gbapy (Growth Balance Analysis for Python)
Project description

Growth Balance Analysis for Python
pip install gba
gbapy is a Python package that provides tools for building and analyzing self-replicating cell (SRC) models based on the growth balance analysis (GBA) mathematical formalism (Dourado et al. 2023).
Features
The module offers two core components:
- :wrench: A builder class, to construct SRC models of any size from first principles,
- :chart_with_upwards_trend: A model class, to manipulate and optimize models once they are built.
Typical workflow
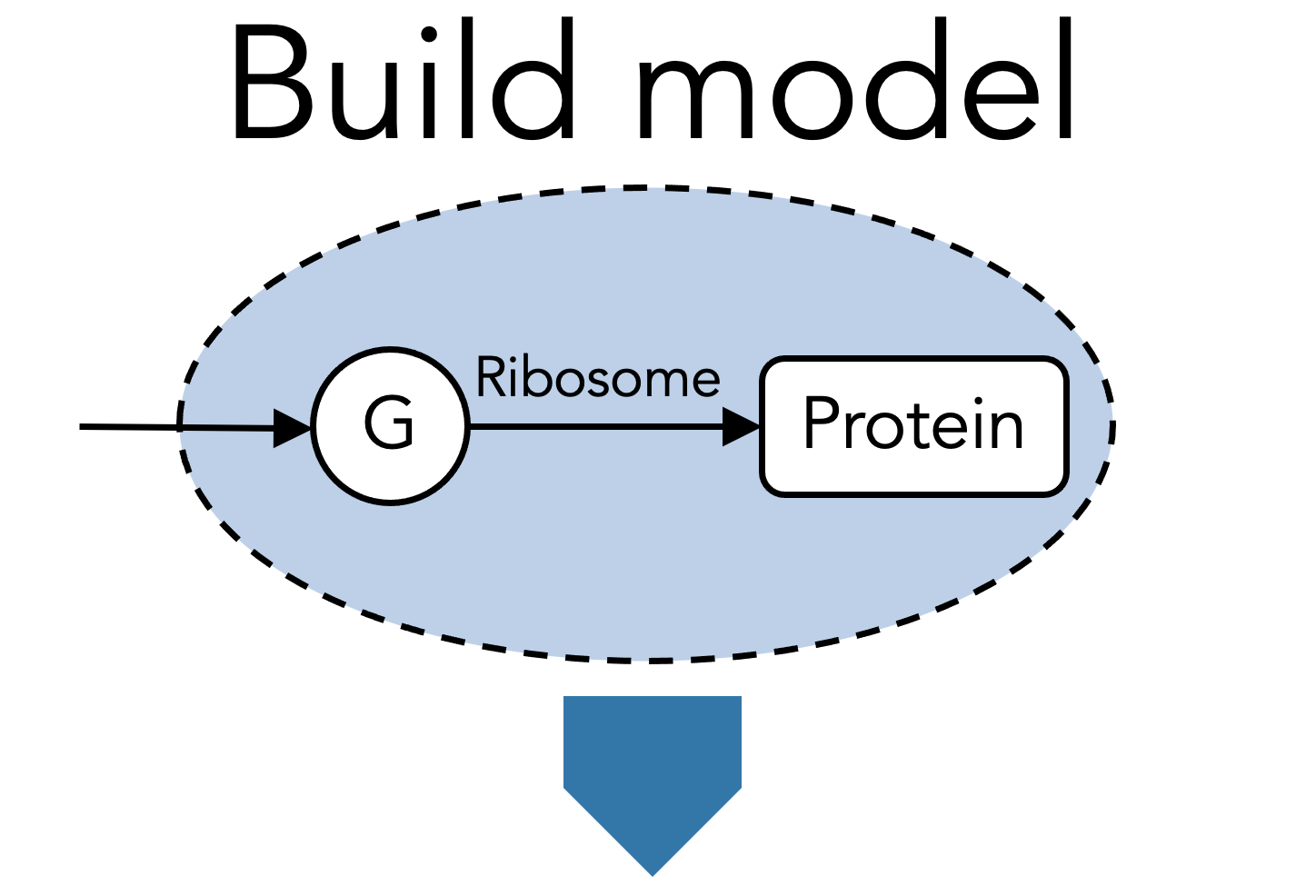
from gba import Builder, Model, Protein, Metabolite, Reaction
from gba import SpeciesLocation, ReactionType, ReactionDirection
builder = Builder(name="toy")
### Add model information (ODS sheet 'Info')
builder.add_info(category="General", key="Name", content="toy")
builder.add_info(category="General", key="Description", content="Toy model")
### Create and add proteins (one protein per enzyme per reaction)
### - Masses in Da.
p1 = Protein(id="p1", mass=1000000.0)
p2 = Protein(id="p2", mass=1000000.0)
builder.add_proteins([p1, p2])
### Create and add metabolites:
### - External and internal glucose
### - One generic Protein product
### - Masses in Da.
x_G = Metabolite(id="x_G", species_location=SpeciesLocation.EXTERNAL, mass=180.0)
G = Metabolite(id="G", species_location=SpeciesLocation.INTERNAL, mass=180.0)
Protein = Metabolite(id="Protein", species_location=SpeciesLocation.INTERNAL,mass=180.0)
builder.add_metabolites([x_G, G, Protein])
### Create and add transporter to import glucose:
### - Enzyme is composed of one protein p1
### - Reaction is irreversible
### - kcat values in 1/h
### - KM values in g/L
rxn1 = Reaction(id="rxn1", lb=0.0, ub=1000.0,
reaction_type=ReactionType.TRANSPORT,
metabolites={"x_G":-1.0, "G": 1.0},
proteins={"p1": 1.0})
rxn1.add_kcat_value(direction=ReactionDirection.FORWARD, kcat_value=45000.0)
rxn1.add_km_value(metabolite_id="x_G", km_value=0.00013)
rxn1.complete(kcat_value=0.0, km_value=0.0)
builder.add_reaction(rxn1)
### Create and add ribosome reaction to produce proteins:
### - Enzyme is composed of one protein p2
### - Reaction is irreversible
ribosome = Reaction(id="Ribosome", lb=0.0, ub=1000.0,
reaction_type=ReactionType.METABOLIC,
metabolites={"G":-1.0, "Protein": 1.0},
proteins={"p2": 1.0})
ribosome.add_kcat_value(direction=ReactionDirection.FORWARD, kcat_value=45000.0)
ribosome.add_km_value(metabolite_id="G", km_value=0.00013)
ribosome.complete(kcat_value=0.0, km_value=0.0)
builder.add_reaction(ribosome)
### Convert the model to GBA formalism (cf. Dourado et al. 2023)
builder.convert(ribosome_mass_kcat=4.55, ribosome_mass_km=8.3)
builder.build_GBA_model()
### Set cell's total density (g/L)
builder.set_rho(340.0)
### Create external conditions (in g/L)
x_G_conc = 1.0
for i in range(25):
builder.add_condition(condition_id=str(i+1), metabolites={"x_G": x_G_conc})
x_G_conc *= 2/3
### Save the model to an ODS file
builder.export_to_ods()

from gba import read_ods_model
### Load the ODS model
model = read_ods_model(name="toy")
### Find a valid initial solution
model.find_initial_solution()
### Optimize the model for all conditions
model.find_optimum_by_condition()
### Make a plot
model.plot(x="x_G", y="mu", title="Growth rate", logx=True)
### Export optimization data in CSV
model.export_optimization_data()
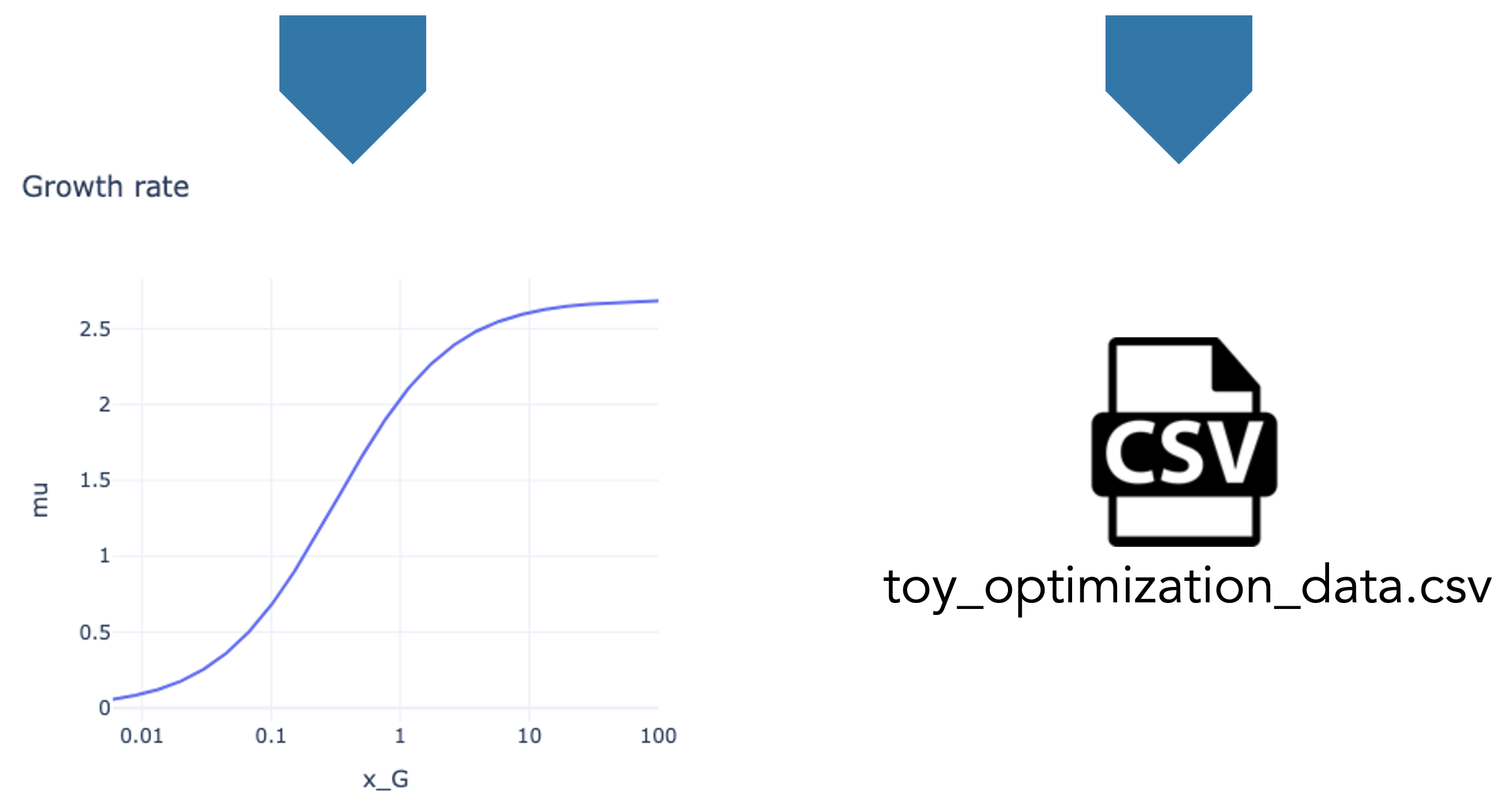
Reference files
Table of contents
1) Installation
The easiest way to install gbapy is from PyPI:
pip install gba
[!IMPORTANT] gbacpp software is required to run optimization tasks.
1.1) Supported platforms
gbapy has been primilary developed for Unix/Linux and macOS systems.
1.2) Dependencies
• Software
- gbacpp is required to run optimization tasks.
• Licensed Python modules
- The Python API of GUROBI optimizer must be installed and requires a user license (free for academics).
• Other Python modules
1.3) Manual installation
If you want to install gbapy manually, download the latest release, and save it to a directory of your choice. Open a terminal and use the cd command to navigate to this directory. Then follow the steps below to compile and build the executables.
sh install.sh
[!TIP] You can later uninstall the module using
sh uninstall.sh.
2) Tutorials
Tutorials coming soon ...
3) Documentation
Documentation coming soon ...
4) Contributing
If you wish to contribute, do not hesitate to reach the developer.
5) Copyright
Copyright © 2024-2026 Charles Rocabert, Furkan Mert.
6) License
This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program. If not, see http://www.gnu.org/licenses/.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gba-0.6.3.tar.gz.
File metadata
- Download URL: gba-0.6.3.tar.gz
- Upload date:
- Size: 60.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d3eee71f0cf81e26a571892e0b3ab5cad6602ccc1b6c52a196effb11069a0667
|
|
| MD5 |
5b1e3a08bd756de09c48069678dd1fef
|
|
| BLAKE2b-256 |
5f7f26db84d31e9a6e94557f05b49a95ea0707f9de5be946ade9fad5f0f72ffc
|
Provenance
The following attestation bundles were made for gba-0.6.3.tar.gz:
Publisher:
python-publish.yml on charlesrocabert/gbapy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
gba-0.6.3.tar.gz -
Subject digest:
d3eee71f0cf81e26a571892e0b3ab5cad6602ccc1b6c52a196effb11069a0667 - Sigstore transparency entry: 1018407980
- Sigstore integration time:
-
Permalink:
charlesrocabert/gbapy@bbe3fd7b35e4fc58155fcd1ddf6547ffd75e445a -
Branch / Tag:
refs/tags/v0.6.3 - Owner: https://github.com/charlesrocabert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@bbe3fd7b35e4fc58155fcd1ddf6547ffd75e445a -
Trigger Event:
release
-
Statement type:
File details
Details for the file gba-0.6.3-py3-none-any.whl.
File metadata
- Download URL: gba-0.6.3-py3-none-any.whl
- Upload date:
- Size: 60.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
97d44327aace8c25b0cde3fff8e6dca8548a0e53933228b7317bbb7bfc551677
|
|
| MD5 |
4b8183a1d538c692f23ca69f796b8e8b
|
|
| BLAKE2b-256 |
b221d1fa49cff3dfd6f5b4c5e532c62888d8f44affb04064142010278b34860c
|
Provenance
The following attestation bundles were made for gba-0.6.3-py3-none-any.whl:
Publisher:
python-publish.yml on charlesrocabert/gbapy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
gba-0.6.3-py3-none-any.whl -
Subject digest:
97d44327aace8c25b0cde3fff8e6dca8548a0e53933228b7317bbb7bfc551677 - Sigstore transparency entry: 1018408008
- Sigstore integration time:
-
Permalink:
charlesrocabert/gbapy@bbe3fd7b35e4fc58155fcd1ddf6547ffd75e445a -
Branch / Tag:
refs/tags/v0.6.3 - Owner: https://github.com/charlesrocabert
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@bbe3fd7b35e4fc58155fcd1ddf6547ffd75e445a -
Trigger Event:
release
-
Statement type:

















