Fast, easy-to-use causal discovery analysis tools
Project description
fastcausal
Fast, easy-to-use causal discovery analysis tools for Python.
Overview
fastcausal provides a unified Python interface for causal discovery analysis, combining the functionality of several earlier packages into one pip-installable tool. It supports both interactive Jupyter notebook workflows and config-driven batch processing of large datasets.
Key features:
- No Java dependency — uses tetrad-port (C++ port of Tetrad algorithms) instead of Java
- Seven causal discovery algorithms — PC, FGES, GFCI, BOSS, BOSS-FCI, GRaSP, GRaSP-FCI
- Prior knowledge support — temporal tiers, forbidden/required edges
- Bootstrapped stability analysis — edge frequency selection across subsampled runs
- SEM fitting — automatic structural equation modeling via semopy
- Flexible graph visualization — node styling with fnmatch patterns, multi-graph comparison with shared layouts
- Batch pipeline — config-driven processing of hundreds of cases via CLI
- Report generation — automated Word document reports with embedded graphs
Installation
pip install fastcausal # core package
pip install fastcausal[sem] # + SEM fitting (semopy)
pip install fastcausal[jupyter] # + Jupyter/matplotlib/seaborn
pip install fastcausal[batch] # + Word report generation
pip install fastcausal[all] # everything
Quick Start
Five lines to your first causal graph:
from fastcausal import FastCausal
fc = FastCausal()
df = fc.load_sample("boston") # bundled EMA dataset
df = fc.standardize(df)
results, graph = fc.run_search(df, algorithm="gfci", alpha=0.01, penalty_discount=1.0)
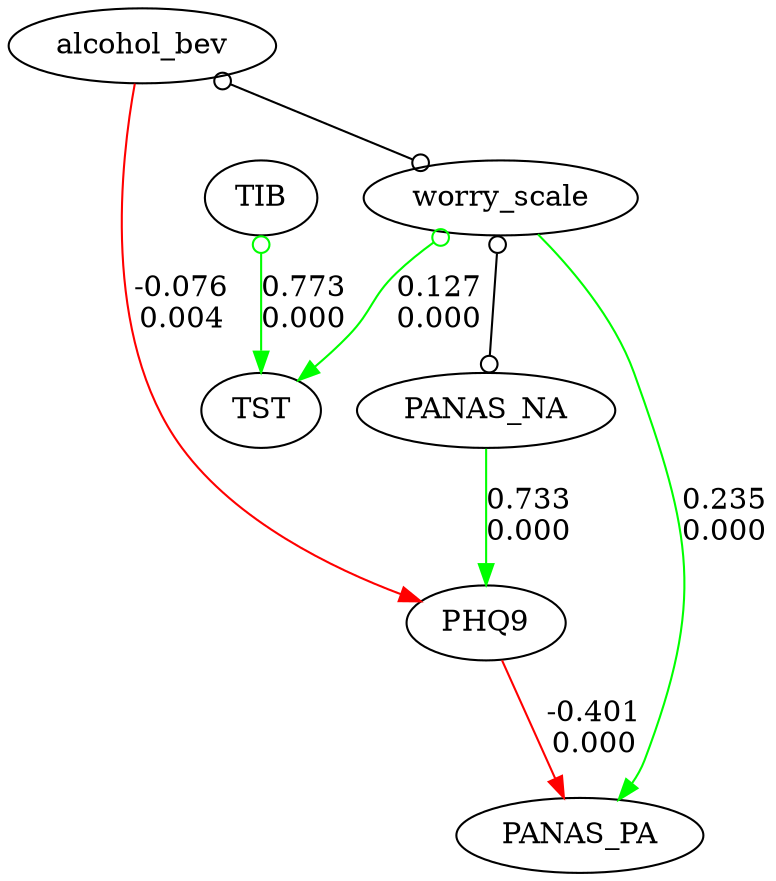
fc.show_graph(graph)
Time-series workflow with prior knowledge
For time-series data, add lagged columns, standardize, and encode temporal ordering so that yesterday's values can only be causes (not effects) of today's:
from fastcausal import FastCausal
fc = FastCausal()
df = fc.load_sample("boston")
# Add lagged columns and standardize
lag_stub = "_lag"
df_lag = fc.add_lag_columns(df, lag_stub=lag_stub)
df_std = fc.standardize(df_lag)
# Build temporal prior knowledge explicitly:
# Tier 0 (lag vars) can only be parents of Tier 1 (current-day vars)
cols = df.columns
knowledge = {
"addtemporal": {
0: [col + lag_stub for col in cols],
1: [col for col in cols],
}
}
# knowledge =>
# {"addtemporal": {
# 0: ["alcohol_bev_lag", "TIB_lag", "TST_lag", "PANAS_PA_lag",
# "PANAS_NA_lag", "worry_scale_lag", "PHQ9_lag"],
# 1: ["alcohol_bev", "TIB", "TST", "PANAS_PA",
# "PANAS_NA", "worry_scale", "PHQ9"]
# }}
# Run GFCI causal discovery with SEM fitting
result, graph = fc.run_search(
df_std,
algorithm="gfci",
alpha=0.01,
penalty_discount=1.0,
knowledge=knowledge,
)
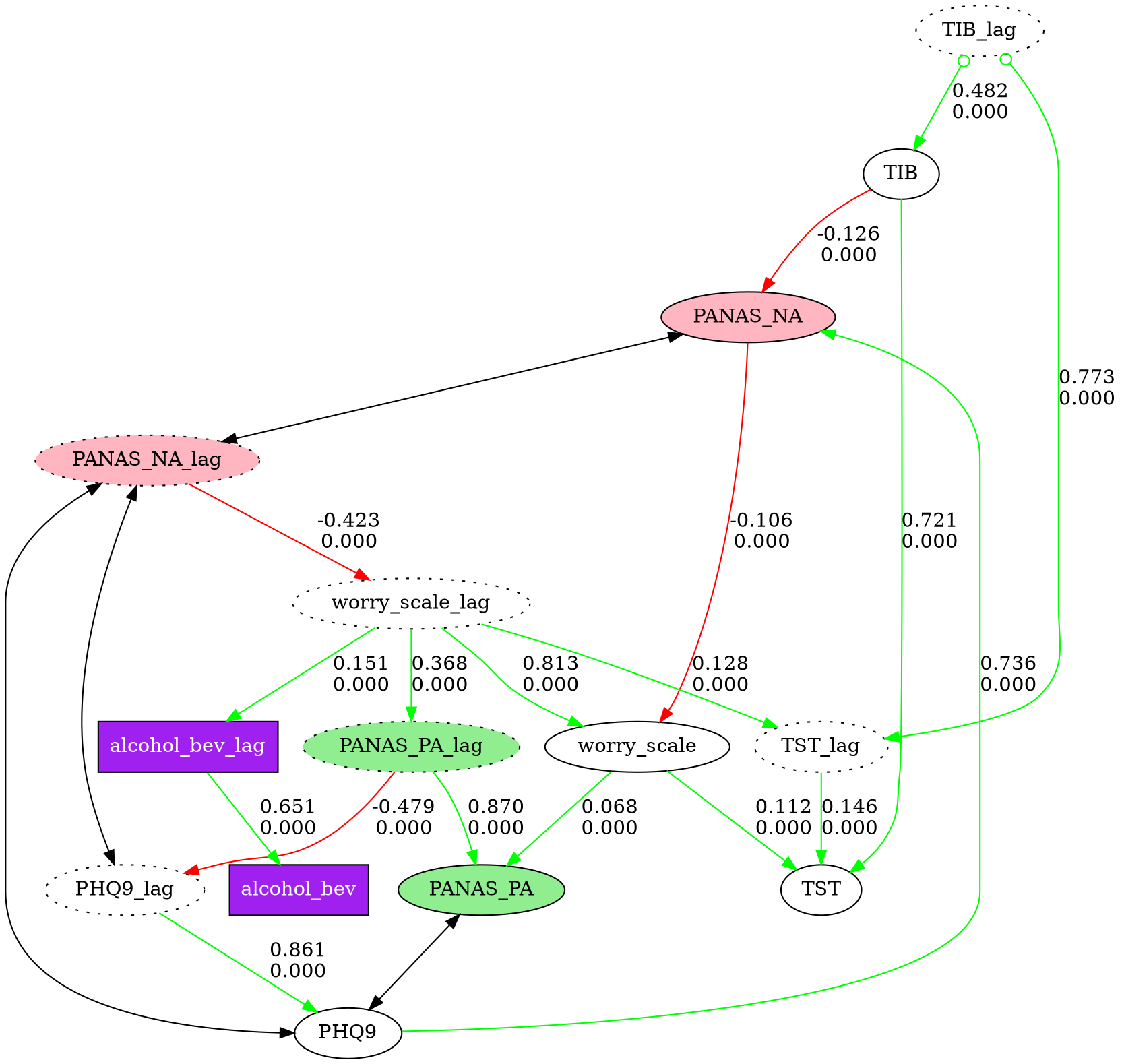
# Visualize with custom node styles
node_styles = [
{"pattern": "*_lag", "style": "dotted"},
{"pattern": "PANAS_PA*", "style": "filled", "fillcolor": "lightgreen"},
{"pattern": "PANAS_NA*", "style": "filled", "fillcolor": "lightpink"},
{"pattern": "alcohol_bev*", "shape": "box", "style": "filled",
"fillcolor": "purple", "fontcolor": "white"},
]
fc.show_graph(graph, node_styles=node_styles)
See fastcausal_demo_short.ipynb for the full interactive demo.
CLI Usage
fastcausal provides a command-line interface for batch processing:
# Data preparation
fastcausal parse --config proj/config.yaml
# Batch causal discovery across cases
fastcausal run --config proj/config.yaml
fastcausal run --config proj/config.yaml --start 0 --end 50
fastcausal run --config proj/config.yaml --list
# Effect size analysis and heatmaps
fastcausal paths --config proj/config.yaml
# Generate Word report
fastcausal report --config proj/config.yaml --mode 2wide
# Quick single-file analysis
fastcausal analyze data.csv --algorithm gfci --output results/
Supported Algorithms
| Algorithm | Type | Output | Key Parameters |
|---|---|---|---|
| PC | Constraint-based (Fisher Z) | CPDAG | alpha |
| FGES | Score-based (BIC) | CPDAG | penalty_discount |
| GFCI | Hybrid (FGES + FCI rules) | PAG | alpha, penalty_discount |
| BOSS | Permutation-based (BIC) | CPDAG | penalty_discount |
| BOSS-FCI | BOSS + FCI rules | PAG | alpha, penalty_discount |
| GRaSP | Permutation-based (tuck DFS) | CPDAG | penalty_discount |
| GRaSP-FCI | GRaSP + FCI rules | PAG | alpha, penalty_discount |
See the Algorithm Guide for detailed parameter reference, edge types, and selection guidance.
Architecture
fastcausal consolidates four earlier codebases into a layered architecture:
pip install fastcausal
|
fastcausal (API + CLI + viz + SEM + batch)
/ \
tetrad-port dgraph_flex
(C++ algorithms) (graph rendering)
- tetrad-port — C++ port of CMU Tetrad algorithms, exposed via nanobind
- dgraph_flex — Graphviz-based directed graph rendering
Project Structure
fastcausal/
├── core.py # FastCausal class (main API)
├── search.py # Algorithm wrapper (PC, FGES, GFCI, BOSS, GRaSP, ...)
├── sem.py # SEM fitting via semopy
├── transform.py # Lag columns, standardization, subsampling
├── knowledge.py # Prior knowledge handling
├── edges.py # Edge parsing, selection, deduplication
├── cli.py # Click-based CLI
├── viz/
│ ├── styling.py # fnmatch-based node styling
│ ├── graphs.py # Graph display and save (single + multi)
│ └── plots.py # Heatmaps and effect size plots
├── pipeline/
│ ├── config.py # YAML config parsing (v4.0 + v5.0)
│ ├── parse.py # Data preparation engine
│ ├── batch.py # Batch causal discovery
│ ├── paths.py # Effect size analysis
│ ├── report.py # Word document generation
│ └── metrics.py # Graph metrics (centrality, ancestors)
└── io/
├── data.py # CSV loading, sample datasets
└── wearables.py # Fitbit/Garmin integration (planned)
Documentation
- Algorithm Guide — Algorithm selection, parameters, edge types, prior knowledge
- Migration from fastcda — API mapping for fastcda users
- Migration from cda_tools2 — CLI mapping for cda_tools2 users
- Consolidation Plan — Implementation plan and phase status
- Project Conventions — Development guidelines and conventions
Config File Format
fastcausal uses YAML config files for batch processing. Version 5.0 is the current format; version 4.0 (from cda_tools2) is accepted with a deprecation warning.
GLOBAL:
version: 5.0
name: my_project
title: "My Causal Analysis"
CAUSAL:
algorithm: gfci
alpha: 0.05
penalty_discount: 1.0
knowledge: prior.txt
standardize_cols: true
Requirements
- Python >= 3.11
- tetrad-port >= 0.1.0
- dgraph_flex >= 0.1.11
- Graphviz (system install for graph rendering)
License
MIT
Citation
If you use fastcausal in your research, please cite the relevant algorithm papers and this package.
The bundled "boston" EMA dataset is from:
Cunningham TJ, Fields EC, Kensinger EA. "Boston College daily sleep and well-being survey data during early phase of the COVID-19 pandemic." Sci Data. 2021. https://www.nature.com/articles/s41597-021-00886-y
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file fastcausal-0.1.6.tar.gz.
File metadata
- Download URL: fastcausal-0.1.6.tar.gz
- Upload date:
- Size: 51.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
03a5b0ee87a5a2eb17ed2b33668c9696d19555a4f3bf21588587a2ff6b0b17fa
|
|
| MD5 |
51bd28e4fd0081dc1155bdde7604e474
|
|
| BLAKE2b-256 |
1b1347352ebb10ac04c3df45a179db3e55e76f3a6a71f7640ba71ba63454becf
|
File details
Details for the file fastcausal-0.1.6-py3-none-any.whl.
File metadata
- Download URL: fastcausal-0.1.6-py3-none-any.whl
- Upload date:
- Size: 47.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
840a1ee91704e1b20f0e7239138fc3892d0d2d6668abbe2a7725760be0c24297
|
|
| MD5 |
4e60d16c37879e940d1a78f8180917e8
|
|
| BLAKE2b-256 |
80ffd88afb71414337fbd6a08c6687df0b380df99956bb9ece54263d6f6b4f8a
|