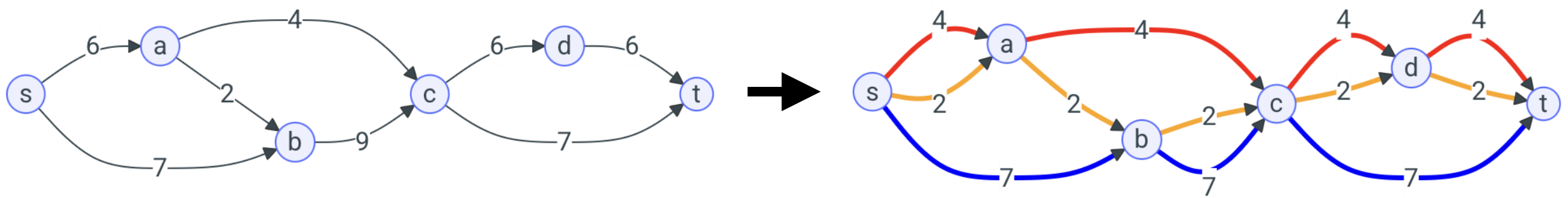
A Python package to quickly decompose weighted graphs into weights paths, under various models.
Project description
The flowpaths Python Package
This package implements various solvers for decomposing a weighted directed acyclic graph (DAG) into weighted paths, based on (Mixed) Integer Linear Programming ((M)ILP) formulations. It also supports the easy creation of solvers for new decomposition models.
Installation
pip install flowpaths
Documentation
The documentation of this package is currently being written at algbio.github.io/flowpaths/.
Basic usage example:
import flowpaths as fp
import networkx as nx
# Create a simple graph
graph = nx.DiGraph()
graph.add_edge("s", "a", flow=2)
graph.add_edge("a", "t", flow=2)
graph.add_edge("s", "b", flow=5)
graph.add_edge("b", "t", flow=5)
# ...
# Create a Minimum Flow Decomposition solver
mfd_solver = fp.MinFlowDecomp(graph, flow_attr="flow")
mfd_solver.solve() # We solve it
if mfd_solver.is_solved(): # We get the solution
print(mfd_solver.get_solution())
# {'paths': [['s', 'b', 't'], ['s', 'a', 't']], 'weights': [5, 2]}
Design principles
-
Easy to use: You just pass a directed graph to the solvers (as a networkx DiGraph), and they return optimal weighted paths. See the examples folder for some usage examples.
-
It just works: You do not need to install an (M)ILP solver. This is possible thanks to the fast open source solver HiGHS, which gets installed once you install this package.
- If you have a Gurobi license (free for academic users), you can install the gurobipy Python package, and then you can run the Gurobi solver instead of the default HiGHS solver by just passing the entry
"external_solver": "gurobi"in thesolver_optionsdictionary.
- If you have a Gurobi license (free for academic users), you can install the gurobipy Python package, and then you can run the Gurobi solver instead of the default HiGHS solver by just passing the entry
-
Easy to implement other decomposition models: We provide an abstract class modeling a generic path-finding MILP (
AbstractPathModelDAG), which encodes a given number of arbitrary paths in the DAG. You can inherit from this class to add e.g. weights to the paths, and specify various constraints that these weighted paths must satisfy, or the objective function they need to minimize or maximize. See a basic example of a solver implemented in this manner. This abstract class interfaces with a wrapper for both MILP solvers, so you do not need to worry about MILP technicalities. The decomposition solvers already implemented in this package use this wrapper. -
Fast: Having solvers implemented using
AbstractPathModelDAGmeans that any optimization to the path-finding mechanisms benefits all solvers that inherit from this class. We implement some "safety optimizations" described in this paper, based on ideas first introduced in this paper, which can provide up to 1000x speedups, depending on the graph instance, while preserving global optimality (under some simple assumptions).
Models currently implemented:
- Minimum Flow Decomposition: Given a DAG with flow values on its edges (i.e. at every node different from source or sink the flow enetering the node is equal to the flow exiting the node), find the minimum number of weighted paths such that, for every edge, the sum of the weights of the paths going through the edge equals the flow value of the edge.
- $k$-Least Absolute Errors: Given a DAG with weights on its edges, and a number $k$, find $k$ weighted paths such that the sum of the absolute errors of each edge is minimized.
- The error of an edge is defined as the weight of the edge minus the sum of the weights of the paths going through it.
- $k$-Minimum Path Error: Given a DAG with weights on its edges, and a number $k$, find $k$ weighted paths, with associated slack values, such that:
- The error of each edge (defined as in $k$-Least Absolute Errors above) is at most the sum of the slacks of the paths going through the edge, and
- The sum of path slacks is minimized.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file flowpaths-0.1.18.tar.gz.
File metadata
- Download URL: flowpaths-0.1.18.tar.gz
- Upload date:
- Size: 57.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f18212a220bf304895de761fa19e0b0b677bbe22bfd7efe7c0434e5837090cdb
|
|
| MD5 |
2314d4241524cd465f8b9483c7c94194
|
|
| BLAKE2b-256 |
3b1c549fa9b535f32dd8064c9f31ab5115834a625e196755602f612032eb6cd9
|
Provenance
The following attestation bundles were made for flowpaths-0.1.18.tar.gz:
Publisher:
python-publish.yml on algbio/flowpaths
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
flowpaths-0.1.18.tar.gz -
Subject digest:
f18212a220bf304895de761fa19e0b0b677bbe22bfd7efe7c0434e5837090cdb - Sigstore transparency entry: 192344819
- Sigstore integration time:
-
Permalink:
algbio/flowpaths@3fd4b020595646a4c1fc0734936b2deca1cfeb4a -
Branch / Tag:
refs/heads/main - Owner: https://github.com/algbio
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@3fd4b020595646a4c1fc0734936b2deca1cfeb4a -
Trigger Event:
push
-
Statement type:
File details
Details for the file flowpaths-0.1.18-py3-none-any.whl.
File metadata
- Download URL: flowpaths-0.1.18-py3-none-any.whl
- Upload date:
- Size: 69.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a8a996ad164e520a0bea2e9eba586b2f3ba8489e5c4de90f1bd74e519f8af573
|
|
| MD5 |
9ba91e34f045912a2883d4e6a4ba978c
|
|
| BLAKE2b-256 |
e531e283841ac7535203994890f1762c0c83b22bf4f7d0caf48452e9d17953da
|
Provenance
The following attestation bundles were made for flowpaths-0.1.18-py3-none-any.whl:
Publisher:
python-publish.yml on algbio/flowpaths
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
flowpaths-0.1.18-py3-none-any.whl -
Subject digest:
a8a996ad164e520a0bea2e9eba586b2f3ba8489e5c4de90f1bd74e519f8af573 - Sigstore transparency entry: 192344822
- Sigstore integration time:
-
Permalink:
algbio/flowpaths@3fd4b020595646a4c1fc0734936b2deca1cfeb4a -
Branch / Tag:
refs/heads/main - Owner: https://github.com/algbio
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@3fd4b020595646a4c1fc0734936b2deca1cfeb4a -
Trigger Event:
push
-
Statement type: