facetpy-2.0.1.tar.gz -
Subject digest:
ef8288aadae63276824bf422f0d2aab4e2a953240a3eed48048f7a935efd7765 - Sigstore transparency entry: 1013076384
- Sigstore integration time:
-
Permalink:
H0mire/FACETpy@a234fec53c347fc8dbe5717906fe0e7ae8332083 -
Branch / Tag:
refs/tags/v2.0.1 - Owner: https://github.com/H0mire
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@a234fec53c347fc8dbe5717906fe0e7ae8332083 -
Trigger Event:
push
-
Statement type:
File details
Details for the file facetpy-2.0.1-py3-none-any.whl.
File metadata
- Download URL: facetpy-2.0.1-py3-none-any.whl
- Upload date:
- Size: 190.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
df6cc18c08883fb256324d46d2e020405c532db92d73a0741e6fbbe4acb750e5
|
|
| MD5 |
361161ac2bd61c65c94a9840d712225e
|
|
| BLAKE2b-256 |
db0a79696dbb91fb4f75022001b9827fa7d10d7eb0b93031613e1ef459d633e6
|
Provenance
The following attestation bundles were made for facetpy-2.0.1-py3-none-any.whl:
Publisher:
publish.yml on H0mire/FACETpy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
facetpy-2.0.1-py3-none-any.whl -
Subject digest:
df6cc18c08883fb256324d46d2e020405c532db92d73a0741e6fbbe4acb750e5 - Sigstore transparency entry: 1013076403
- Sigstore integration time:
-
Permalink:
H0mire/FACETpy@a234fec53c347fc8dbe5717906fe0e7ae8332083 -
Branch / Tag:
refs/tags/v2.0.1 - Owner: https://github.com/H0mire
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@a234fec53c347fc8dbe5717906fe0e7ae8332083 -
Trigger Event:
push
-
Statement type:












