A PyQt/PySide widget to display crystal structures
Project description
fastmolwidget
A PyQt/PySide6 widget to display crystal structures
fastmolwidget is a lightweight, embeddable Qt widget that renders molecular and crystal structures in both 2D projection and 3D OpenGL. It supports anisotropic displacement parameter (ADP) ellipsoids, ball-and-stick diagrams, and plain sphere representations. The 2D backend uses a pure-Python QPainter renderer (no OpenGL required); the 3D backend uses hardware-accelerated OpenGL with sphere and ellipsoid impostors.
Screenshots
| 2D (QPainter) | 3D (OpenGL) |
|---|---|
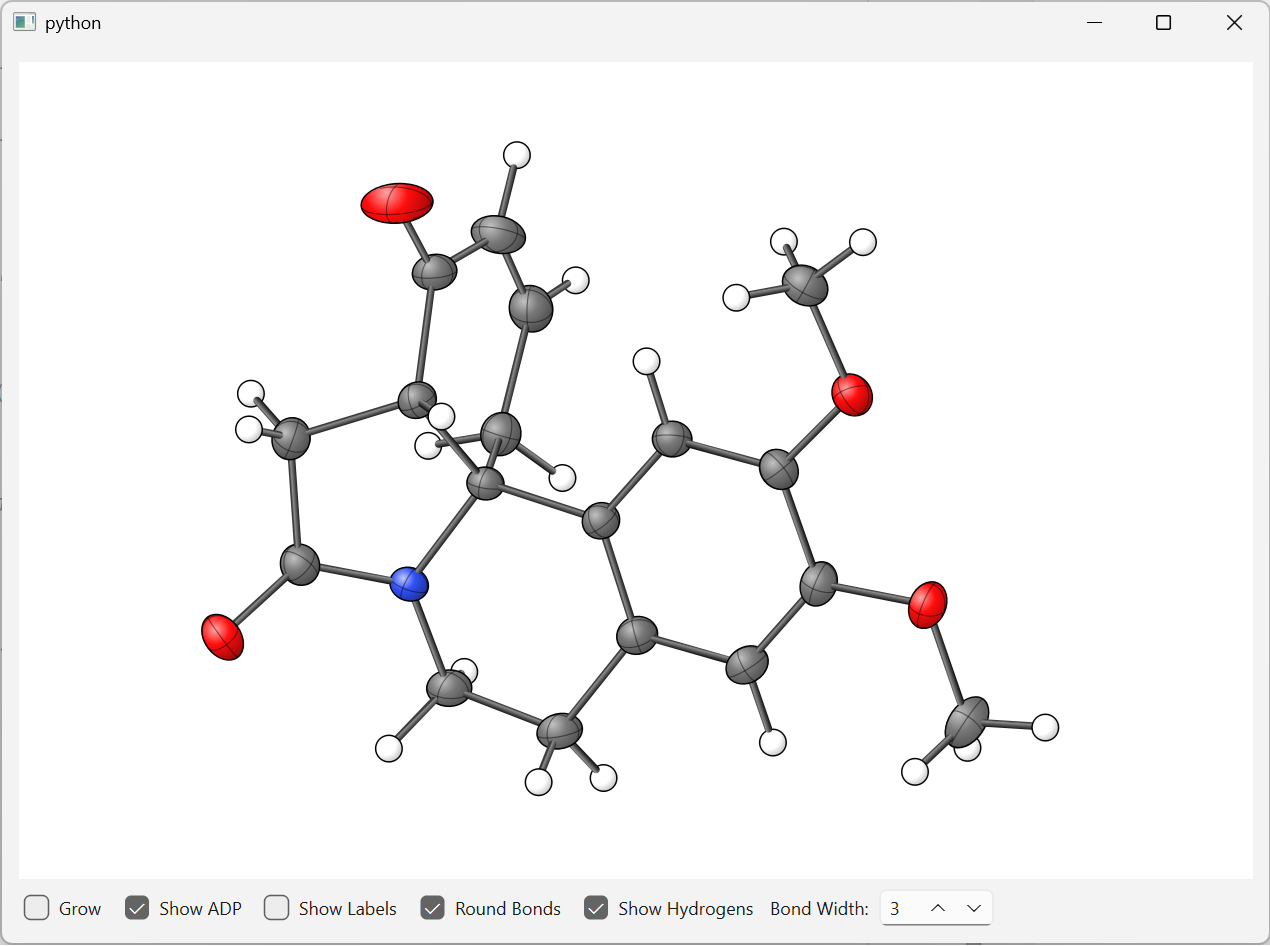 |
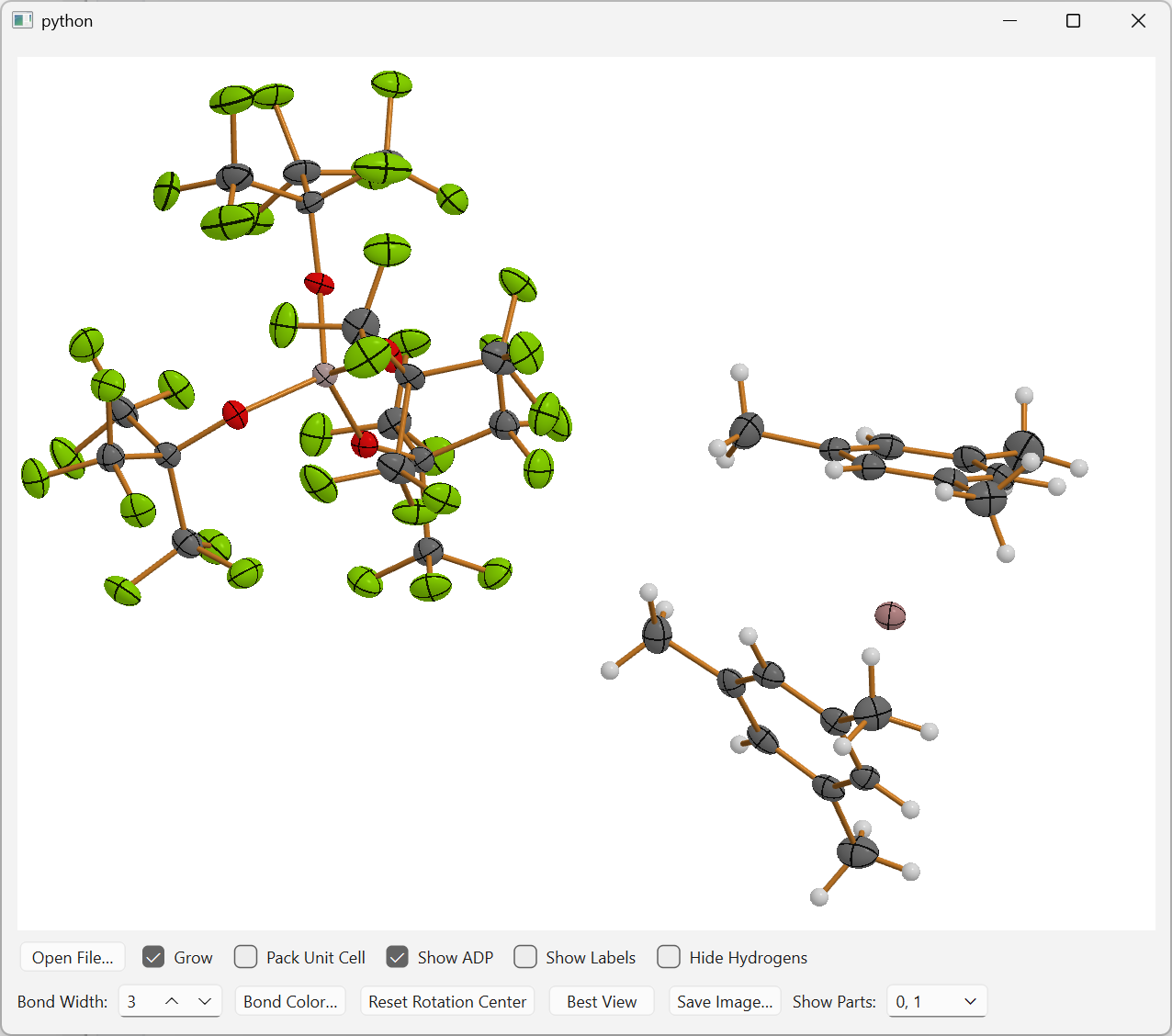 |
| ORTEP-style crystal structure with ADP ellipsoids (2D QPainter backend) | Real-time 3D ball-and-stick view with depth-shaded spheres and cylinder bonds (OpenGL backend) |
Features
- ADP ellipsoids at the 50 % probability level
- Ball-and-stick and isotropic sphere
- Real-time 3D rendering via
MoleculeWidget3D— sphere impostors and tessellated cylinder bonds in hardware-accelerated OpenGL - Interactive mouse controls: rotate (left-drag), zoom (right-drag), pan (middle-drag), scroll wheel to resize labels
- Atom and bond selection: single click or Ctrl+click for multi-selection; emits
atomClicked/bondClickedQt signals - Hover labels: hovering over an atom shows its label; hovering over a bond shows the distance in Ångströms
- Hydrogen visibility toggle
- Atom label display toggle with adjustable font size
- Bond width adjustment via spin box
- Configurable bond color — set programmatically or via the control-bar color picker
- Multiple file formats: CIF, SHELX
.res/.ins, and plain XYZ. More to come... - Embeddable — both
MoleculeWidget(2D) andMoleculeWidget3D(3D) are plainQWidgetsubclasses; drop either into any layout - Ready-to-use viewers —
MoleculeViewerWidget(2D) andMoleculeViewer3DWidget(3D) bundle the renderer with a full control bar - Common protocol —
MoleculeWidgetProtocollets you write code that works with either widget interchangeably
Supported File Formats
| Extension | Format | Notes |
|---|---|---|
.cif |
Crystallographic Information File | Reads atoms, unit cell, and ADPs |
.res / .ins |
SHELXL instruction file | Reads atoms and unit cell via shelxfile |
.xyz |
Standard XYZ coordinate file | Cartesian coordinates, no cell or ADPs |
Installation
# with PySide6 (recommended)
uv add "fastmolwidget[pyside6]"
# or PyQt6
uv add "fastmolwidget[pyqt6]"
# add 3D OpenGL support (optional, requires Qt ≥ 6.7 and pyopenGL installed in the Python environment)
uv add "fastmolwidget[pyside6,gl3d]"
Optional C++ Acceleration (sdm_cpp)
The symmetry-growing step (SDM) has an optional C++ extension that uses pybind11 and OpenMP for a significant speed-up on large structures. The pure-Python fallback is always available.
uv pip install pybind11
uv pip install -e . --no-build-isolation
# macOS: optionally install libomp for multi-threaded acceleration
brew install libomp
Requirements: Python ≥ 3.12, NumPy, gemmi, shelxfile, qtpy, and either PySide6 or PyQt6.
Quick Start
Standalone 2D viewer
from qtpy.QtWidgets import QApplication
from fastmolwidget import MoleculeViewerWidget
app = QApplication([])
viewer = MoleculeViewerWidget()
viewer.load_file("structure.cif")
viewer.show()
app.exec()
Standalone 3D viewer
from qtpy.QtWidgets import QApplication
from fastmolwidget import MoleculeViewer3DWidget
app = QApplication([])
viewer = MoleculeViewer3DWidget()
viewer.load_file("structure.cif")
viewer.show()
app.exec()
Embedding the 3D widget in your own layout
from fastmolwidget import MoleculeWidget3D
mol = MoleculeWidget3D(parent=self)
mol.open_molecule(atoms, cell=cell)
layout.addWidget(mol)
Embedding the 2D widget in your own layout
from fastmolwidget import MoleculeWidget, MoleculeLoader
mol = MoleculeWidget(parent=self)
loader = MoleculeLoader(mol)
# The loader recognizes the file format from the extension and populates `mol` accordingly
loader.load_file("structure.cif")
# drop `mol` into any QLayout
layout.addWidget(mol)
Loading a different file at runtime
viewer.load_file("new_structure.res")
Reacting to atom / bond clicks
mol.atomClicked.connect(lambda label: print(f"Clicked atom: {label}"))
mol.bondClicked.connect(lambda a, b: print(f"Clicked bond: {a}–{b}"))
Mouse Controls
| Action | Effect |
|---|---|
| Left-drag | Rotate the molecule |
| Right-drag | Zoom in / out |
| Middle-drag | Pan the view |
| Middle-click | Recentre the rotation pivot on the clicked atom (3D only) |
| Alt/Option + Left-click | On systems without a middle mouse button, Alt/Option + Left-click recentres the rotation pivot on the clicked atom (same as Middle-click) |
| Scroll wheel | Increase / decrease label font size |
| Left-click | Select a single atom or bond |
| Ctrl + Left-click | Toggle multi-selection |
| Hover over atom | Show the atom label (enlarged when persistent labels are on) |
| Hover over bond | Show the bond distance (Å) in a rounded tooltip near the cursor |
Keyboard Shortcuts
The widget must have keyboard focus (click on it once) for these shortcuts to work.
| Key | Effect |
|---|---|
| F1 | Align the view so that the reciprocal axis a* points towards the viewer (requires a unit cell) |
| F2 | Align the view so that the reciprocal axis b* points towards the viewer (requires a unit cell) |
| F3 | Align the view so that the reciprocal axis c* points towards the viewer (requires a unit cell) |
Note: The F-key shortcuts are available in both the 2D (
MoleculeWidget) and 3D (MoleculeWidget3D) renderers. They have no effect when no unit cell is loaded (e.g. plain XYZ files).
Control Bar Options
MoleculeViewerWidget (2D) and MoleculeViewer3DWidget (3D)
Both viewers expose the same two-row control bar:
Row 1 — structure toggles
| Control | Default | Description |
|---|---|---|
| Open File… | — | Opens a file dialog to load a structure file |
| Grow | ✗ | Expand the asymmetric unit to complete molecules (mutually exclusive with Pack Unit Cell) |
| Pack Unit Cell | ✗ | Generate all symmetry-equivalent positions within one unit cell (mutually exclusive with Grow) |
| Show ADP | ✓ | Toggle ORTEP ellipsoid / isotropic sphere rendering |
| Show Labels | ✗ | Toggle non-hydrogen atom labels |
| Hide Hydrogens | ✗ | When checked, hydrogen atoms and their bonds are hidden |
Row 2 — bond and view controls
| Control | Default | Description |
|---|---|---|
| Bond Width | 3 | Stroke width / cylinder radius for bonds (2D: 1–15, 3D: 0–15) |
| Bond Color | — | Opens a colour picker to change the default bond colour |
| Reset Rotation Center | — | Restores the rotation pivot to the molecule's geometric centre (both 2D and 3D) |
| Best View | — | Rotates the current structure to a visibility-optimized orientation (PCA on visible atoms) |
| Save Image… | — | Opens a file-save dialog and writes the current view to a PNG or JPEG file |
| Parts | All | Filter displayed disorder parts; shown when multiple part values are present |
When Pack Unit Cell is active, a unit-cell axis indicator (a = red, b = green, c = blue) is drawn in the bottom-left corner of the widget and rotates with the view.
API Overview
MoleculeViewer3DWidget(parent=None)
A self-contained 3D viewer combining MoleculeWidget3D with the control bar.
load_file(path)— load a structure file (format auto-detected from extension:.cif,.res,.ins,.xyz)grow()— expand the asymmetric unit to complete molecules using crystal symmetry; deactivates Pack Unit Cell if active; no-op for XYZ files or when no file is loadedset_bond_color(color)— set the default color for non-selected bondsrender_widget— read-only property exposing the underlyingMoleculeWidget3D
MoleculeWidget3D(parent=None)
Hardware-accelerated OpenGL renderer. A QOpenGLWidget (Qt ≥ 6) or QWidget subclass that can be dropped into any layout.
Rendering technique
| Primitive | Technique |
|---|---|
| Atoms | Billboard sphere impostors — each atom is a quad; the fragment shader ray-casts a sphere and writes corrected depth values |
| ADP ellipsoids | Impostor quads — the fragment shader ray-casts an exact ellipsoid using the inverse U_cart tensor passed as a mat3 uniform |
| Bonds | Tessellated cylinder mesh (8-segment, 4-segment for angular style) built on the CPU and uploaded as a single VBO |
| Labels | QPainter overlay drawn after the OpenGL pass |
GLSL shader targets are platform-aware: #version 120 on macOS (OpenGL 2.1 / GLSL 1.20) and #version 140 on Windows/Linux (OpenGL 3.1+ / GLSL 1.40).
Qt Signals
| Signal | Signature | Emitted when |
|---|---|---|
atomClicked |
(label: str) |
The user clicks on an atom |
bondClicked |
(label1: str, label2: str) |
The user clicks on a bond |
Data Methods
-
open_molecule(atoms, cell=None, keep_view=False)
Load a new set of atoms and redraw.atoms— list ofAtomtuple(label, type, x, y, z, part, adp=None)in Cartesian coordinates (Å); embedadp=(U11,U22,U33,U23,U13,U12)directly in the tuple for anisotropic atomscell— optional(a, b, c, α, β, γ)tuple; required for ADP renderingkeep_view— preserve current zoom, rotation, and pan whenTrue
-
grow_molecule(atoms, cell=None)
Replace atoms while preserving the view. Equivalent toopen_molecule(..., keep_view=True). -
clear()
Remove all atoms and bonds.
Display Methods
show_adps(value: bool)— toggle ADP ellipsoid rendering; falls back to isotropic spheres whenFalseshow_labels(value: bool)— show / hide atom labelsshow_hydrogens(value: bool)— show / hide hydrogen atoms and bondsset_visible_parts(parts: set[int] | None)— filter by disorder part;Noneshows all atoms; an empty set hides all atoms; e.g.set_visible_parts({0, 1})shows only Part 0 and Part 1set_bond_width(width: int)— set cylinder radius scale (0–15)set_bond_color(color)— set the default color for non-selected bonds; acceptsQColor, hex string, or an RGB tupleset_labels_visible(visible: bool)— alias forshow_labelssetLabelFont(font_size: int)— set label font pixel sizeset_background_color(color: QColor)— change background colourreset_view()— reset zoom, rotation, and pan to defaultsalign_best_view()— rotate the structure so the widest face points towards the viewer (PCA on visible atoms; H/D excluded when hydrogen visibility is off)reset_rotation_center()— restore the rotation pivot to the molecule's geometric center (undoes a middle-click recentring)save_image(filename: Path, image_scale: float = 1.5)— capture the current OpenGL framebuffer and write it to a PNG or JPEG file (format inferred from the file extension). The captured image is then scaled byimage_scaleusing smooth bilinear filtering before saving. Labels appear in the saved image if they are active at the time of the call.
Example — feeding atom data directly to MoleculeWidget3D
from fastmolwidget import MoleculeWidget3D, Atomtuple
mol = MoleculeWidget3D(parent=self)
# Embed ADP tensors directly in each Atomtuple (None = isotropic / no ADP)
atoms = [
Atomtuple(label="C1", type="C", x=0.0, y=0.0, z=0.0, part=0,
adp=(0.02, 0.02, 0.02, 0.0, 0.0, 0.0)),
Atomtuple(label="O1", type="O", x=1.22, y=0.0, z=0.0, part=0,
adp=(0.03, 0.03, 0.03, 0.0, 0.0, 0.0)),
Atomtuple(label="H1", type="H", x=-0.5, y=0.94, z=0.0, part=0),
]
cell = (5.0, 5.0, 5.0, 90.0, 90.0, 90.0)
mol.open_molecule(atoms=atoms, cell=cell)
mol.atomClicked.connect(lambda label: print(f"Selected: {label}"))
layout.addWidget(mol)
MoleculeViewerWidget(parent=None)
A self-contained 2D viewer combining MoleculeWidget with the control bar.
load_file(path)— load a structure file (format auto-detected from extension)grow()— expand the asymmetric unit to complete molecules using crystal symmetry; deactivates Pack Unit Cell if active; no-op for XYZ files or when no file is loadedset_bond_color(color)— set the default color for non-selected bondsrender_widget— read-only property exposing the underlyingMoleculeWidget
MoleculeWidget(parent=None)
The 2D QPainter renderer. A plain QWidget subclass you can drop into any layout.
Qt Signals
| Signal | Signature | Emitted when |
|---|---|---|
atomClicked |
(label: str) |
The user clicks on an atom; label is the atom name (e.g. "C1") |
bondClicked |
(label1: str, label2: str) |
The user clicks on a bond; both atom labels are passed |
Data Methods
-
open_molecule(atoms, cell=None, keep_view=False)
Load a new set of atoms and reset (or optionally preserve) the view.atoms— list ofAtomtuple(label, type, x, y, z, part, adp=None)in Cartesian coordinates (Å); embedadp=(U11,U22,U33,U23,U13,U12)for anisotropic atomscell— optional(a, b, c, α, β, γ)tuple of unit-cell parameters (Å / °); required for ADP renderingkeep_view— whenTrue, the current zoom, pan, and rotation are preserved (useful for live updates)
-
grow_molecule(atoms, cell=None)
Replace the atom set while always preserving the current view.
Equivalent to callingopen_molecule(..., keep_view=True). -
clear()
Remove all atoms and bonds from the display.
Display Methods
-
show_adps(value: bool)
Toggle ORTEP-style ADP ellipsoid rendering. WhenFalse, atoms are drawn as isotropic spheres. -
show_labels(value: bool)
Show or hide non-hydrogen atom labels. -
show_hydrogens(value: bool)
Show or hide hydrogen / deuterium atoms and their bonds. -
set_visible_parts(parts: set[int] | None)
Filter by disorder part number.None(the default) shows all parts. Pass a set of integers to restrict rendering to those parts; an empty set hides every atom. Example:widget.set_visible_parts({0, 1})shows Part 0 and Part 1. -
set_bond_width(width: int)
Set the stroke width for bonds in pixels (valid range: 1–15). -
set_bond_color(color)
Set the default color for non-selected bonds. AcceptsQColor, hex string (e.g."#d1812a"), or an RGB tuple (floats in[0..1]or integers in[0..255]). -
set_labels_visible(visible: bool)
Alias forshow_labels. -
setLabelFont(font_size: int)
Set the pixel size used for atom labels. -
set_background_color(color: QColor)
Change the widget background color. -
reset_view()
Reset zoom, pan, and rotation to their defaults. -
align_best_view()
Rotate the structure to the orientation that maximises atom visibility for screenshots. Uses PCA on the currently visible atom positions: the thinnest axis of the atom cloud points towards the camera so the widest face faces the viewer. Hydrogen / deuterium atoms are excluded when their visibility is turned off. -
save_image(filename: Path, image_scale: float = 1.5)
Render the current structure view to an image file.
The widget is redrawn off-screen atwidget_size × image_scale; the result is saved as PNG or JPEG (format inferred from the file extension).
Labels appear in the saved image if they are active at the time of the call.
Example — feeding atom data directly to MoleculeWidget (2D)
from fastmolwidget import MoleculeWidget, Atomtuple
mol = MoleculeWidget(parent=self)
# Embed ADP tensors directly in each Atomtuple (omit or use None = isotropic)
atoms = [
Atomtuple(label="C1", type="C", x=0.0, y=0.0, z=0.0, part=0,
adp=(0.02, 0.02, 0.02, 0.0, 0.0, 0.0)),
Atomtuple(label="O1", type="O", x=1.22, y=0.0, z=0.0, part=0,
adp=(0.03, 0.03, 0.03, 0.0, 0.0, 0.0)),
Atomtuple(label="H1", type="H", x=-0.5, y=0.94, z=0.0, part=0),
]
cell = (5.0, 5.0, 5.0, 90.0, 90.0, 90.0) # optional
mol.open_molecule(atoms=atoms, cell=cell)
mol.atomClicked.connect(lambda label: print(f"Selected: {label}"))
layout.addWidget(mol)
Advanced API
MoleculeWidgetProtocol
The core rendering interface is defined by MoleculeWidgetProtocol. Both MoleculeWidget (2D) and MoleculeWidget3D (3D) satisfy this protocol, making them drop-in replacements for each other.
from fastmolwidget.molecule_base import MoleculeWidgetProtocol
from fastmolwidget import MoleculeWidget3D
def do_something_with_widget(widget: MoleculeWidgetProtocol):
...
3D Application Example
import sys
from qtpy.QtWidgets import QApplication
from fastmolwidget import MoleculeViewer3DWidget
app = QApplication(sys.argv)
viewer = MoleculeViewer3DWidget()
viewer.load_file("examples/test_molecule.res")
viewer.show()
sys.exit(app.exec_())
3D Generic Widget Example
import sys
from qtpy.QtWidgets import QApplication, QMainWindow, QVBoxLayout, QWidget
from fastmolwidget import MoleculeWidget3D
from fastmolwidget.loader import MoleculeLoader
app = QApplication(sys.argv)
main_window = QMainWindow()
central_widget = QWidget(main_window)
layout = QVBoxLayout(central_widget)
# Create and configure the 3D molecule widget
molecule_widget = MoleculeWidget3D()
molecule_widget.set_bond_color("#FF5733") # Example: set bond color to a shade of orange
# Load a molecule file (CIF, RES, or XYZ format)
loader = MoleculeLoader(molecule_widget)
loader.load_file("examples/test_molecule.res")
layout.addWidget(molecule_widget)
main_window.setCentralWidget(central_widget)
main_window.show()
sys.exit(app.exec_())
Running the Examples
To run the provided examples, you can use the following commands:
# 2D Viewer example
python -m fastmolwidget.examples.viewer_2d_example
# 3D Viewer example
python -m fastmolwidget.examples.viewer_3d_example
# Generic 3D Widget example
python -m fastmolwidget.examples.generic_3d_widget_example
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file fastmolwidget-0.9.7-cp314-cp314-win_amd64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp314-cp314-win_amd64.whl
- Upload date:
- Size: 179.3 kB
- Tags: CPython 3.14, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8abc4c2d203ff02384a75636bd20d1a5b0516e798e6866d0ea808893e22fbb40
|
|
| MD5 |
a9afe7e8a220836c8b95be9186fd597c
|
|
| BLAKE2b-256 |
d6e8ac0f5e07af4bedfe5a25e6d316c4082d3f57f887d311042e670b4fc5f5b5
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp314-cp314-win_amd64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp314-cp314-win_amd64.whl -
Subject digest:
8abc4c2d203ff02384a75636bd20d1a5b0516e798e6866d0ea808893e22fbb40 - Sigstore transparency entry: 1519685308
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp314-cp314-musllinux_1_2_x86_64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp314-cp314-musllinux_1_2_x86_64.whl
- Upload date:
- Size: 2.5 MB
- Tags: CPython 3.14, musllinux: musl 1.2+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
481047ac4ddfa24fb16015ca530b60422e88b65df5618b634504b22d74287ed2
|
|
| MD5 |
86d0d38d0fcd4811f9ba36c7aadef345
|
|
| BLAKE2b-256 |
d29b4a0fb3f3fbdcb5cf12866e29fc5f35e25dc70370699b58704303d9bc39cd
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp314-cp314-musllinux_1_2_x86_64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp314-cp314-musllinux_1_2_x86_64.whl -
Subject digest:
481047ac4ddfa24fb16015ca530b60422e88b65df5618b634504b22d74287ed2 - Sigstore transparency entry: 1519685202
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp314-cp314-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp314-cp314-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl
- Upload date:
- Size: 1.4 MB
- Tags: CPython 3.14, manylinux: glibc 2.24+ x86-64, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
212398ad7d176ce2fc1bf9a333b5872a27bde902b48f85af6c6e395f1737b9a2
|
|
| MD5 |
4110569e0c9c04cd3931c1710a658f5f
|
|
| BLAKE2b-256 |
00e83784334f116c2d0cf11fc83e7dfdfb03b75a4f279f5d65f1a94745742a7c
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp314-cp314-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp314-cp314-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl -
Subject digest:
212398ad7d176ce2fc1bf9a333b5872a27bde902b48f85af6c6e395f1737b9a2 - Sigstore transparency entry: 1519685536
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp314-cp314-macosx_14_0_universal2.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp314-cp314-macosx_14_0_universal2.whl
- Upload date:
- Size: 262.4 kB
- Tags: CPython 3.14, macOS 14.0+ universal2 (ARM64, x86-64)
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c86b00b88c179870460b68b5b0103f426338ad29be3012b238be93938e1a2fa6
|
|
| MD5 |
dcb497888fd14b7a13205a1c9306f45d
|
|
| BLAKE2b-256 |
67a05939f012f0b066a3444eae3450f8e5ab616f00e5173e6c1c495a1e452e91
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp314-cp314-macosx_14_0_universal2.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp314-cp314-macosx_14_0_universal2.whl -
Subject digest:
c86b00b88c179870460b68b5b0103f426338ad29be3012b238be93938e1a2fa6 - Sigstore transparency entry: 1519685482
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp313-cp313-win_amd64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp313-cp313-win_amd64.whl
- Upload date:
- Size: 176.5 kB
- Tags: CPython 3.13, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5fe68241b15c42d5b19ce8a3448fcdcbe46d5940ec227c54ccaee68a2998e4be
|
|
| MD5 |
281f40a988c2256d1ce38a3286d5ff4d
|
|
| BLAKE2b-256 |
9c88f361b7cd6bcd02b65645f5ff1d05f477f7fa75c0ee9ce985264178e2a84c
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp313-cp313-win_amd64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp313-cp313-win_amd64.whl -
Subject digest:
5fe68241b15c42d5b19ce8a3448fcdcbe46d5940ec227c54ccaee68a2998e4be - Sigstore transparency entry: 1519685276
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp313-cp313-musllinux_1_2_x86_64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp313-cp313-musllinux_1_2_x86_64.whl
- Upload date:
- Size: 2.5 MB
- Tags: CPython 3.13, musllinux: musl 1.2+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cc7e78e855346fc3e0c7f961256708dcdad3d078a79aea405f596d9bb67e4512
|
|
| MD5 |
a8e537f663a1bad79625c78cb0a1b416
|
|
| BLAKE2b-256 |
d57aaabe3142ac98da1a8b88ab4ef5a20f5f74a55894fae115ab4688d32535d0
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp313-cp313-musllinux_1_2_x86_64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp313-cp313-musllinux_1_2_x86_64.whl -
Subject digest:
cc7e78e855346fc3e0c7f961256708dcdad3d078a79aea405f596d9bb67e4512 - Sigstore transparency entry: 1519685186
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp313-cp313-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp313-cp313-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl
- Upload date:
- Size: 1.4 MB
- Tags: CPython 3.13, manylinux: glibc 2.24+ x86-64, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
64fe933745cbf8ac07a987119b6bede0a2edf1e3e2630f93a704e6a0a7324007
|
|
| MD5 |
8a26bedfd78e7d089c305b77ae74345c
|
|
| BLAKE2b-256 |
e6ab5f4db7645c403ff9495eaabb718878fb4de2f7d879a9fb526ffbab4d557a
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp313-cp313-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp313-cp313-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl -
Subject digest:
64fe933745cbf8ac07a987119b6bede0a2edf1e3e2630f93a704e6a0a7324007 - Sigstore transparency entry: 1519685250
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp313-cp313-macosx_14_0_universal2.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp313-cp313-macosx_14_0_universal2.whl
- Upload date:
- Size: 262.4 kB
- Tags: CPython 3.13, macOS 14.0+ universal2 (ARM64, x86-64)
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5b66fae4e237866d04fa3d9e1d4d80ee1bea6c7459d8f569ab549092fa5e3669
|
|
| MD5 |
4aa335b36f4a67a703b2d09be5a12347
|
|
| BLAKE2b-256 |
e8190b7c865f47bd620b795a697257077b0de7c553ffbc962a429cff10131552
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp313-cp313-macosx_14_0_universal2.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp313-cp313-macosx_14_0_universal2.whl -
Subject digest:
5b66fae4e237866d04fa3d9e1d4d80ee1bea6c7459d8f569ab549092fa5e3669 - Sigstore transparency entry: 1519685172
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp312-cp312-win_amd64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp312-cp312-win_amd64.whl
- Upload date:
- Size: 176.5 kB
- Tags: CPython 3.12, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4f9602bc6c7ab6468c1f4356b823b4bd95a41a28ec5683ee8fbb508b0e717d9e
|
|
| MD5 |
e1b42717793fe621dbd573fa2aefbf74
|
|
| BLAKE2b-256 |
981d2996591288cec2d09dc355ddce67794e5d5e80af9cc8e38667e511820480
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp312-cp312-win_amd64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp312-cp312-win_amd64.whl -
Subject digest:
4f9602bc6c7ab6468c1f4356b823b4bd95a41a28ec5683ee8fbb508b0e717d9e - Sigstore transparency entry: 1519685213
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp312-cp312-musllinux_1_2_x86_64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp312-cp312-musllinux_1_2_x86_64.whl
- Upload date:
- Size: 2.5 MB
- Tags: CPython 3.12, musllinux: musl 1.2+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f0f19d9a20b97b8b16c039949b132dc01f455bd845019606f0a42cb9db0d6918
|
|
| MD5 |
ee3694c6aa822d45cf07f6ee5de345f8
|
|
| BLAKE2b-256 |
d8a253382cee4bf7c81f9a65493795b3d705060dd04cee5572d38db9e7185e75
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp312-cp312-musllinux_1_2_x86_64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp312-cp312-musllinux_1_2_x86_64.whl -
Subject digest:
f0f19d9a20b97b8b16c039949b132dc01f455bd845019606f0a42cb9db0d6918 - Sigstore transparency entry: 1519685395
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp312-cp312-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp312-cp312-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl
- Upload date:
- Size: 1.4 MB
- Tags: CPython 3.12, manylinux: glibc 2.24+ x86-64, manylinux: glibc 2.28+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c71da72a28a1f4369cbbc936b55a7f04a2107ec7c5261c8e1a1b67c9a3f71cd5
|
|
| MD5 |
685c7a9f98028cc6cac60069a3c27daa
|
|
| BLAKE2b-256 |
4e680fb4af119f4cca670fddceb926d21bb97909dcd76dd9852541321aeaf378
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp312-cp312-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp312-cp312-manylinux_2_24_x86_64.manylinux_2_28_x86_64.whl -
Subject digest:
c71da72a28a1f4369cbbc936b55a7f04a2107ec7c5261c8e1a1b67c9a3f71cd5 - Sigstore transparency entry: 1519685459
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
File details
Details for the file fastmolwidget-0.9.7-cp312-cp312-macosx_14_0_universal2.whl.
File metadata
- Download URL: fastmolwidget-0.9.7-cp312-cp312-macosx_14_0_universal2.whl
- Upload date:
- Size: 262.3 kB
- Tags: CPython 3.12, macOS 14.0+ universal2 (ARM64, x86-64)
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
293ec1151c98d23fa6439155ee43a1cc9bc72b6d2d20e7f0049d8456a5152401
|
|
| MD5 |
d0b6d95fbca56e2a06eeeaf8d69ffb3a
|
|
| BLAKE2b-256 |
66b030dc05c34a7e8904eb3c056be08ed6ef45b6f3d8c53d4d91430ec1530c48
|
Provenance
The following attestation bundles were made for fastmolwidget-0.9.7-cp312-cp312-macosx_14_0_universal2.whl:
Publisher:
python-publish.yml on dkratzert/Fastmolwidget
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
fastmolwidget-0.9.7-cp312-cp312-macosx_14_0_universal2.whl -
Subject digest:
293ec1151c98d23fa6439155ee43a1cc9bc72b6d2d20e7f0049d8456a5152401 - Sigstore transparency entry: 1519685231
- Sigstore integration time:
-
Permalink:
dkratzert/Fastmolwidget@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Branch / Tag:
refs/tags/version-0.9.7 - Owner: https://github.com/dkratzert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@31bc75e991909ac922b8c8f60daf988b2da96db3 -
Trigger Event:
push
-
Statement type:
















