GASTON: interpretable deep learning model for spatial transcriptomics
Project description
GASTON
GASTON is an interpretable deep learning model for learning a topographic map of a tissue slice from spatially resolved transcriptomics (SRT) data. Specifically, GASTON models gene expression topography by learning the isodepth, a 1-D coordinate that smoothly varies across a tissue slice.
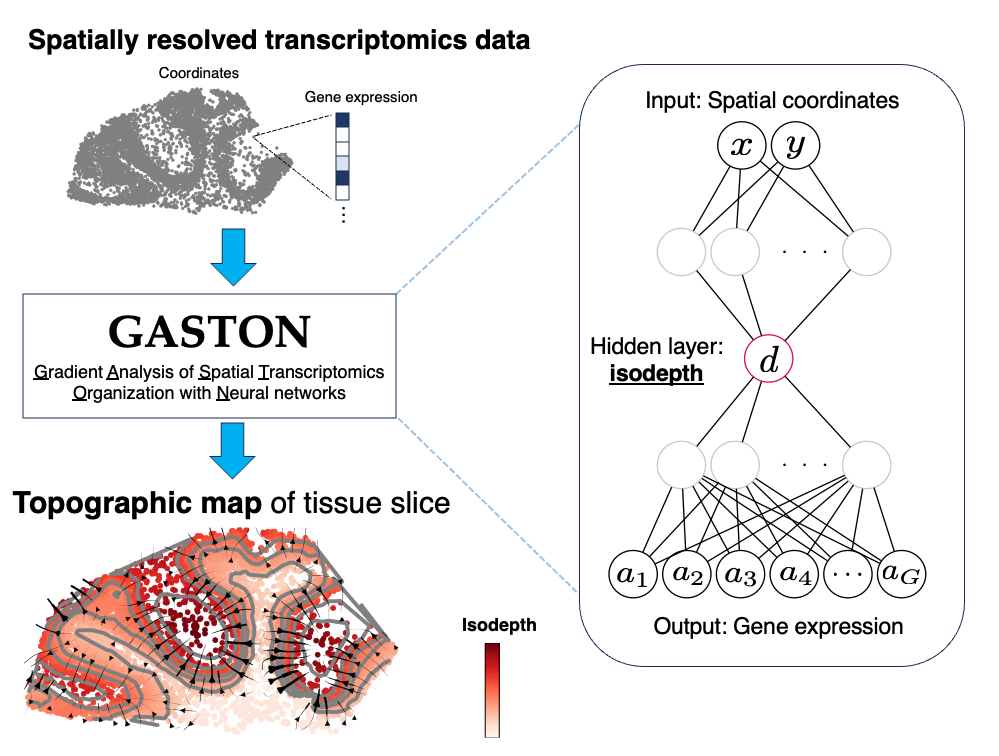
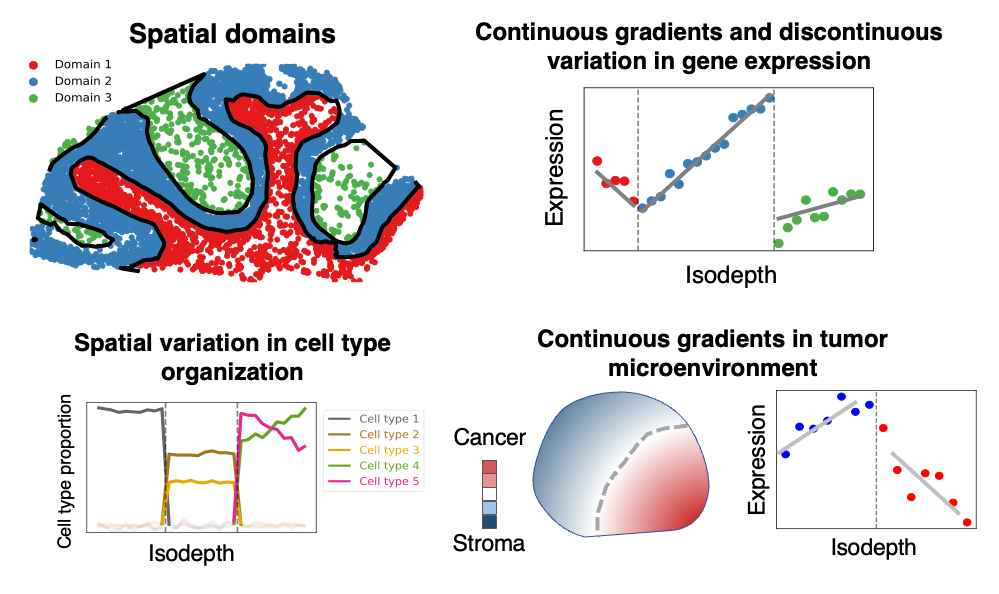
The isodepth and topographic map learned by GASTON have many downstream applications including
- identifying spatial domains,
- inferring continuous gradients and discontinuous jumps in gene expression,
- deriving maps of spatial variation in cell type organization,
- analyzing continuous gradients in the tumor microenvironment
Installation
GASTON is on pypi: https://pypi.org/project/gaston-spatial/.
pip install gaston-spatial
You can also directly install the conda environment from the environment.yml file:
cd GASTON
conda env create -f environment.yml
Then install GASTON using pip.
conda activate gaston-package
pip install -e .
Installation should take <10 minutes.
We will add GASTON to bioconda soon!
Getting started
Check out our readthedocs, which includes tutorials for four analyses:
- mouse cerebellum (Slide-SeqV2)
- human colorectal tumor (10x Visium)
- human breast cancer tumor (10x Xenium)
- mouse cortex (MERFISH).
Training GASTON by following the tutorials should take <1 hour.
GASTON requires an NxG gene expression matrix (e.g. UMI counts) and an Nx2 spatial location matrix, which can be provided or read from Space Ranger output.
The data to run the tutorials is located in docs/notebooks/tutorials/. Note that due to Github size constraints, you have to download the counts matrices for both analyses from Google Drive.
Software dependencies
- torch (=2.0.0)
- matplotlib (=3.8.0)
- pandas (=2.1.1)
- scikit-learn (=1.3.1)
- numpy (=1.23.4)
- jupyterlab (=4.0.6)
- seaborn (=0.12.2)
- tqdm (=4.66.1)
- scipy (=1.11.2)
- scanpy (=1.9.5)
- squidpy (=1.3.1)
See full list in environment.yml file. GASTON can be run on CPU or GPU.
Citations
The GASTON manuscript is published at Nature Methods. If you use GASTON for your work, please cite our paper.
@article{Chitra2025,
date = {2025/01/23},
date-added = {2025-01-27 12:52:11 -0500},
date-modified = {2025-01-27 12:52:11 -0500},
doi = {10.1038/s41592-024-02503-3},
id = {Chitra2025},
isbn = {1548-7105},
journal = {Nature Methods},
title = {Mapping the topography of spatial gene expression with interpretable deep learning},
url = {https://doi.org/10.1038/s41592-024-02503-3},
year = {2025},
bdsk-url-1 = {https://doi.org/10.1038/s41592-024-02503-3}}
Support
For questions or comments, please file a Github issue and/or email Uthsav Chitra (uchitra@princeton.edu).
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gaston_spatial-0.0.3.tar.gz.
File metadata
- Download URL: gaston_spatial-0.0.3.tar.gz
- Upload date:
- Size: 29.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f6fc88b983b066391c2e77c42dd7144d49a60843219058a77868162c62c40189
|
|
| MD5 |
702fbcc10c38fa174066c25496661d4b
|
|
| BLAKE2b-256 |
0ea3f9cbc4763c132ba85c2cb31de889d010c8eb2c83236d65fc6d9be44fdc16
|
File details
Details for the file gaston_spatial-0.0.3-py3-none-any.whl.
File metadata
- Download URL: gaston_spatial-0.0.3-py3-none-any.whl
- Upload date:
- Size: 32.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d8cf6a88cf051d11c3e3f3f95eaa58a1e6720bdfa5745f6524506a1b7277cf56
|
|
| MD5 |
7599ed0e127bb10ab16728b70cbe545d
|
|
| BLAKE2b-256 |
ef8ed9d1b00063552d7a6ae8e8aa99c00f6f8b1d10a8a52bc6e03ebbd1349acb
|











